There are many useful visualizations, annotations, and biological interpretation tools that can operate on a gene list. In order for these features work with an imported list, an annotation file must be associated with the gene list. Additionally, many operations that work with a list of significant genes (like GO- or Pathway-Enrichment) require comparison against a background of “non-significant” genes. The quickest way to accomplish both is to use the background of “all genes” for that organism provided by an annotation source like RefSeq, Ensembl, etc. in .pannot (Partek annotation), .gff, .gtf, .bed, tab- or comma-delimited format. If the file is not already in a tab-separated or comma delimited format, you may import, modify, and save the file in the proper file format.
Associating a spreadsheet with an annotation file
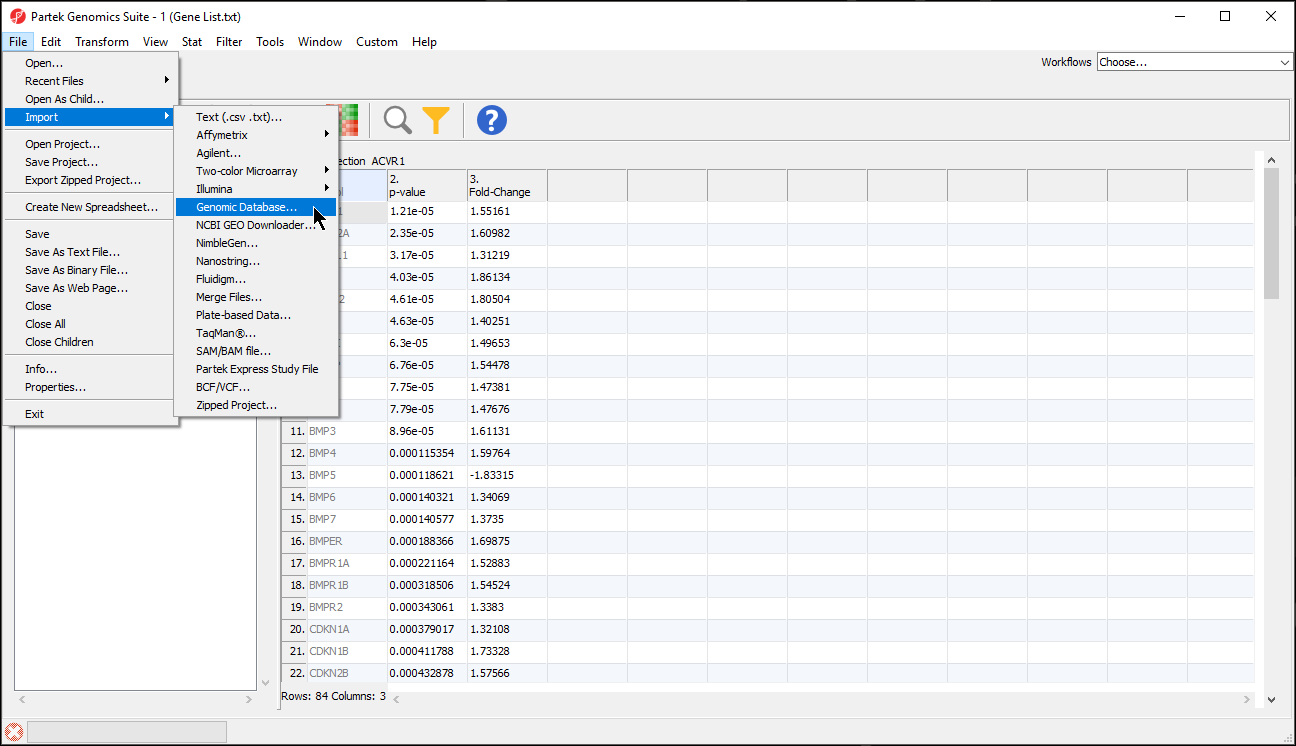
- Select File from the main toolbar
- Select Genomic Database under Import (Figure 1)
- Select the annotation file; in this example, we select a .pannot file downloaded from Partek distributed library file repository – hg19_refseq_14_01_03_v2.pannot
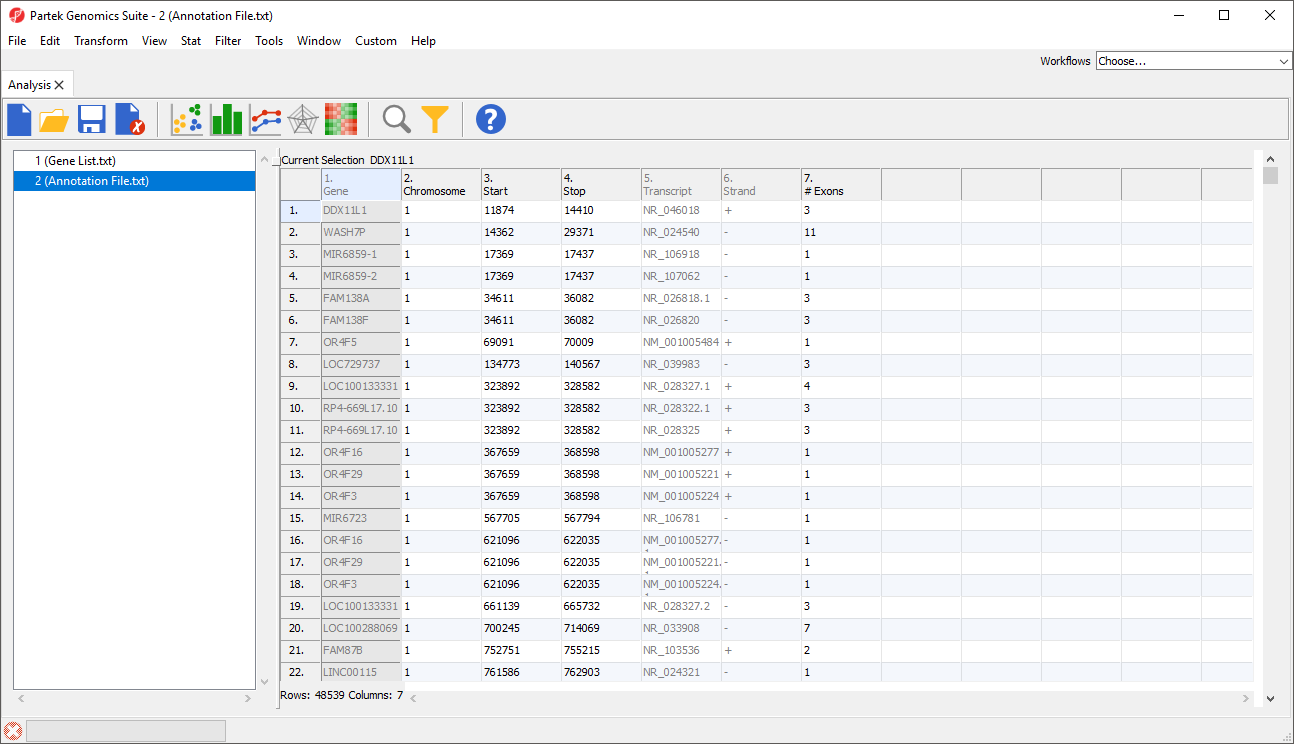
- Delete or rearrange the columns as necessary; we have placed the column with identifiers (should be unique ID) that correspond to our gene list first
- Select File then Save As Text File... to save the annotation file; we have named it Annotation File (Figure 2)
- Select () to close the annotation file
Now we can add the annotation file to our imported gene list.
- Right click 1 (gene_list.txt) in the spreadsheet tree
- Select Properties from the pop-up menu
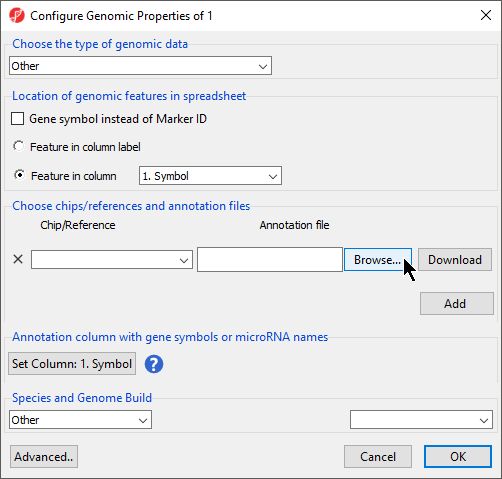
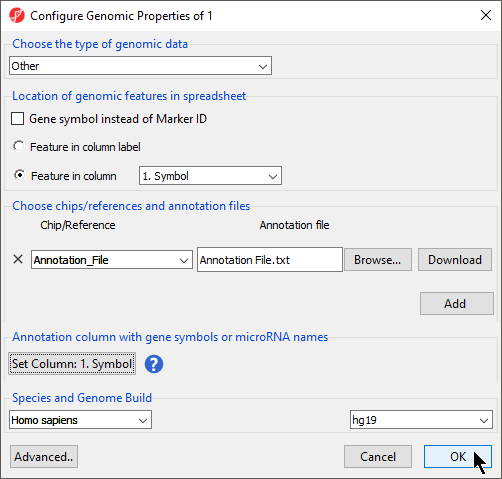
This brings up the Configure Genomic Properties dialog (Figure 3).
- Select Browse under Annotation File
- Choose the annotation file; we have chosen Annotation File.txt
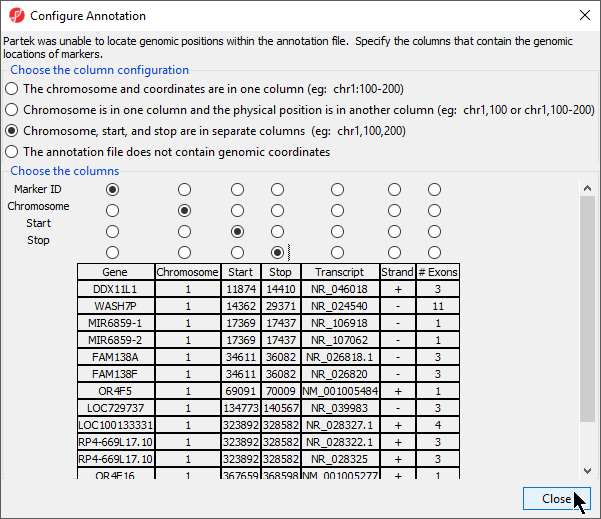
If this is the first time you have used an annotation, the Configure Annotation dialog will launch. This is used to choose the columns with the chromosome number and position information for each feature. Our example annotation file has chromosome, start, and stop in separate columns.
- Select the proper column configuration options (Figure 4)
- Select Close to return to the Configure Genomic Properties dialog
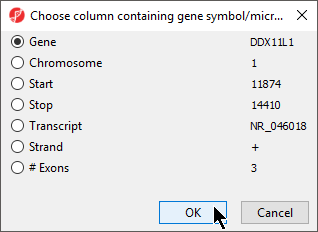
- Select Set Column: to open the Choose column with gene symbols or microRNA names dialog (Figure 5)
- Select the appropriate column; here the default choice of 1. Symbol is appropriate
- Select OK to return to the Configure Genomic Properties dialog
- Select the appropriate species and genome build options; we have selected Homo sapiens and hg19 (Figure 6)
- Select OK
- Select () to save the spreadsheet
The annotation file has been associated with the spreadsheet and additional tasks can now be performed on the data, e.g. since the annotation has genomic location, you can draw chromosome view on this data.
Adding annotations to a spreadsheet
Inserting annotations from an annotation file
If an annotation file has been associated with a spreadsheet, annotations from the file can be added as columns in the spreadsheet when each identifier is on a row.
- Right click on a column header
- Select Insert Annotation
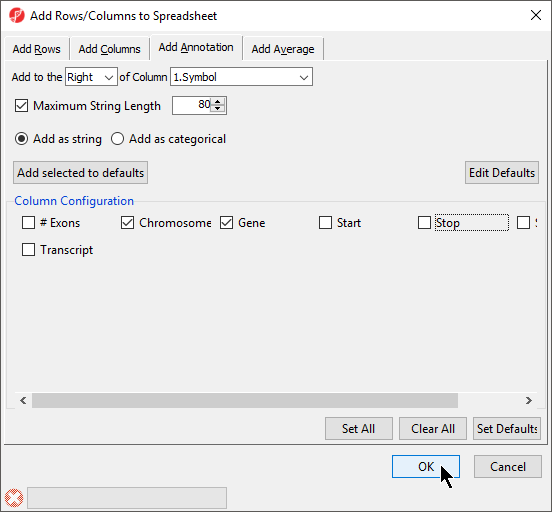
- Select columns to add from Column Configuration; we have selected Chromosome, Start, and Stop (Figure 7)
- Select OK
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
35 | rates |