This tutorial outlines how to analyze miRNA expression data in Partek Genomics Suite and outlines how miRNA expression data can be integrated with mRNA expression data from gene expression microarrays.
This tutorial illustrates how to:
Note: the workflow described below is enabled in Partek Genomics Suite version 7.0 software. Please fill out the form on Our support page to request this version or use the Help > Check for Updates command to check whether you have the latest released version. The screenshots shown within this tutorial may vary across platforms and across different versions of Partek Genomics Suite.
Description of the data set
The data set for this tutorial includes miRNA from 3 human brain samples and 3 heart samples quantified using the Affymetrix GeneChip miRNA 1.0 array. The same sample set was also processed on GeneChip Human Gene 1.0 ST arrays for mRNA expression.
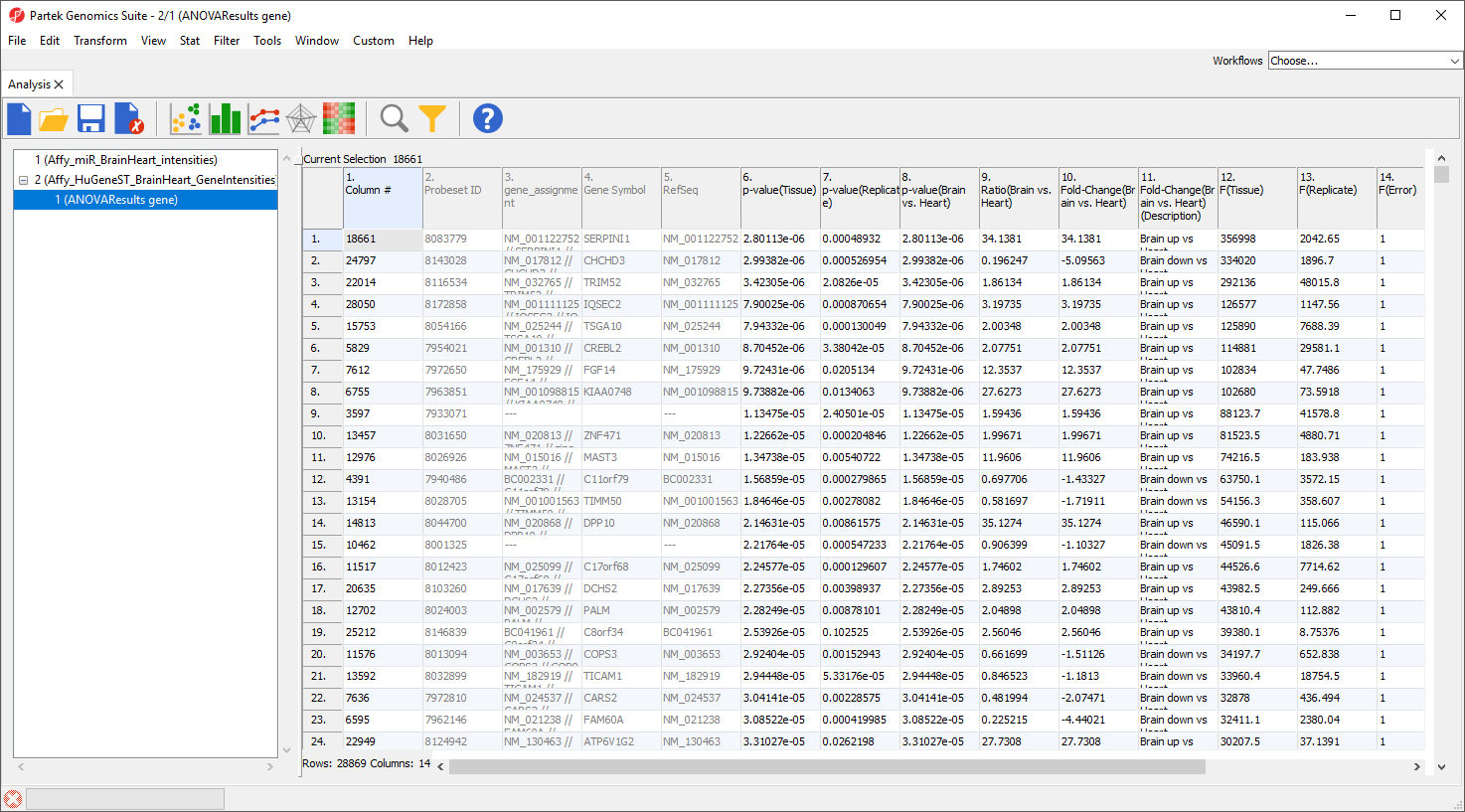
For this tutorial, the gene expression and miRNA expression studies have been analyzed and stored in Partek Genomics Suite project (ppj) format as miRNAmRNA integration. The project contains two Partek format files: Affy_miR_BrainHeart_intensities.fmt with the miRNA data and Affy_HuGeneST_BrainHeart_GeneIntensities.fmt with the analyzed mRNA data. There is also an ANOVA results spreadsheet open as a child spreadsheet of Affy_HuGeneST_BrainHeart_GeneIntensities.fmt.
- Download the miRNA Expression and Integration with Gene Expression data set and save it in an easily accessible location on your computer
We can now open the project in Partek Genomics Suite.
- Select File
- Select Import
- Select Zipped Project...
- Select the miRNA_tutorial_data.zip zipped folder
The project files will open in the Analysis tab (Figure 1).
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
36 | rates |