Data tracks section of the Select tracks dialog enables you to specify the tracks for visualization on the canvas. An overview of the available track types is provided in Figure 1. Note that not all tracks are visible at all times and that their presence depends whether specific data types are present in the project as well as the zoom level. The tracks can be customized and their appearance changed by using the control panel on the left. Different track also has it own specific configuration settings which allow you to pin the track to move the track, pin the track, change the styles of the track and hide the track using the settings options on the left of each track ( )
Alignments track
Isoform proportion track
Variants track
Amino acids track
Reads pileup track
Probe intensities track
Peaks track
Alignments Track
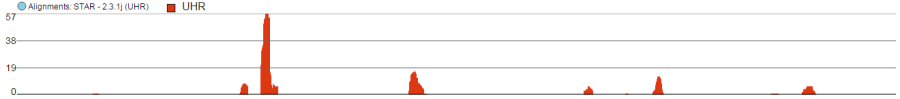
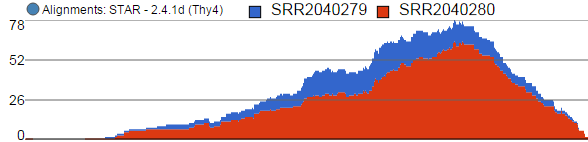
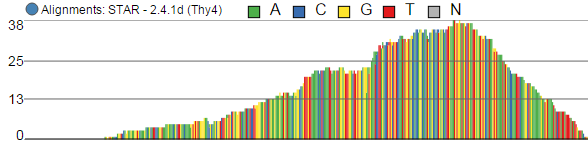
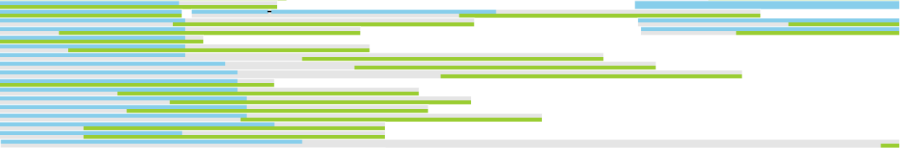
Alignments track displays a histogram view of alignments present in .bam files in a stacked histogram fashion (similar to Partek® Genomics Suite®). The y-axis shows number of (raw) base calls per position. By default, reads are coloured by sample. Variations on the track are displayed below. They can be configured in the following ways:
- The histogram can be colored by sample, attribute groups or by base call. By default, it is colored by sample except when the chromosome viewer is invoked from a variant data table.
- When colouring reads by sample, the reads are stacked (on top of each other), i.e. in the example below there are more reads in the red sample than in the blue sample. This is an example of a Sum histogram type. This can also be configured to display overlays or averages.
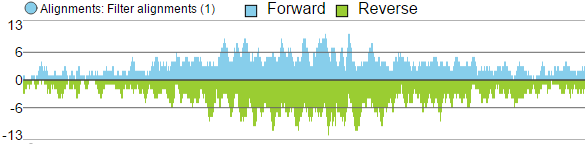
- Reads can be split into two tracks corresponding to the strand that they map to. This can be invoked by clicking the Split read histogram by strand checkbox.
- Y-axis can be scaled for all samples to have the same max or each sample has its own specific max
For more information on configuring tracks see our page on Customizing the chromosome viewer
Reads coloured by sample
Reads coloured by base calls
Reads split by strand
Isoform Proportion Track
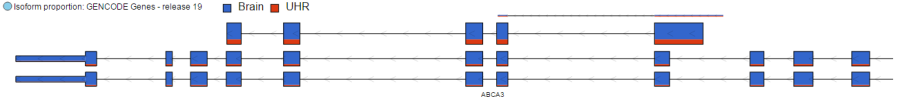
The Isoform proportion track displays the reads mapped to transcripts and helps to visualize differential expression and alternative splicing, this track is only available on the feature list data node which is generated from differential expression analysis. It uses standard symbols for exons (boxes) and introns (lines connecting the boxes). The height and color of each transcript is proprotional to its LS mean value. Figure 3 shows a gene with two transcripts in RefSeq database; the top transcript is more abundant than the bottom transcript and is preferentially expressed in the "blue" condition (labeled as 0 uM). The bottom transcript, on the other hand, seems to be expressed at the same level across all three conditions (i.e. 0 uM, 5 uM, 10 uM). The number and structure of transcripts on the plot depend on the transcript model that was used for mapping.
For more information on configuring tracks see our page on Customizing the chromosome viewer
Variants Track
Variant tracks show single nucleotide variants (SNVs) and indels, and appear in the Select track dialog if Detect variants task has been performed. Presentation of variants depends on the level of zoom. With low power magnification, SNVs are seen as purple columns and indels are bars (insertions: green bars; deletions: red bars) (Figure 4).
Upon zoom-in, SNVs are drawn as pie charts, representing the proportion of each base call at that locus (Figure 5)
At higher modification, insertions are seen as green boxes, with individual inserted bases presented using a pie chart, while deletions look like red boxes and the affected bases are also presented by a pie (Figure 6).
Insertion
Deletion
Amino Acids Track
Amino acids track becomes available in the Select tracks dialog after completing the Annotate variants task. The actual appearance of the track depends on the zoom level. With low-power magnification, you will see a message View not available at this zoom level, Please zoom in to view amino acids.
When you zoom closer to the genome, all the amino acids become visible as colored boxes (Figure 7) and labeled using the single-letter amino acid code. Alternative amino acids are depicted as additional box on the top of the consensus sequence.
If an amino acid spans two exons, its box will be truncated and the line connecting the exons will be dashed. An example is in Figure 8.
An empty gray box on the top of consensus sequence is used to indicate a STOP codon, which is a consequence of a mutation (Figure 9).
Untranslated bases, such as ones downstream of a STOP codon are depicted by lighter shades. Figure 10 shows two transcripts in an amino acid track; the direction is from left to right, so amino acids downstream of a STOP codon (P > G > L) are lightly shaded.
Reads Pileup Track
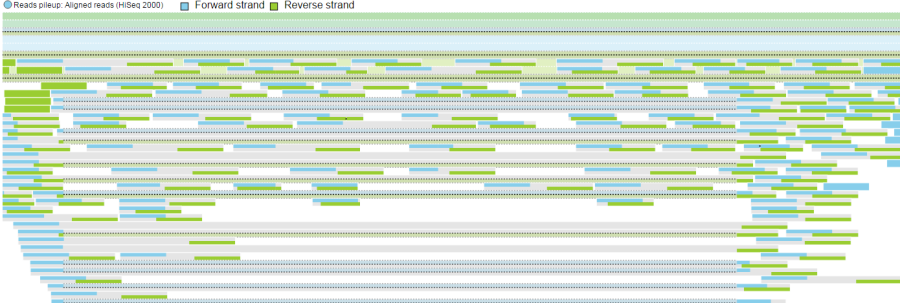
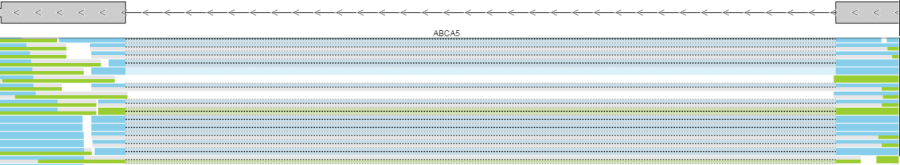
Reads pileup track plots the short sequencing reads, as present in the .bam file. The track is not on by default (go to Select tracks to turn it on) and its appearance depends on the magnification; if you are zoomed out a message - Zoom in to view individual reads - will be displayed.
Forward strand reads are in sky blue, while reverse strand reads are in parakeet green. If paired-end chemistry was used, the paired reads will be depicted as half reads within a gray rectangle encompassing the pair (Figure 11). Singletons will be depicted as thicker reads.
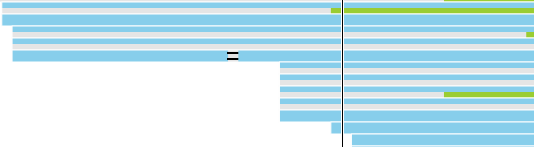
If you used a junction-aware aligner (such as TopHat or STAR), the junction reads will be depicted using dashed lines, which connect exon-spanning parts of the same read (Figure 12).
Deleted bases can also be seen on a Reads pileup track, as fat black lines (Figure 13).
Probe Intensities Track
Microarray probes are visualised by the Probe intensities track. The probes are shown as bars and their colour depends on the probe intensity, ranging from white (low) to admiral blue (high) (Figure 14).
As with the Reads pileup track, probes may not be visible with low power magnification and you will see a message - Zoom in to view individual microarray probes.
Peaks Track
The Peaks track displays the results of peak caller tasks. It displays a bar that spans genomic location of each peak call. If summits are identified by the peak caller, such as the MACS2 algorithm, then its genomic location is marked by a vertical line. The color marks either the pair being compared by the peak caller or, if present, the sample attribute associated with the sample.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
34 | rates |





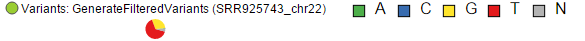

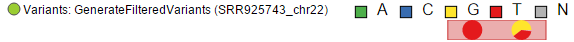





1 Comment
Melissa del Rosario
author: ilukic