Create a new Project
Let's start by creating a new project.
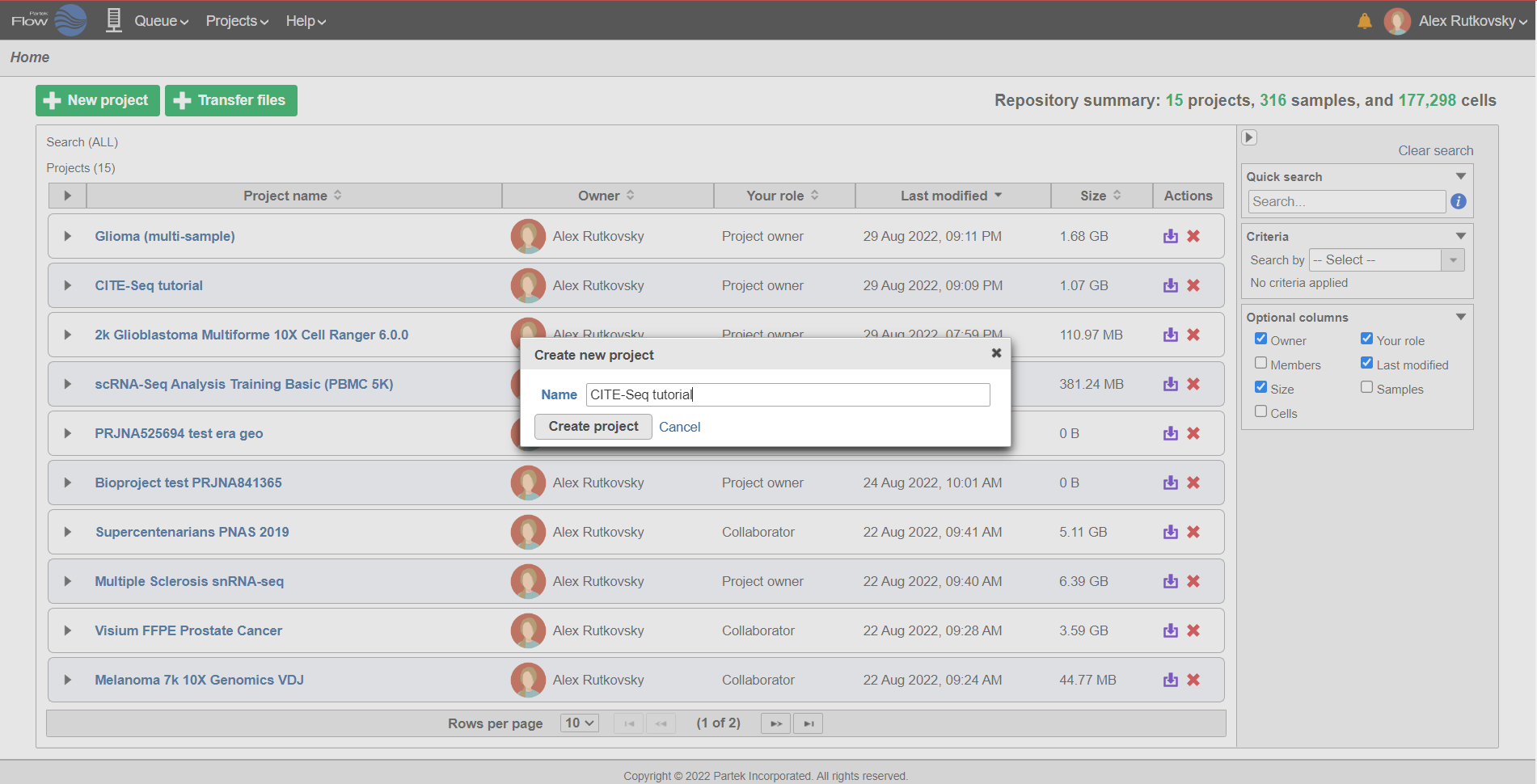
- On the Home page, click New project (Figure 1)
- Give the project a name
- Click Create project
Import data
- Under the Data tab, click Import data
- Click Import single cell data
- Choose the filtered HDF5 file for the MALT sample produced by Cell Ranger
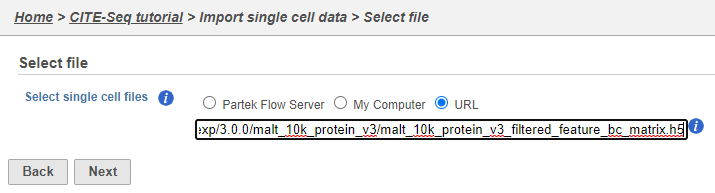
Move the .h5 file to where Partek Flow is installed using , then choose the Partek Flow Server radio button and browse to its location. Otherwise, select the URL radio button and paste in the following link (Figure 2):
http://cf.10xgenomics.com/samples/cell-exp/3.0.0/malt_10k_protein_v3/malt_10k_protein_v3_filtered_feature_bc_matrix.h5
- Click Next
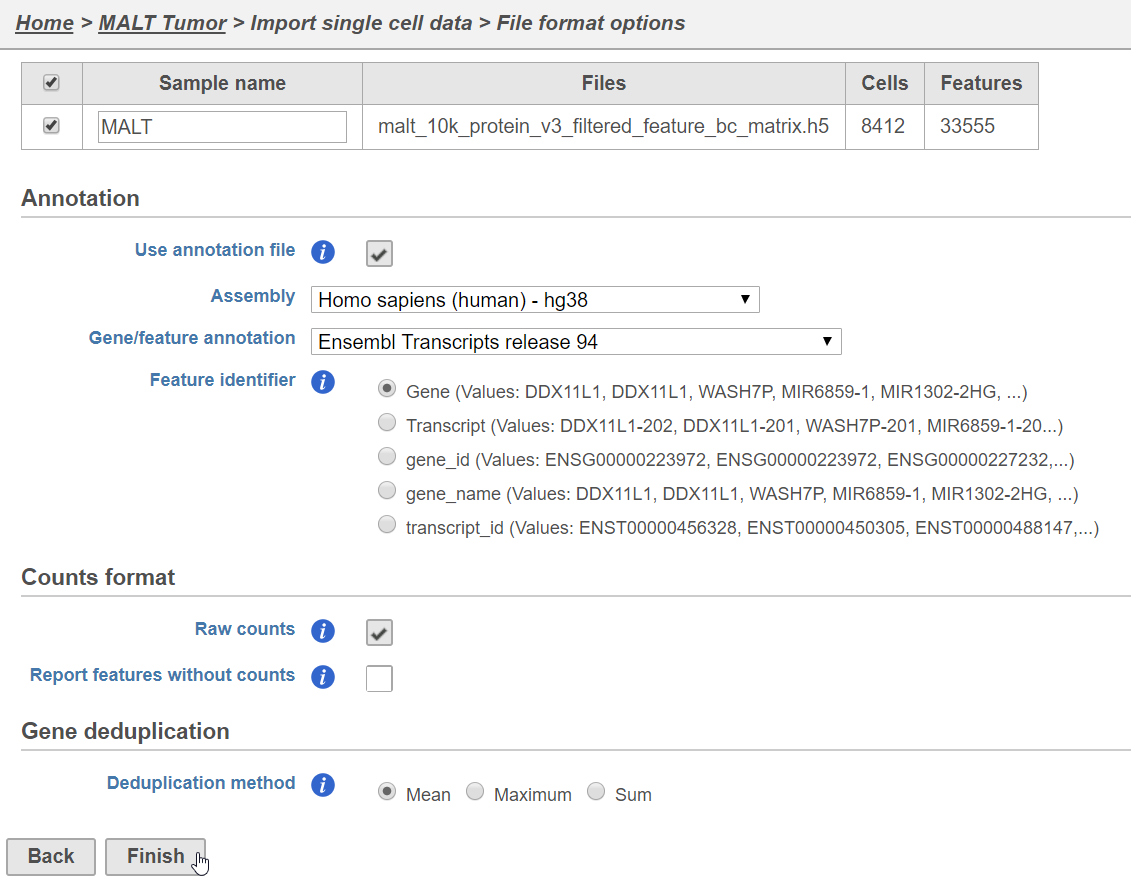
- Name the sample MALT (the default is the file name)
- Specify the annotation used for the gene expression data (here, we choose Homo sapiens (human) - hg38 and Ensembl Transcripts release 94). If Ensembl 94 is not available from the drop-down list, choose Add annotation and download it.
- Uncheck Report features without counts
- Click Finish (Figure 3)
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |