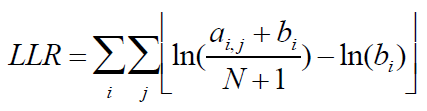Motif detection tasks allow to extract binding motifs from peak region generated from ChIP-seq data. It allows users to search for know motifs from provided database as well as detect noval motifs. These tasks are available when click on region data node
Detect de novo motifs
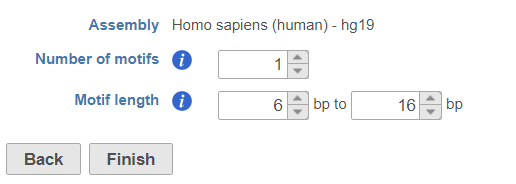
Click on Peak data node, select Detect de novo motifs from Motif detection section in the pop-up menu (Figure 1), specify the number of motifs to detect and the length of the motifs.
This is done by repeating the below two steps until conveergences:
A. Given the alignment matrix from step B, search for location in the set of regions that score highly compared to the alignment matrix using equation
Search for know motifs
Reference
- Neuwald, A.F., Liu, J.S., & Lawrence, C.E. Gibbs motif sampling: detection of bacterial outer membrane protein repeats. Protein Science 1995, 4: 1618-1632.


| Your Rating: |
    
|
Results: |
    
|
1 | rates |