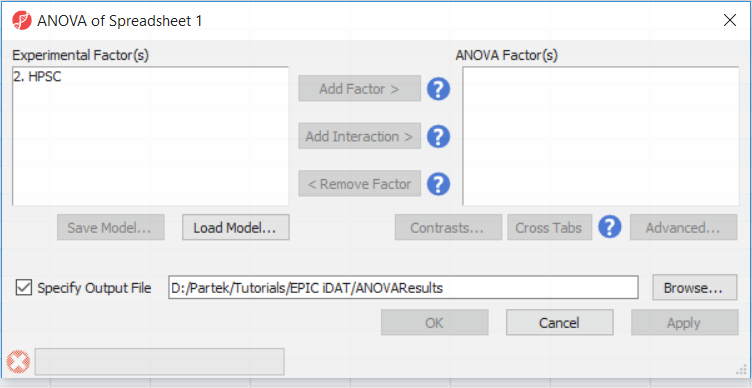
To detect differential methylation between CpG loci in different HSCP populations, go to Analysis > Detect Differential Methylation. In the ANOVA dialog (Figure 1), select Add Factor to move the factor 2. HPSC from Experimental Factor(s) to the ANOVA Factor(s) box.
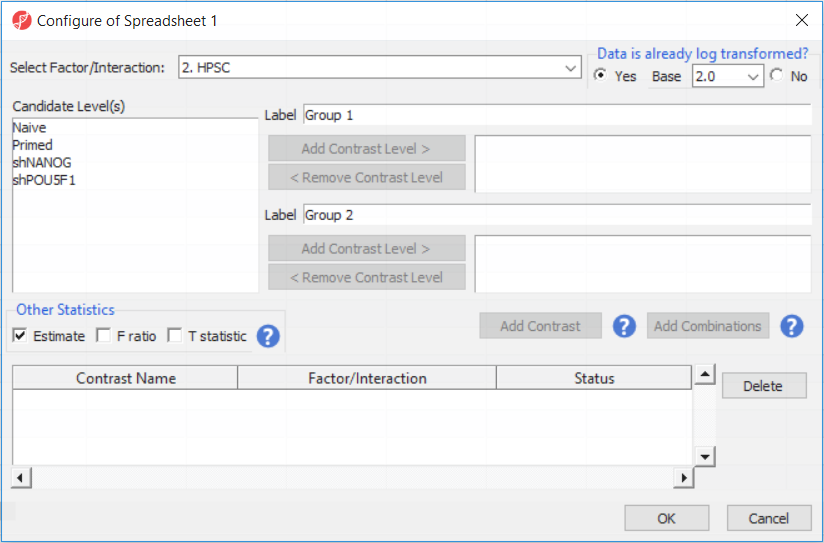
Depending on the data in the top level (i.e. parent) spreadsheet you may want to manually specify whether the data is already log transformed: you should select Yes for M-values (but otherwise No for β-values) (Figure 2). By default, Partek Genomics Suite will calculate fold-change value for each contrast. In addition, to include the difference in methylation levels at each CpG site in the output (i.e. ΔM or Δβ), check the Estimate box in the Other Statistics section of the dialog (Figure 2).
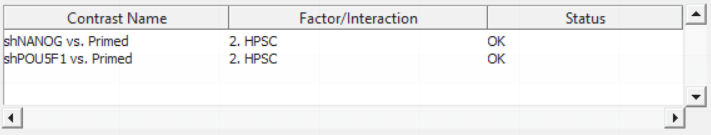
For this exercise, we shall compare primed HPSC with suppression of OCT4 (shPOU5F1) and primed HPSC with suppression of NANOG (shNANOG) with the baseline primed cells. To start with, select the Primed group in the Candidate Level(s) box and push Add Contrast Level > to move the Primed group to Group 2 (lower box). Then Ctrl & select both shPOU5F1 and shNANOG in the Candidate Level(s) box and push Add Contrast Level > to move them to Group 1 (upper box). Then click Add Combinations and confirm that two contrast have been created as seen in Figure 3.
Push OK to confirm the contrast (and close the contrast dialog) and again to start the ANOVA calculation.
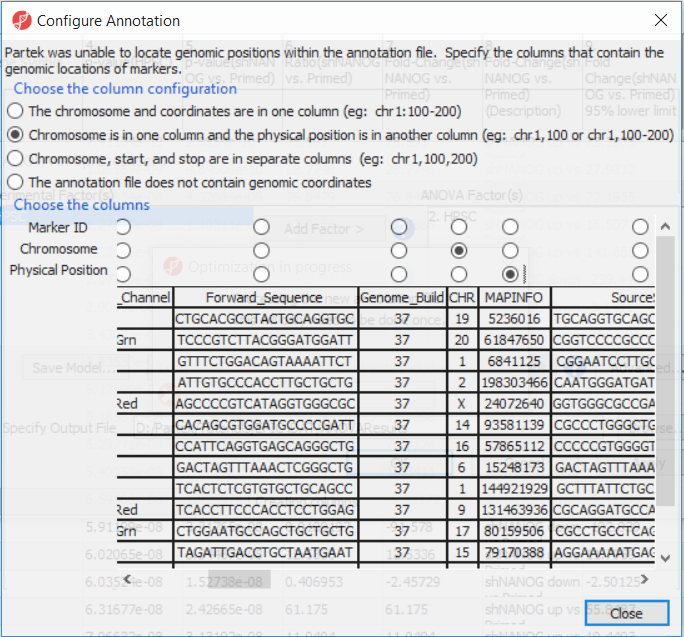
The first time you use MethylationEPIC array, the manifest file needs to be configured and the window like the one in Figure 4 will pop up. First select the second option (Chromosome is in one column and the physical location is in another column). Marker ID is on the first column (Ilmn ID), Chromosome is on the column CHR, while the Physical Position is on the MAPINFO column. Set according to Figure 4 and select Close. This enable Partek Genomics Suite to parse out probe annotation from the manifest file.
Section Heading
Section headings should use level 2 heading, while the content of the section should use paragraph (which is the default). You can choose the style in the first dropdown in toolbar.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |