Page History
...
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5600148/
Running
...
Salmon
Salmon will run on any FASTQ file input.
- Click the data node containing your FASTQ files
- Click the Quantification section in the toolbox
- Click Count feature barcodes
- Choose the Prep kit
For more details on adding Prep kits, please see our documentation on Trim tags. The prep kit should include cell barcode and unique molecular identifiers (UMIs) locations.
- Choose the Barcode location
For BioLegend TotalSeq-A, choose bases 1-15. For BioLegend TotalSeq-B/C, choose bases 11-25. For other locations, select Custom and specify the start and stop positions.
- Choose a Sequences text file
This tab-delimited text file should have the feature ID in the first column and the nucleotide sequence in the second column. Do not include column headers.
- Click Finish to run
The output of Count feature barcodes is a Single cell counts data node.
How does Count feature barcodes work?
Count feature barcodes uses a series of tasks available independently in Partek Flow to process the input FASTQ files. The output files generated by these tasks are not retained in the Count feature barcodes output.
Trim tags identifies the UMI and cell barcode sequences. The Prep kit is specified using the Prep kit setting.
Trim bases trims the insert read to include only the feature barcode sequence. Trim bases is set to Both ends for Trim based on with the start and stop set by the Barcode location preference and the Min read length set to 1.
Bowtie is used to align the reads to the sequences specified in the Sequences text file. Bowtie is set to Ignore quality limit for the Alignment mode. Other settings are default.
Deduplicate UMIs consolidates duplicate reads based on UMIs. Deduplicate UMIs is set to Retain only one alignment per UMI.
Filter barcodes filters the cell barcodes to include cells and not empty droplets. Filter barcodes is set to Automatic.
Quantify barcodes counts the number of UMIs per cell for each feature in the Sequences file. Quantify barcodes uses default settings.
To perform these steps individually instead of using the Count feature barcodes task, you will need to generate a FASTA and GTF file containing the feature barcode IDs and sequences instead of a text file and build an index file for the Bowtie aligner.
For more information about library file management, please see Library File Management.
- Salmon
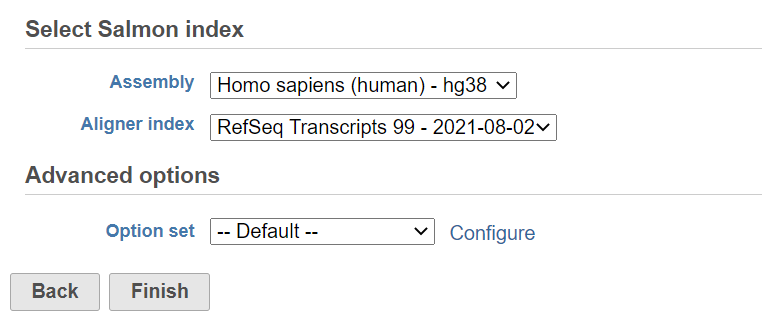
- Choose Assembly and Aligner index on gene annotation model (Figure 1)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Salmon index can't be built on reference assembly, you need to provide transcript annotation file beforehand, the index will be built on transcript annotation model. For more information about adding aligner index based on annotation model, see this chapter in library file management.
The output data node contains relative abundance of the transcripts in units of transcripts per million (TPM).
Note: If you want to perform differential analysis, you need to add offset to deal with 0 values.
References
Rob Patro, Geet Duggal, Michael Love, Rafeal A Irizarry, Carl Kingsford. Salmon: fast and bias-aware quantification of transcript expression using dual-phase inference. Published online 2017 Mar 6. doi:10.1038/nmeth.4197
...