Page History
| Table of Contents | ||||||
|---|---|---|---|---|---|---|
|
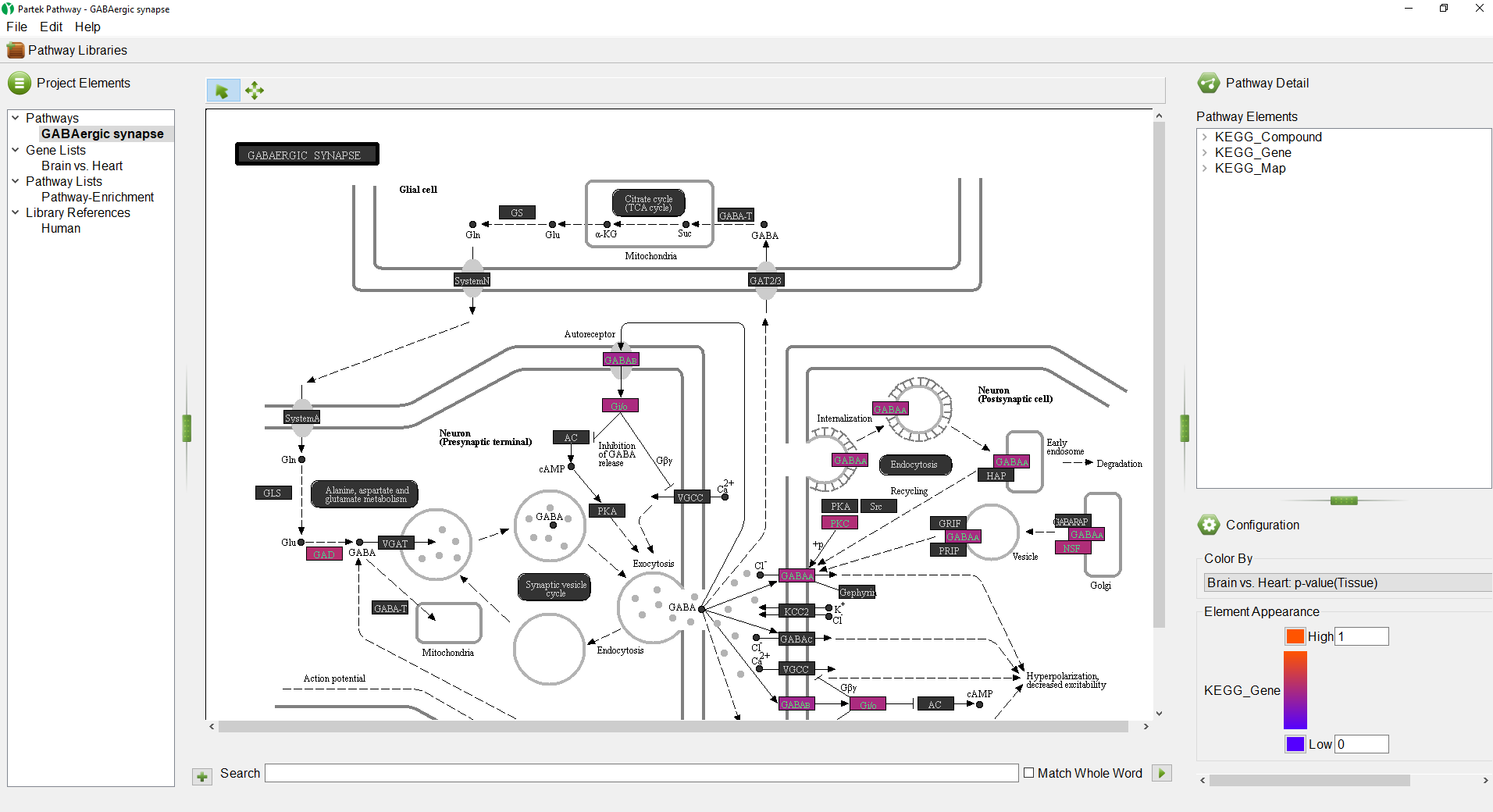
Partek Pathway is a standalone separate program from Partek Genomics Suite with a distinct user interface (Figure 1). The size of each panel can be adjusted using the green vertical and horizontal grab bars.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
The Project Elements panel (Figure 2) displays the selected pathway, the original gene list, the Pathway Enrichment spreadsheet, and the library references that were used for the pathway analysis. The Project Elements panel is used to navigate between open pathway diagrams and spreadsheets.
...
The Pathway-Enrichment.txt spreadsheet also seen in Partek Genomics Suite will open (Figure 4). Selecting This spreadsheet has the same contents as the Pathway-Enrichment.txt spreadsheet in Partek Genomics Suite. Selecting any of the pathway names will open its pathway diagram. The spreadsheet can be sorted by any column.
...
The GABAergic synapse pathway diagram will open. Genes in the pathway are shown as boxes. The color of the box boxes is set by the Configuration panel (Figure 5).
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
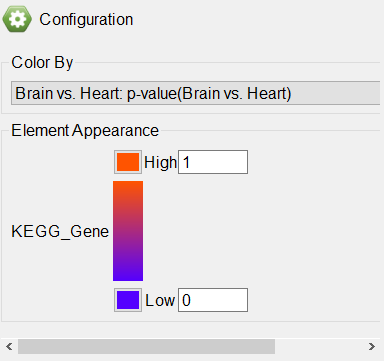
Any numerical column from the source gene list can be used to color the gene boxes. While significant p-values indicate a difference between the categories, they give no information about upregulation or downregulation of the pathway. We can overlay fold-change information on the pathway diagram.
- Select Brain vs. Heart: Fold-Change(Brain vs. Heart) from the drop-down menu
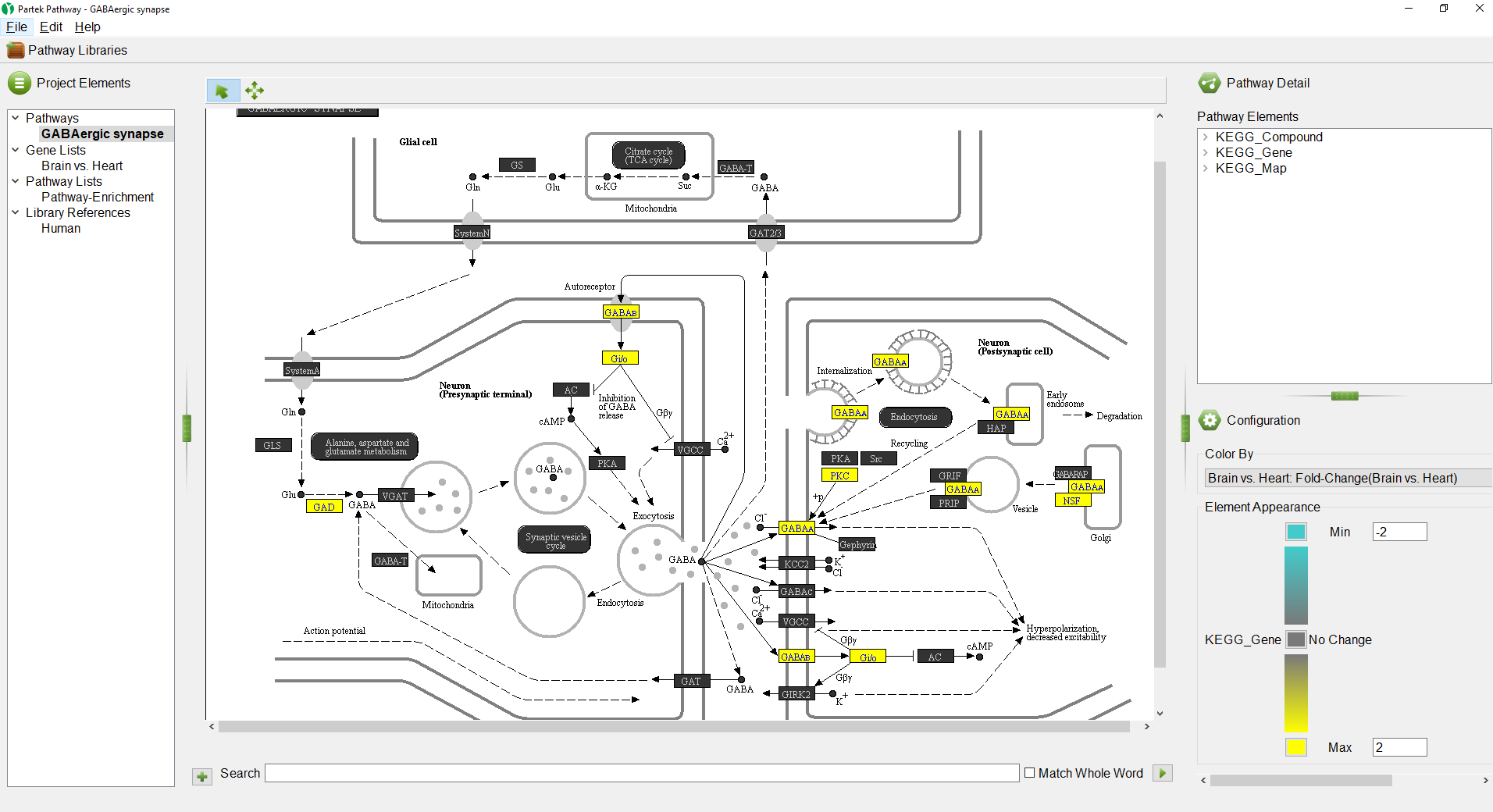
The pathway diagram now shows fold change for each gene in the pathway included in the gene list (Figure 6). Genes not in the gene list remain black.
...
- Select the red color square next to Max
- Select a yellow from the color picker interface
- Select OK
- Select the green color squre square next to Min
- Select teal pale blue from the color picker interface
- Select OK
We can see that all the colored genes in the GABAergic synapse pathway are yellow (Figure 7), indicating that they are upregulatedup-regulated.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
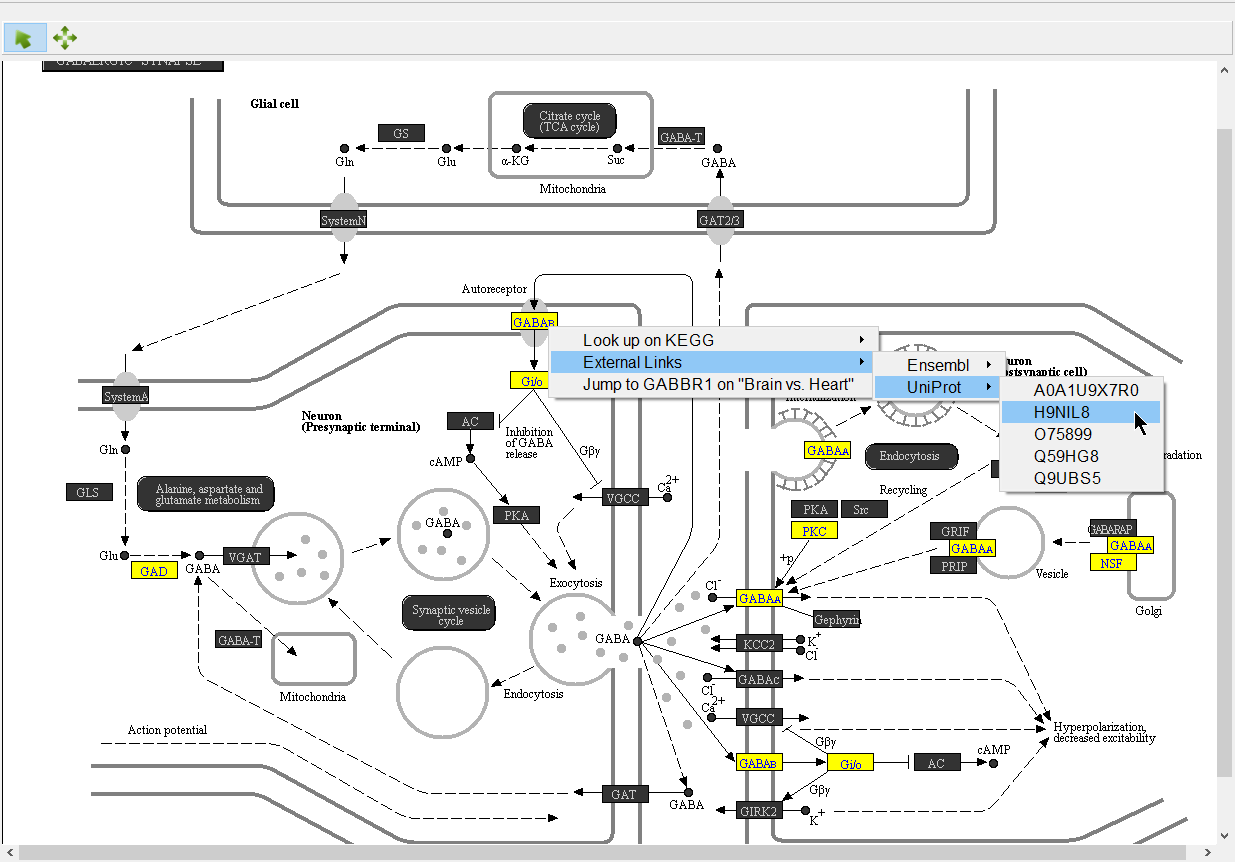
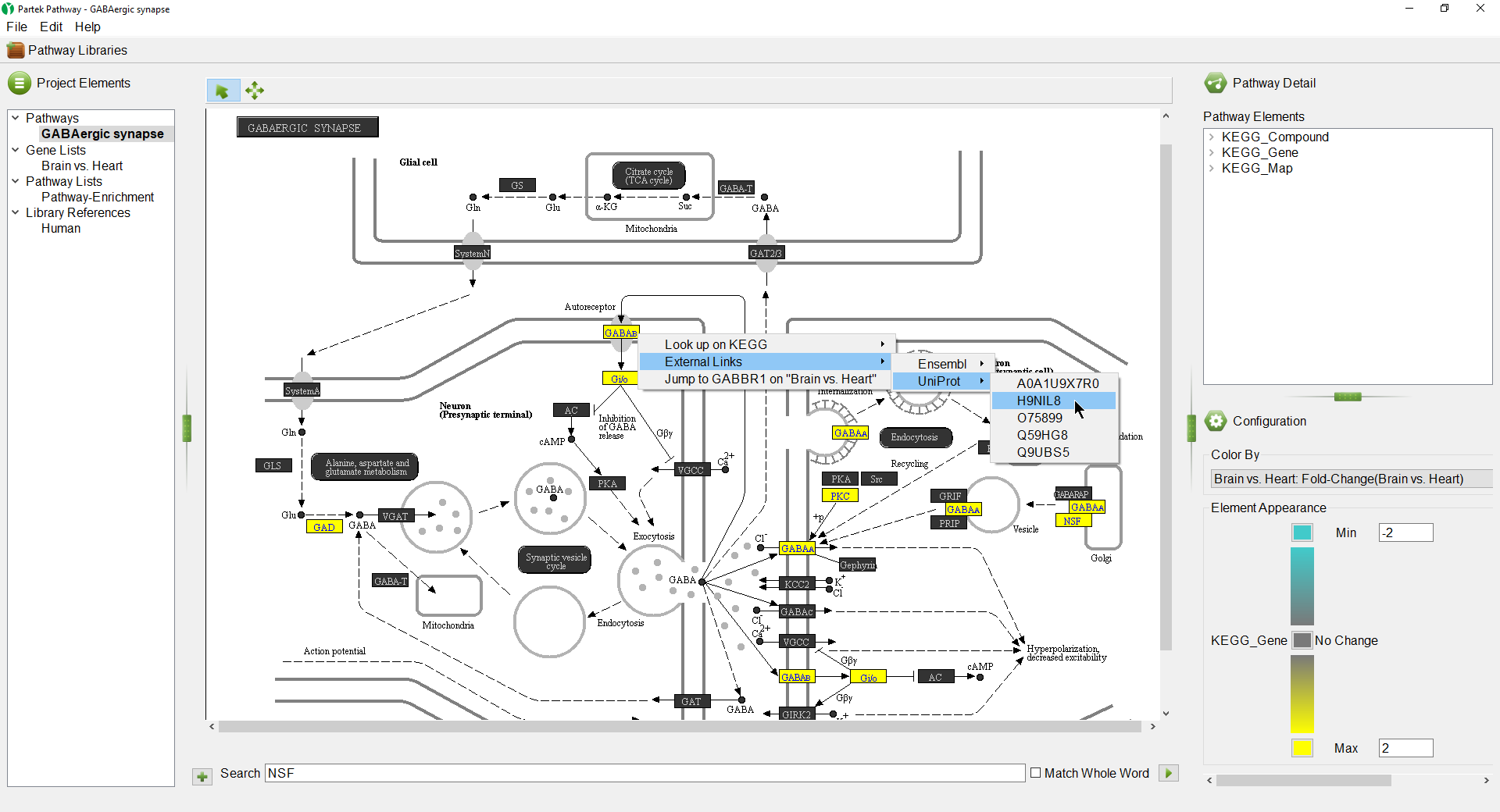
We can select a gene to learn more about it.
- Select () to activate selection modeSelection Mode
- Right-click GABAB (Figure 8)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Opions Options available include:
Look up on KEGG - opens the KEGG page for the pathway on GenomeNet (genome.jp) in your web browser
...
Jump to ___ on "___" - opens the source gene list in Partek Pathway to the row of the selected gene
Selecting () activates Navigation Mode. This enables navigation on large pathway diagrams by left-clicking and dragging to move the view.
The pathway database can be searched for genes of interest using the Search panel.
- Select () to open the search panel
- Type NSF in the search box
- Select () to search
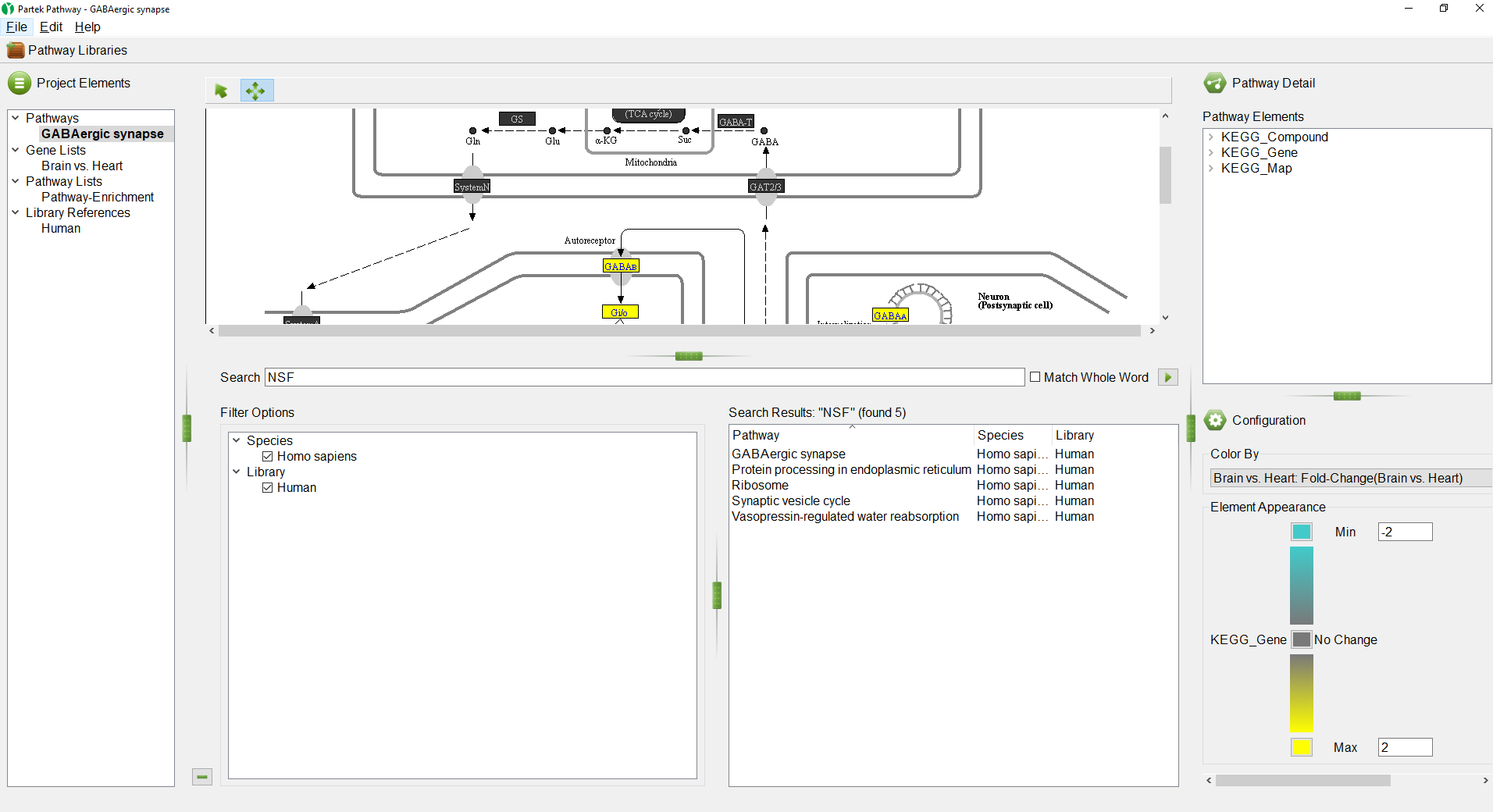
Pathways containing NSF appear on the right-hand side in the Search Results section in alphabetical order (Figure 9).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
If multiple species or libraries have been loaded, the Filter Options section on the left-hand side can be used to choose which species and libraries to search.
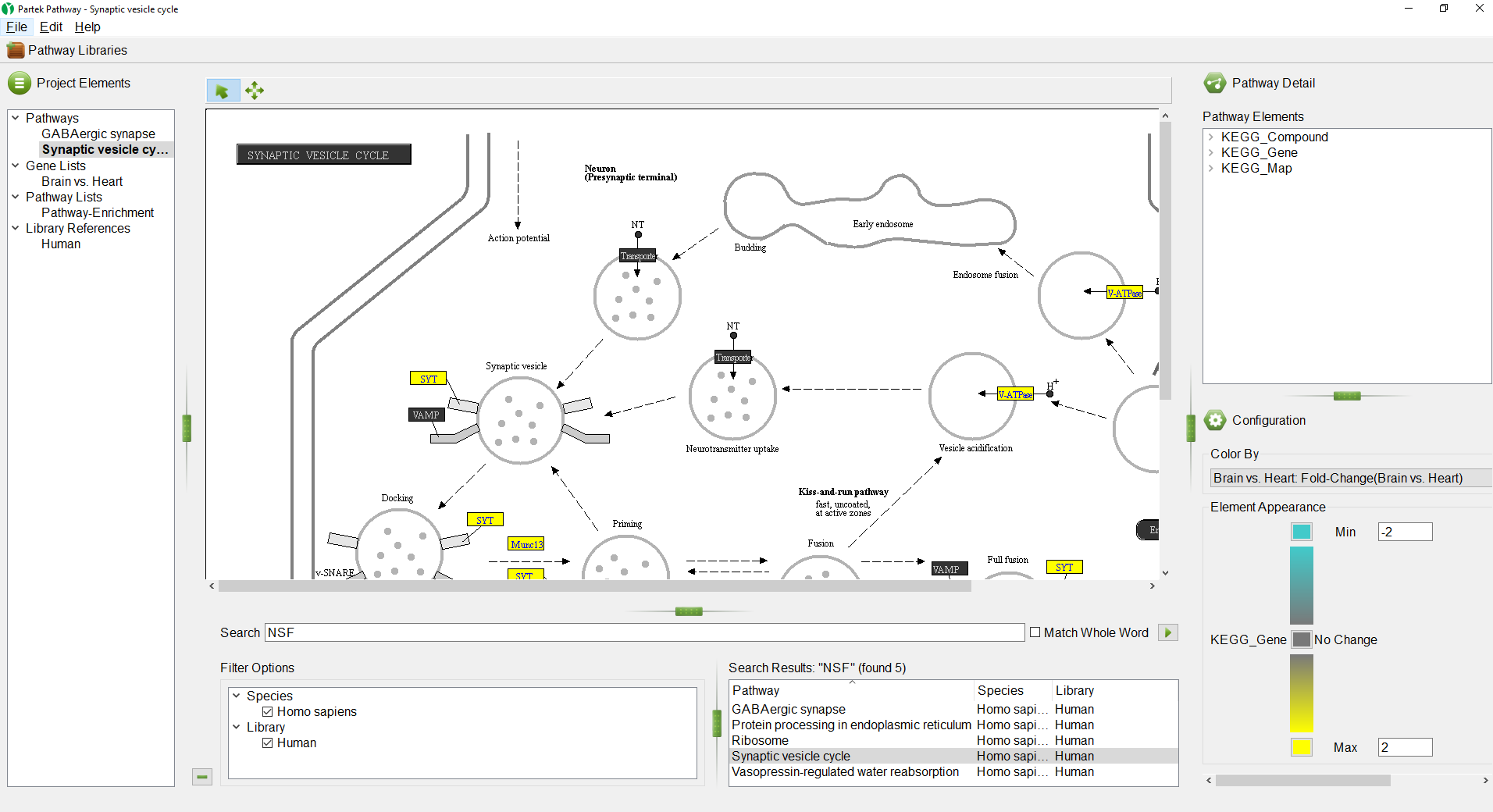
- Double click on Synaptic vesicle cycle in the Search Results section
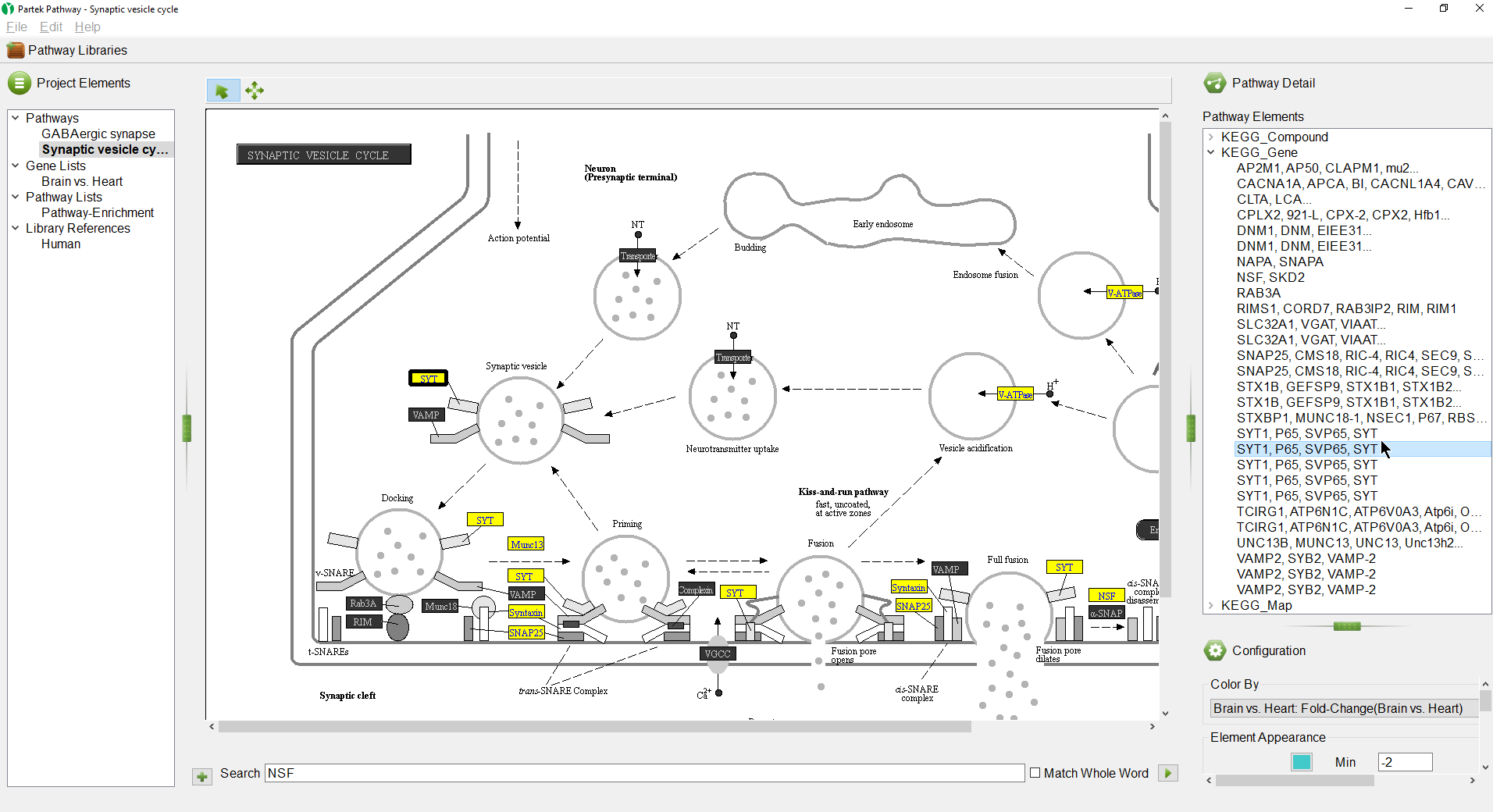
The selected pathway, Synaptic vesicle cycle, will open in the Pathway Diagram panel (Figure 10).
- Select () to minimize the Search panel
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
On the right-hand side of the Partek Pathway window, we see the Pathway Detail panel (Figure 11).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select KEGG_Gene to open the list of genes in the pathway
Selecting a gene in the list will highlight it in the pathway diagram (Figure 12).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Another way to select and view a pathway is browsing the Pathway Libraries.
- Select () in the upper left-hand corner of the Partek Pathway window
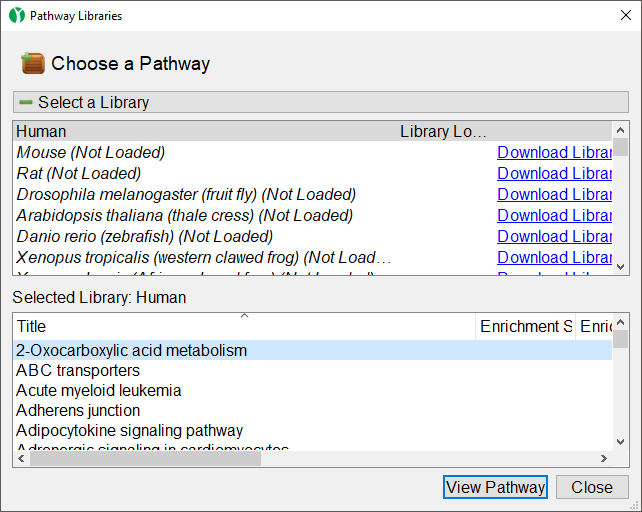
The Pathway Libraries dialog will open (Figure 13).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
In the upper section of the dialog, you can view available KEGG libraries and download them by selecting the Download Library link. Selecting a pathway opens it in the lower section of the dialog.
In the lower section of the dialog, you can view a list of all the pathways in the selected pathway library. You can also open any pathway from the selected library in the Pathway Diagram panel.
- Select Adherens Junction
- Select View Pathway to open the pathway diagram
We can use the Project Elements panel to close an open pathway diagram or list.
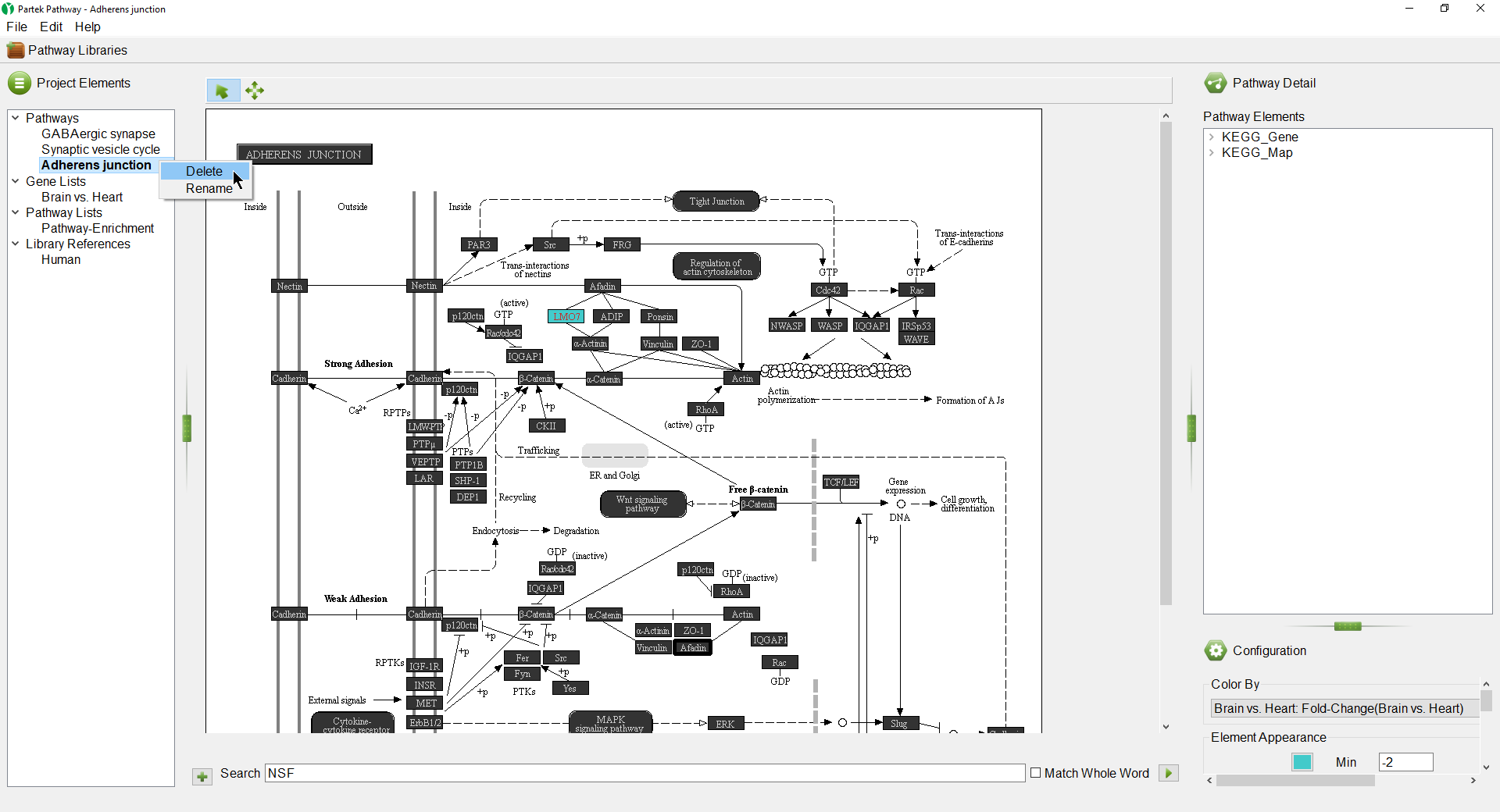
- Right-click Adherens Junction in the Project Elements panel
- Select Delete from the pop-up menu to close the diagram (Figure 14)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
The Search panel and Pathway Libraries can also be used to open pathway diagrams for pathways without any open gene or pathway lists.
| Additional assistance |
|---|
| Rate Macro | ||
|---|---|---|
|