| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| minLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
Import single cell data for different assay types and formats
Select Single cell, choose the assay type (scRNA-Seq, Spatial transcriptomics, scATAC-Seq, V(D)J, Flow/Mass cytometry), and select the data format (Figure 1). Use the Next button to proceed with import.
| Numbered figure captions |
|---|
| SubtitleText | Choose import single cell data option |
|---|
| AnchorName | Import_sc_count |
|---|
|
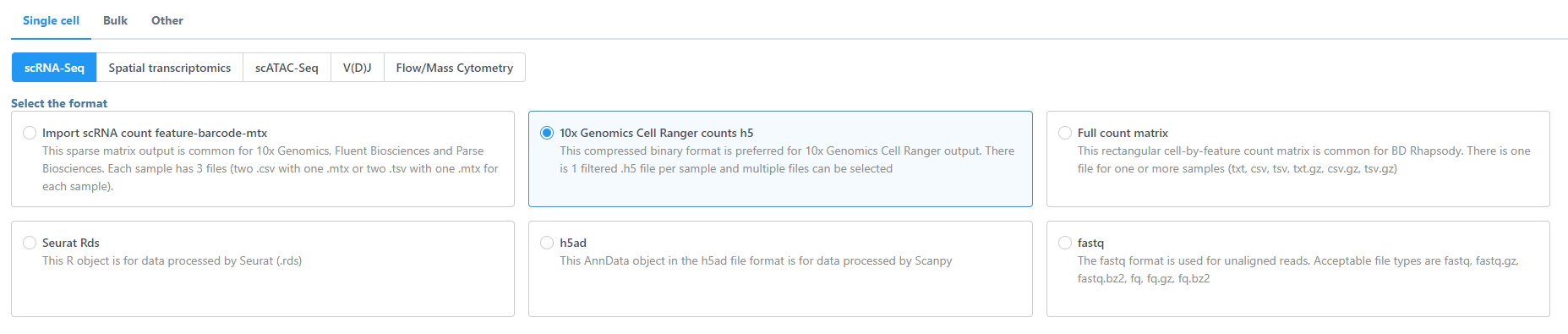 Image Added Image Added
|
Import single cell data from count matrix text file(s)
...
using Full count matrix as the data format
Partek Flow supports single cell data analysis in count matrix text format using the Full count matrix data format (Figure 2). Each matrix text file is assumed to represent on sample, each value in the matrix represents expression value of a feature (e.g. a gene, or a transcript) in a cell. The expression value can be raw count, or normalized count. The requirment of requirement of the format of each text file should be the same as count matrix data.
To import a sample during import data, choose Import single cell data option (Figure 1)
| Numbered figure captions |
|---|
| SubtitleText | Choose import single cell data option |
|---|
| AnchorName | Import_sc_count |
|---|
|
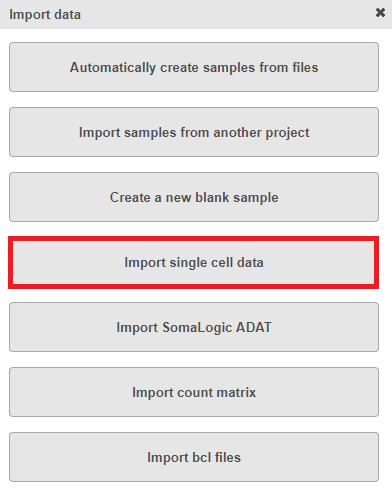 Image Removed Image Removed
|
Specify text file location, only one text file (in other words one sample) can be imported at once, preview of the file will be displayed, configuration of the file format is the same as Import count matrix data. In addition, you need to specify Sample information and Gene deduplication method (Figure 2)the details about this file.
| Numbered figure captions |
|---|
| SubtitleText | Specify sample name and gene deduplication methodChoose Full count matrix as the data format |
|---|
| AnchorName | sample_info |
|---|
| 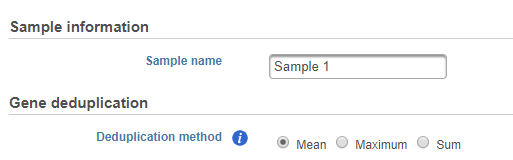 Image Removed
Image Removed |
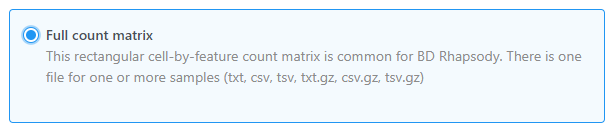 Image Added Image Added
|
Click Finish, the sample will be imported, on the data tab, number of cells in the sample will be displayed.
To import multiple samples, repeat the above steps by clicking Import data on the data tab Metadata tab or within the task menu (toolbox) on the Analyses tab. Make the same previous selections using the cascading menu.
...



