Page History
...
- Select Create Marker List from the Analysis section of the Illumina BeadArray Methylation workflow
- Select 1/anova-1way_autosomal_only (ANOVA-1way_autosomal_only) as the source spreadsheet from the left-hand panel
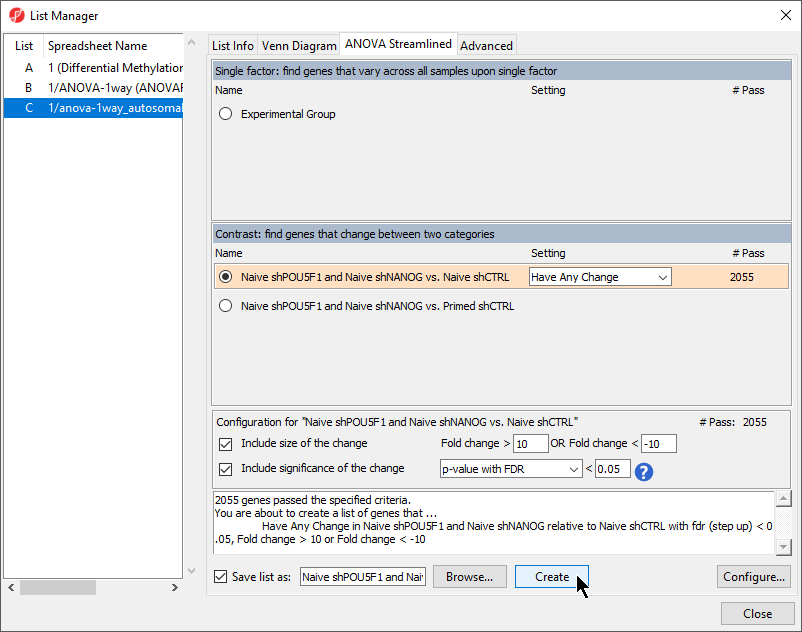
- Select Naive shPOU5F1 and Naive shNANOG vs. Naive shCTRL from the Contrast: find genes that change between two categories section of the ANOVA Streamlined tab of the List Manager dialog
- Set Fold change > to 10
- Set OR Fold change < to -10
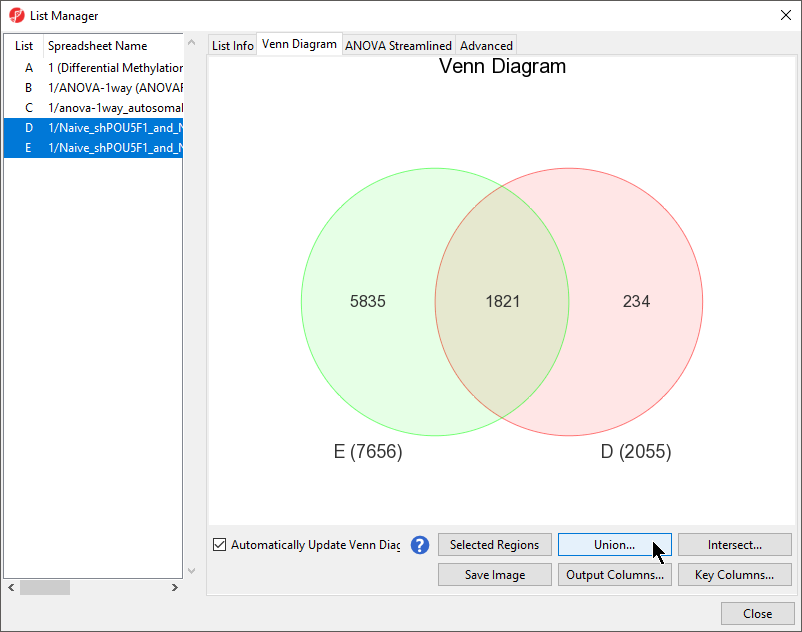
- Leave the rest of the option set to defaults Select LCLs vs. B cells (Figure 1)
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
|
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
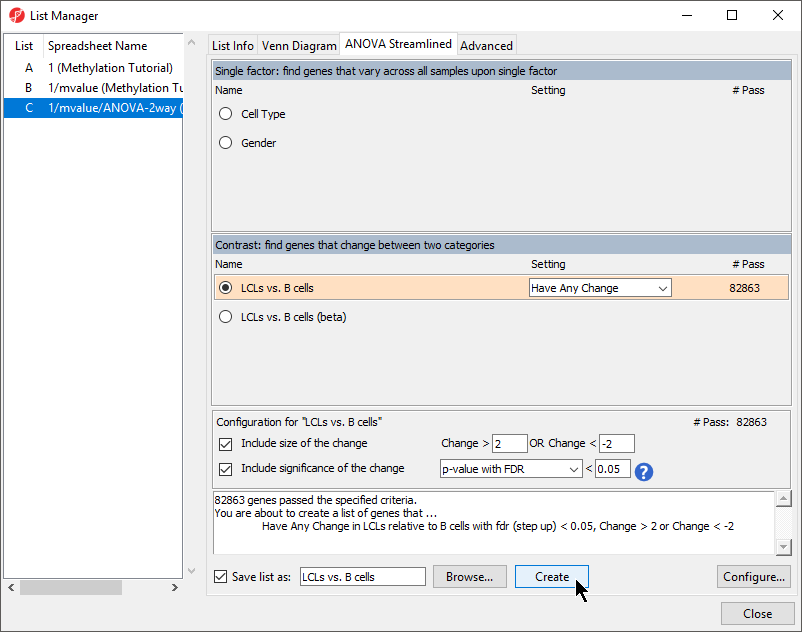
- Leave Include size of the change selected and set to Change > 2 OR Change < -2
- Leave Include significance of the change selected and set to p-value with FDR < 0.05
- Select Create
- Select Close to exit the list manager
The new spreadsheet LCLs vs. B cells (LCLs vs. B cells) will open in the Analysis tab.
It is best practice to occasionally save the project you are working on. Let's take the opportunity to do this now.
- Select File from the main command toolbar
- Select Save Project...
- Specify a name for the project, we chose Methylation Tutorial, using the Save File dialog
- Select Save to save the spreadsheet
...
- project
Saving the project before proceeding to the next section of the tutorial.saves the identity and child-parent relationships of all spreadsheets displayed in the spreadsheet tree. This allows us to open all relevant spreadsheets for our analysis by selecting the project file.
| Page Turner | ||
|---|---|---|
|
...
Overview
Content Tools