...
The input for Annotate peaks is a Peaks type data node.
- Click a Peaks the Filtered features data node
- Click the Peak analysis section in the toolbox
- Click Annotate regions
- Set the Genomic overlaps parameter
...
| Numbered figure captions |
|---|
| SubtitleText | Annotate regions in Partek Flow. |
|---|
| AnchorName | Annotate regions |
|---|
|
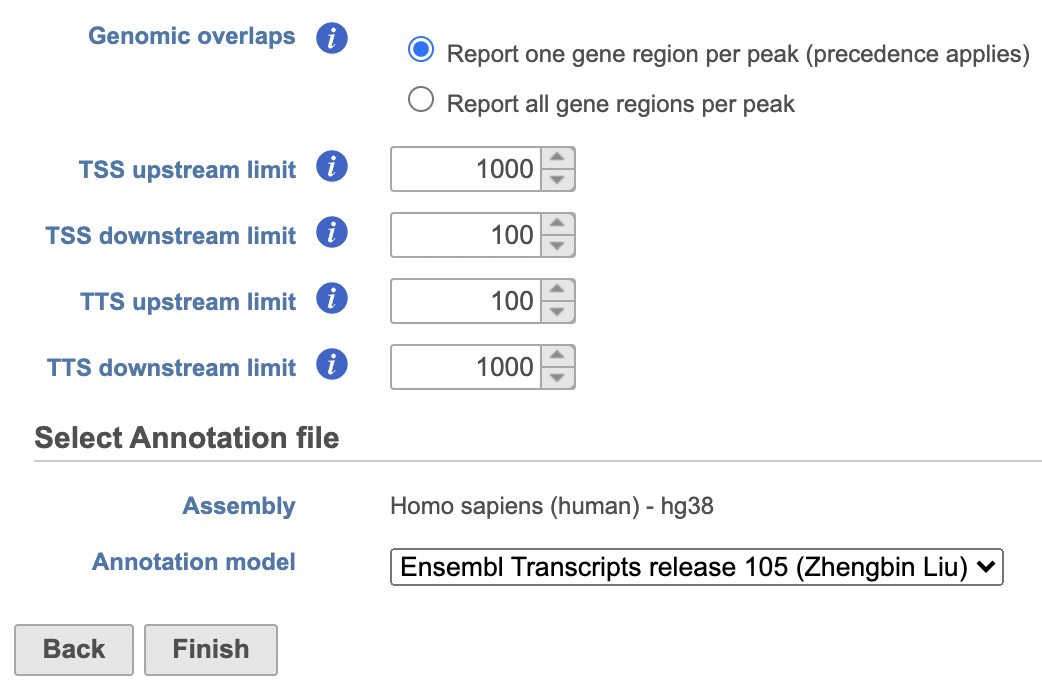 Image Removed Image Removed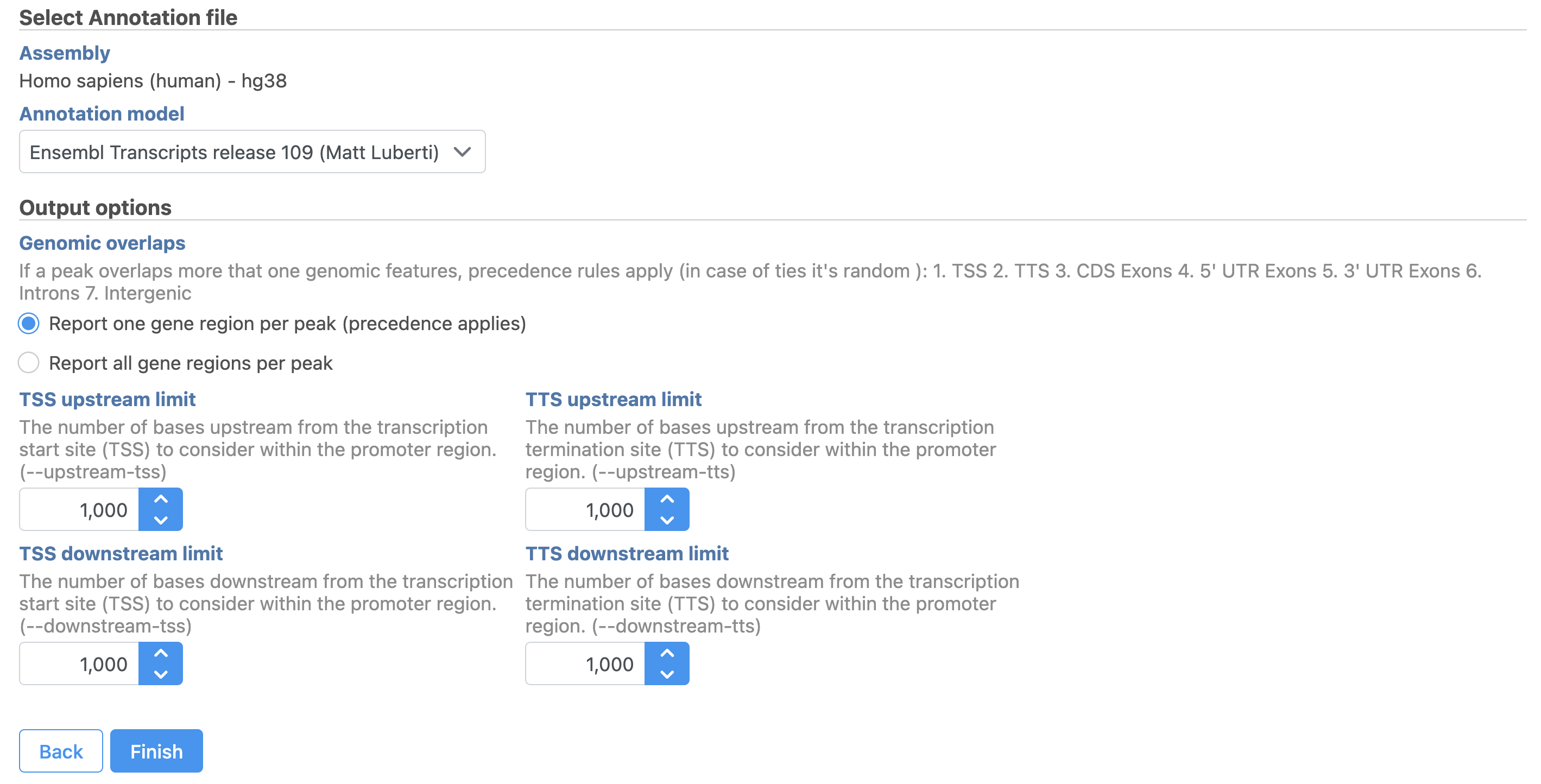 Image Added Image Added
|
Users are able to define the transcription start site (TSS) and transcription termination site (TTS) limit in the unit of bp.
...
| Numbered figure captions |
|---|
| SubtitleText | TF-IDF normalization for scATAC-Seq in Flow. |
|---|
| AnchorName | Normalization |
|---|
|
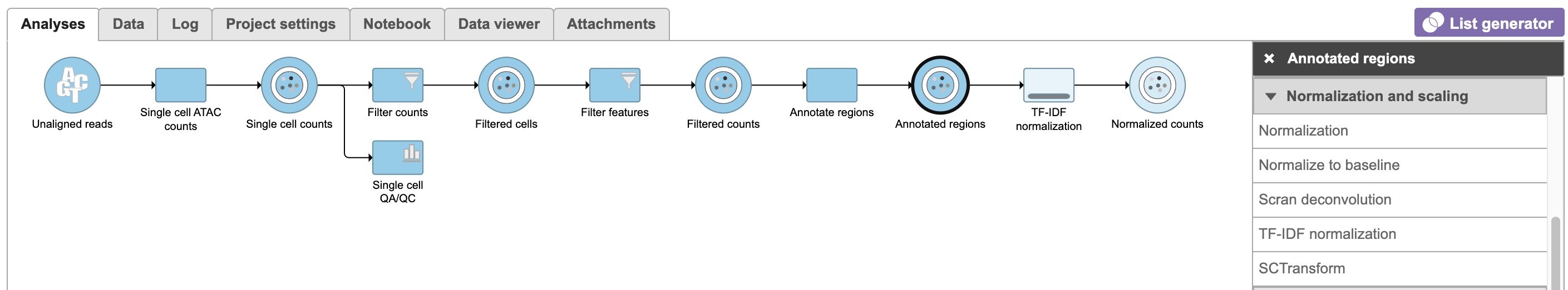 Image Removed Image Removed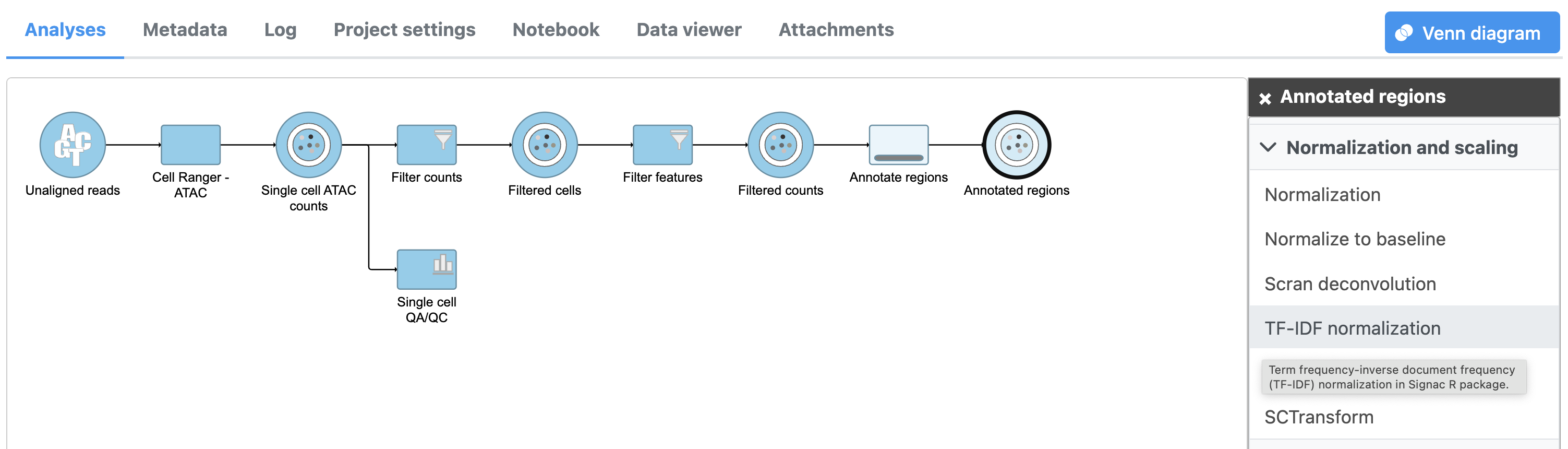 Image Added Image Added
|
To run TF-IDF normalization,
- Click a single Single cell counts data node, in this case the Annotated regions node
- Click the Normalization and scaling section in the toolbox
- Click TF-IDF normalization
...
| Numbered figure captions |
|---|
| SubtitleText | SVD task configuration dialog in Partek Flow. |
|---|
| AnchorName | SVD |
|---|
|
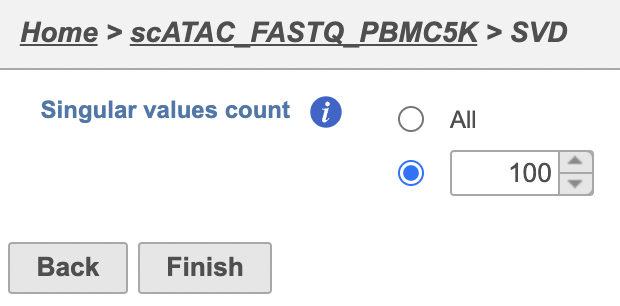 Image Removed Image Removed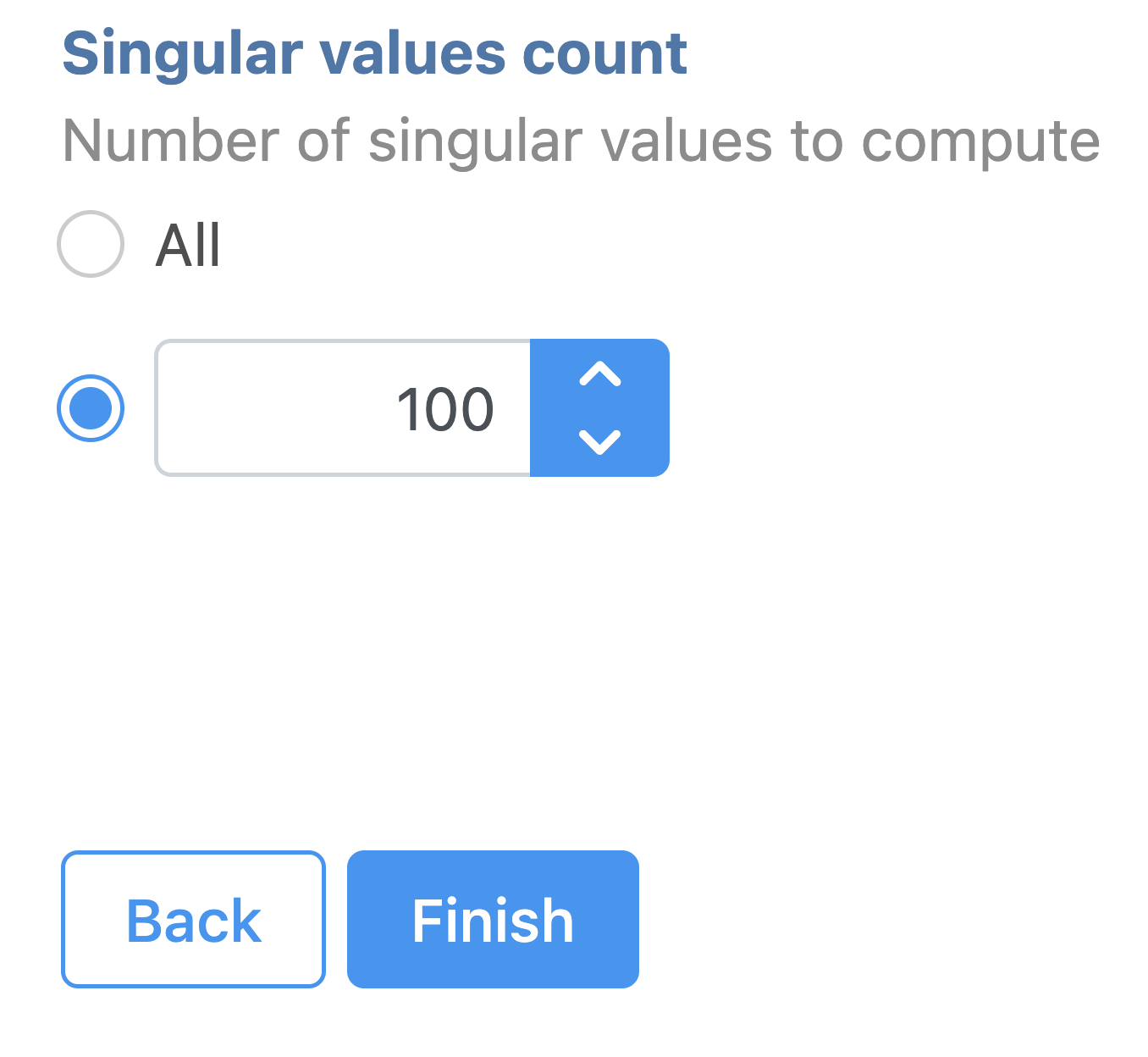 Image Added Image Added
|
Graph-based clustering
...
| Numbered figure captions |
|---|
| SubtitleText | Configure Graph-based clustering in Flow. |
|---|
| AnchorName | Configure Graph-based clustering |
|---|
|
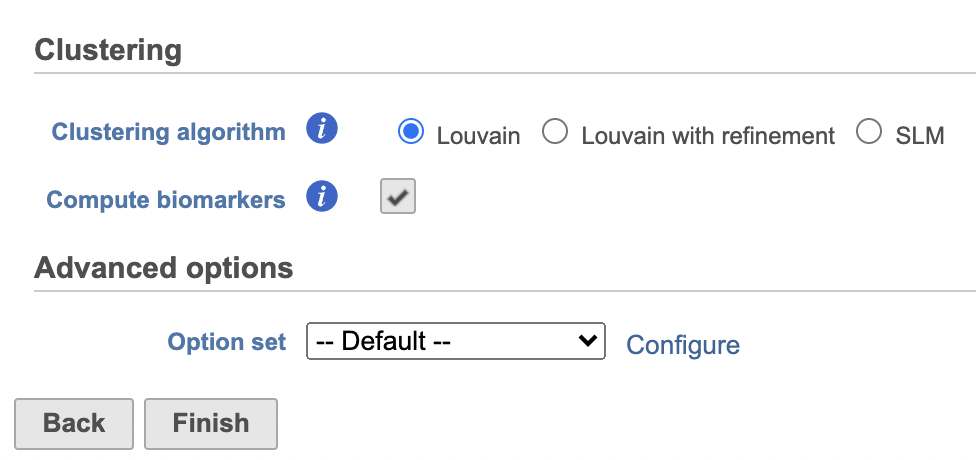 Image Removed Image Removed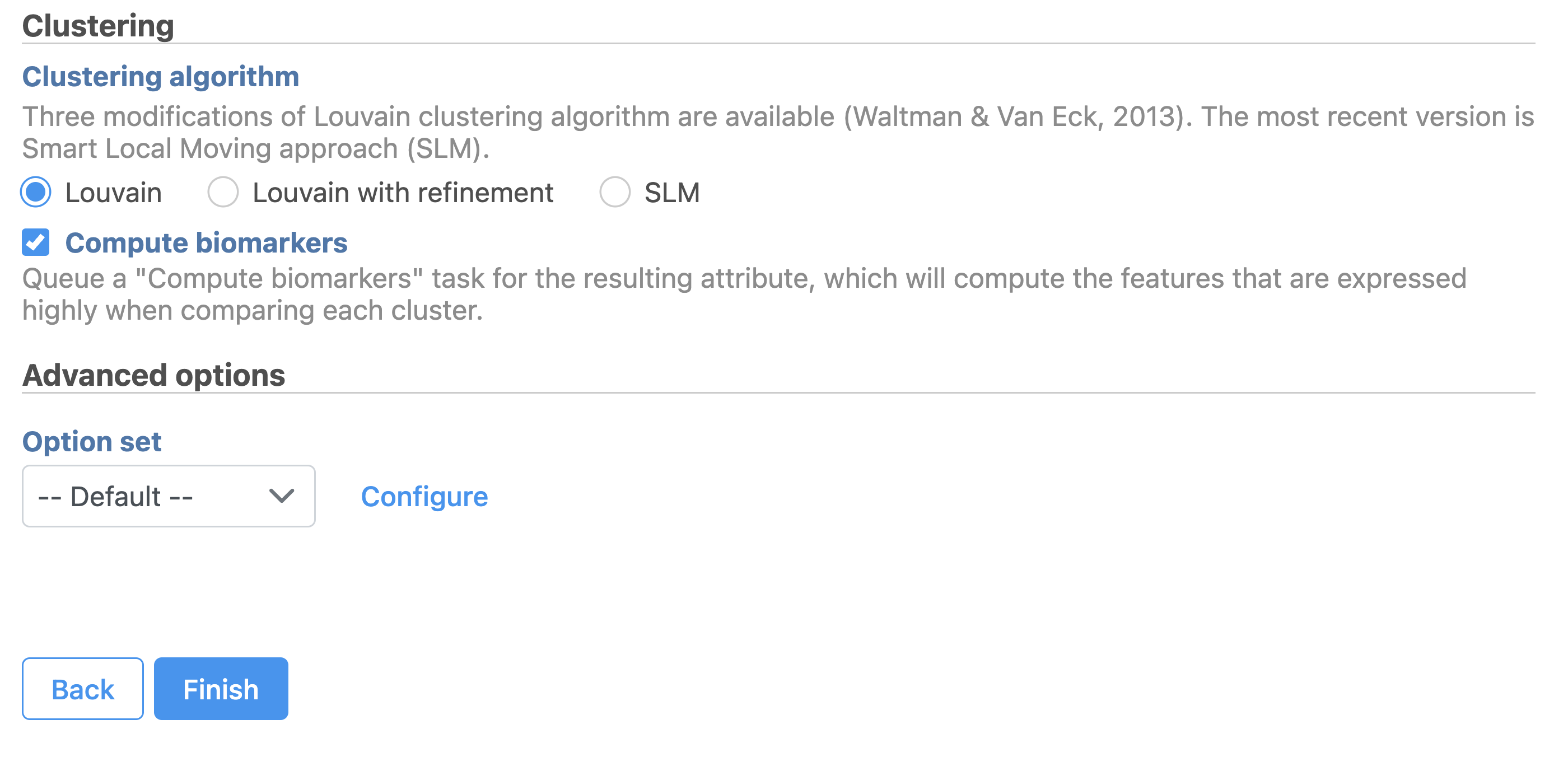 Image Added Image Added
|
A new Graph-based clusters data and a Biomarkers data node will be generated.
...
| Numbered figure captions |
|---|
| SubtitleText | Graph-based clustering results in Flow. |
|---|
| AnchorName | Graph-based clustering results |
|---|
|
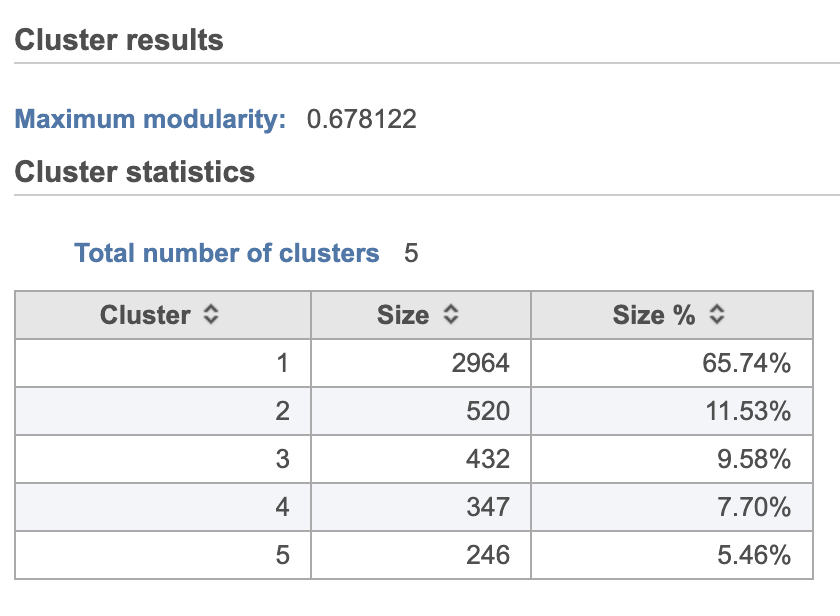 Image Removed Image Removed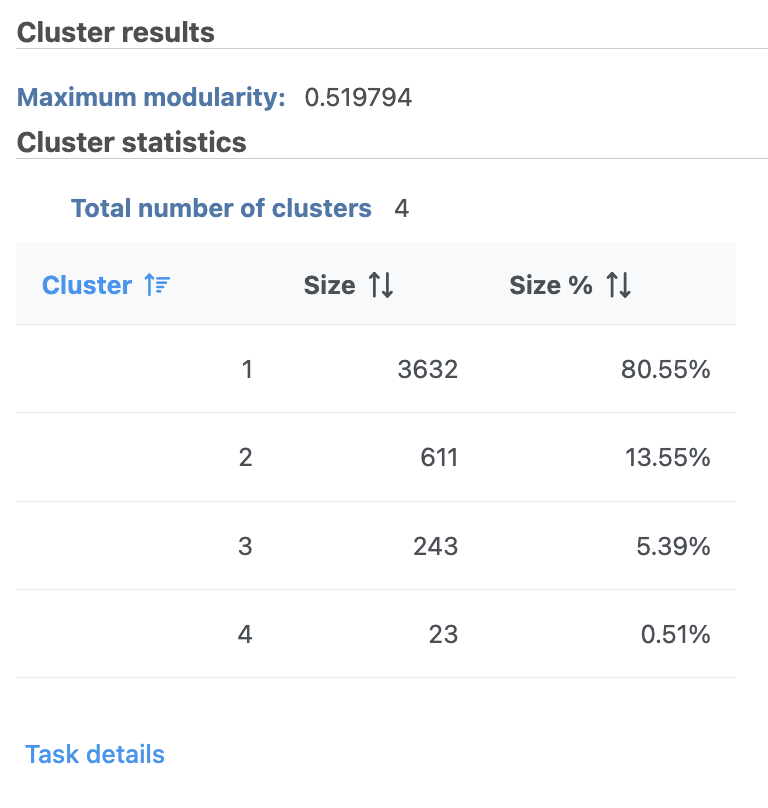 Image Added Image Added
|
| Numbered figure captions |
|---|
| SubtitleText | Computer biomarkers results in Flow. |
|---|
| AnchorName | Computer biomarkers results |
|---|
|
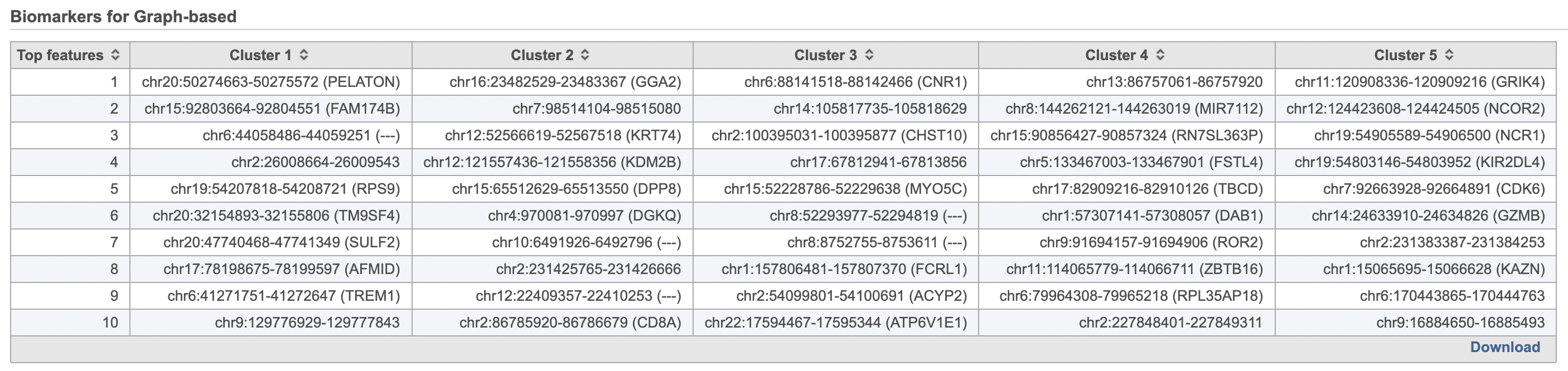 Image Removed Image Removed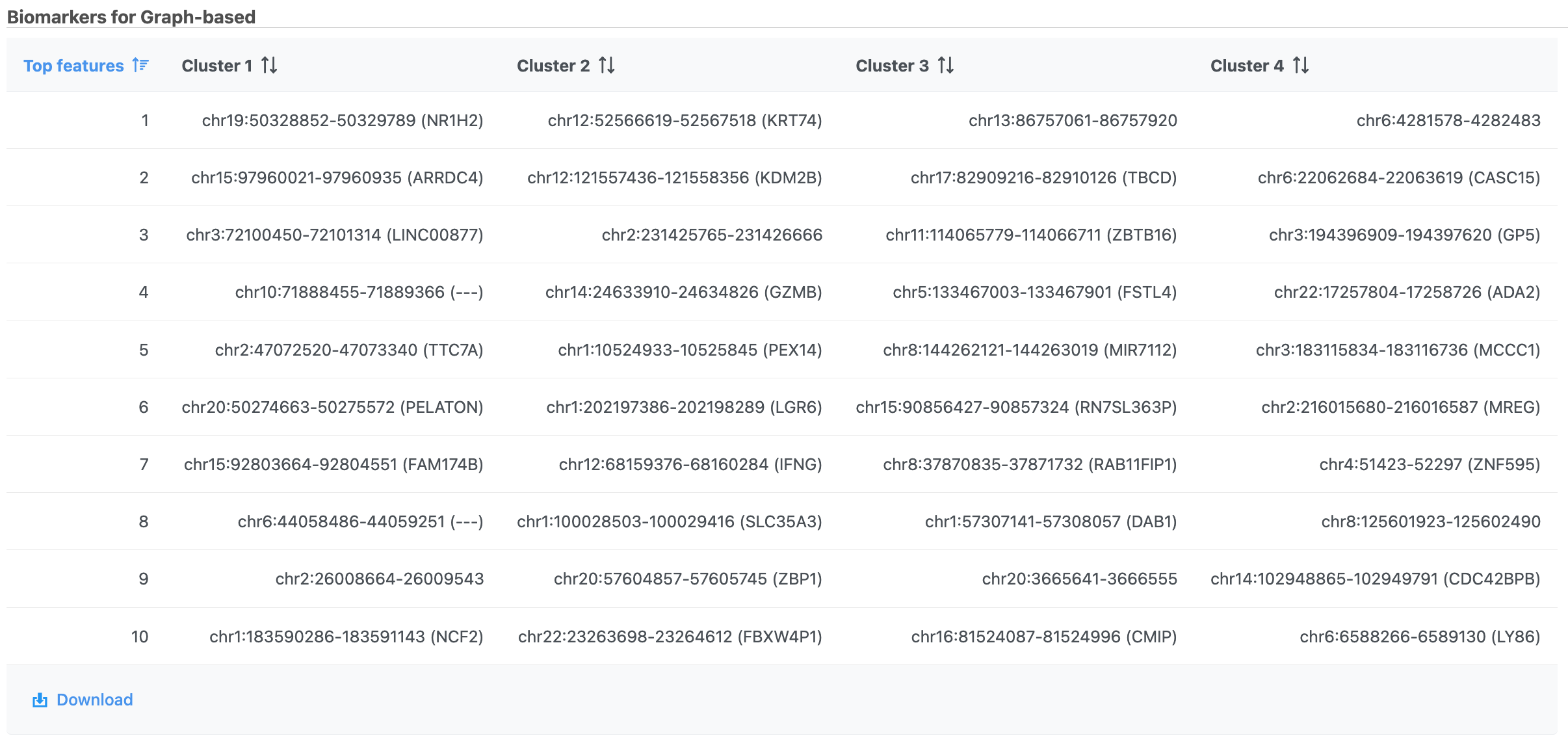 Image Added Image Added
|
UMAP
Similar to t-SNE, Uniform Manifold Approximation and Projection (UMAP) is a dimensional reduction technique. UMAP aims to preserve the essential high-dimensional structure and present it in a low-dimensional representation. UMAP is particularly useful for visually identifying groups of similar samples or cells in large high-dimensional data sets.
...
| Numbered figure captions |
|---|
| SubtitleText | UMAP configuration in Partek Flow. |
|---|
| AnchorName | UMAP configuration |
|---|
|
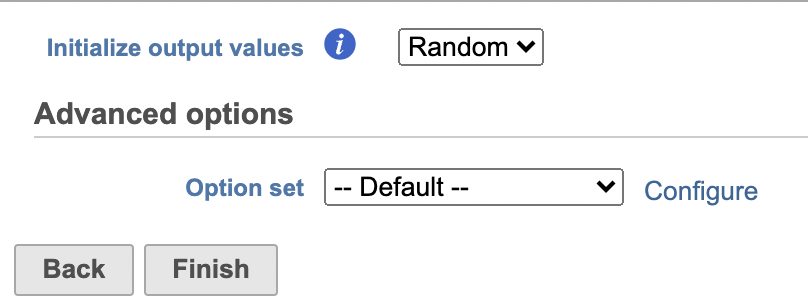 Image Removed Image Removed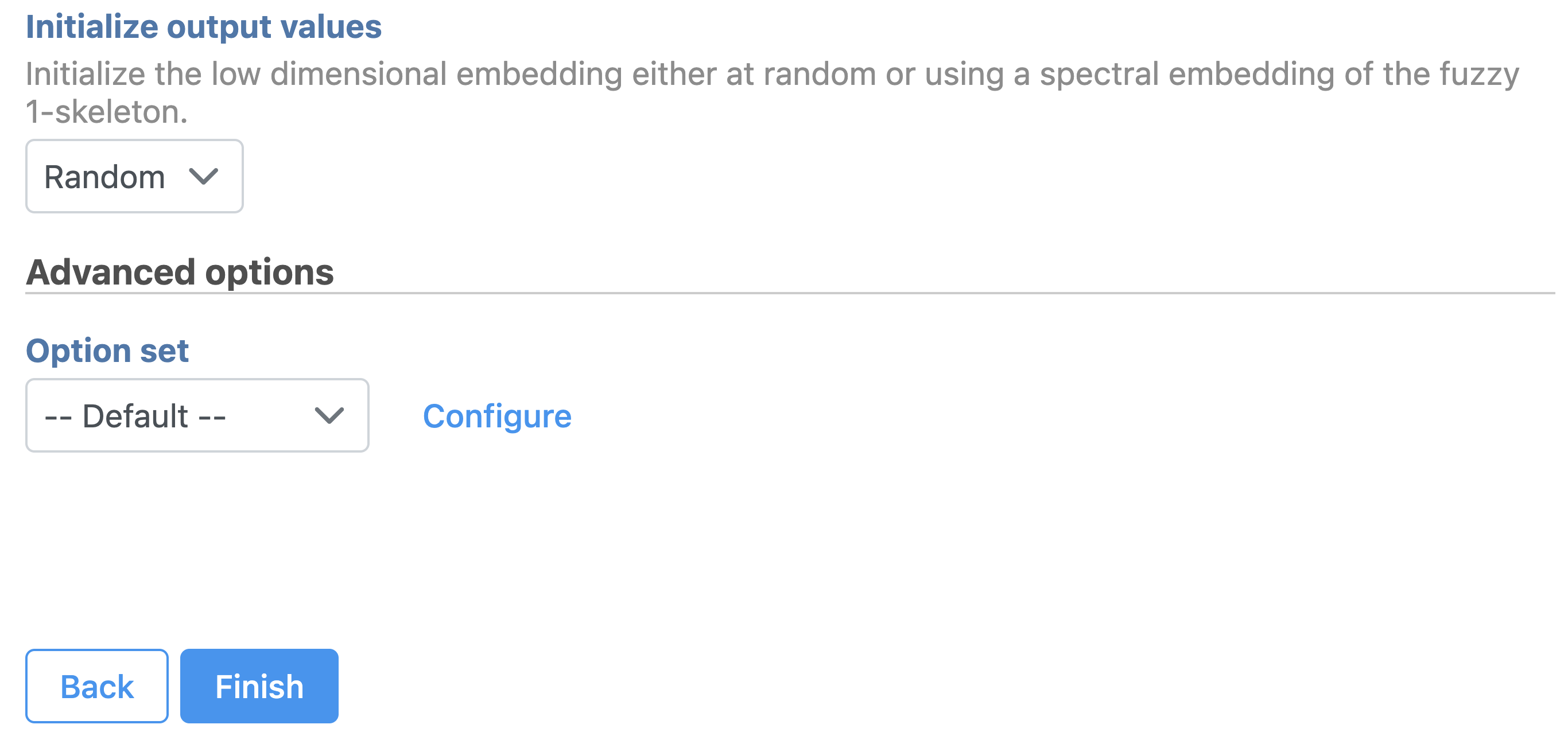 Image Added Image Added
|
Promoter sum matrix
...
| Numbered figure captions |
|---|
| SubtitleText | Promoter sum matrix in Flow. |
|---|
| AnchorName | Promoter sum matrix |
|---|
|
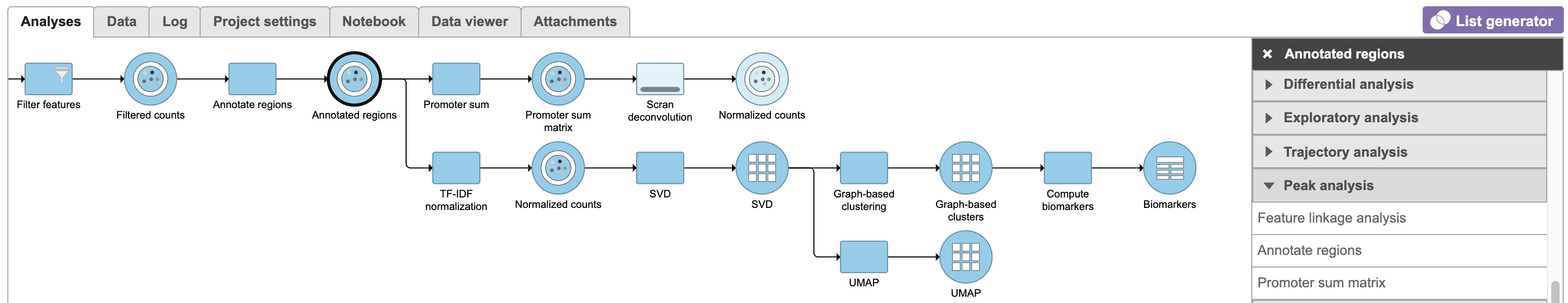 Image Removed Image Removed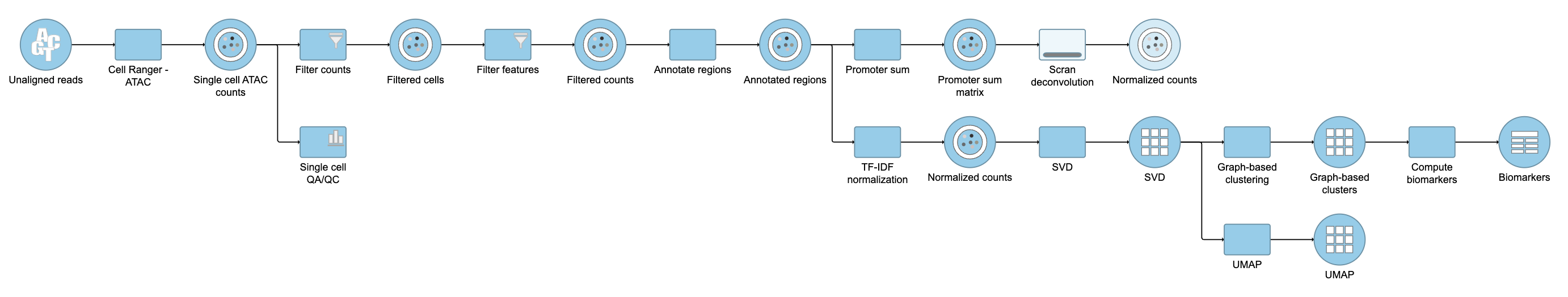 Image Added Image Added
|
Classifying cells
...
- Make sure the right data source has been selected. For scATAC-seq data, it shall be the normalized counts of promoter sum values in most cases (Figure 17)
- Set Color by in the Style configuration to the normalized counts node
- Type MS4A1 in the search box and select it. Rotate the 3D plot if you need to see this cluster more clearly.
- Click Click
 Image Added to activate Lasso mode
Image Added to activate Lasso mode - Draw a lasso around the cluster of MS4A1-expressing cells
- Click Classify selection under Tools in the left panel
- Type B cells for the Name
- Click Save (Figure 18)
...
| Numbered figure captions |
|---|
| SubtitleText | Select the data source in Data Viewer. |
|---|
| AnchorName | Select the data node |
|---|
|
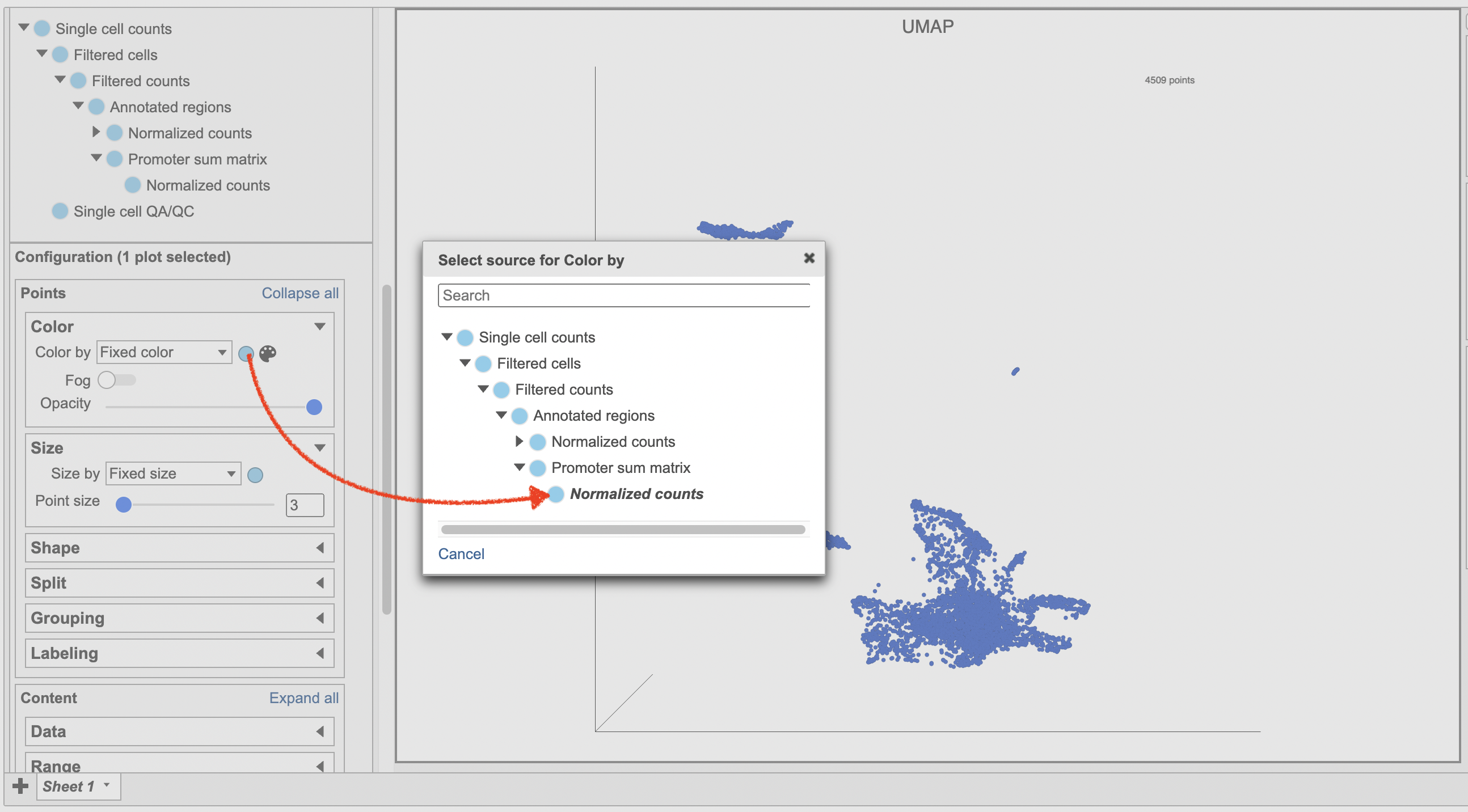 Image Removed Image Removed Image Added Image Added
|
| Numbered figure captions |
|---|
| SubtitleText | Color cells in UMAP by MS4A1 in Flow. |
|---|
| AnchorName | Coloring by MS4A1 |
|---|
|
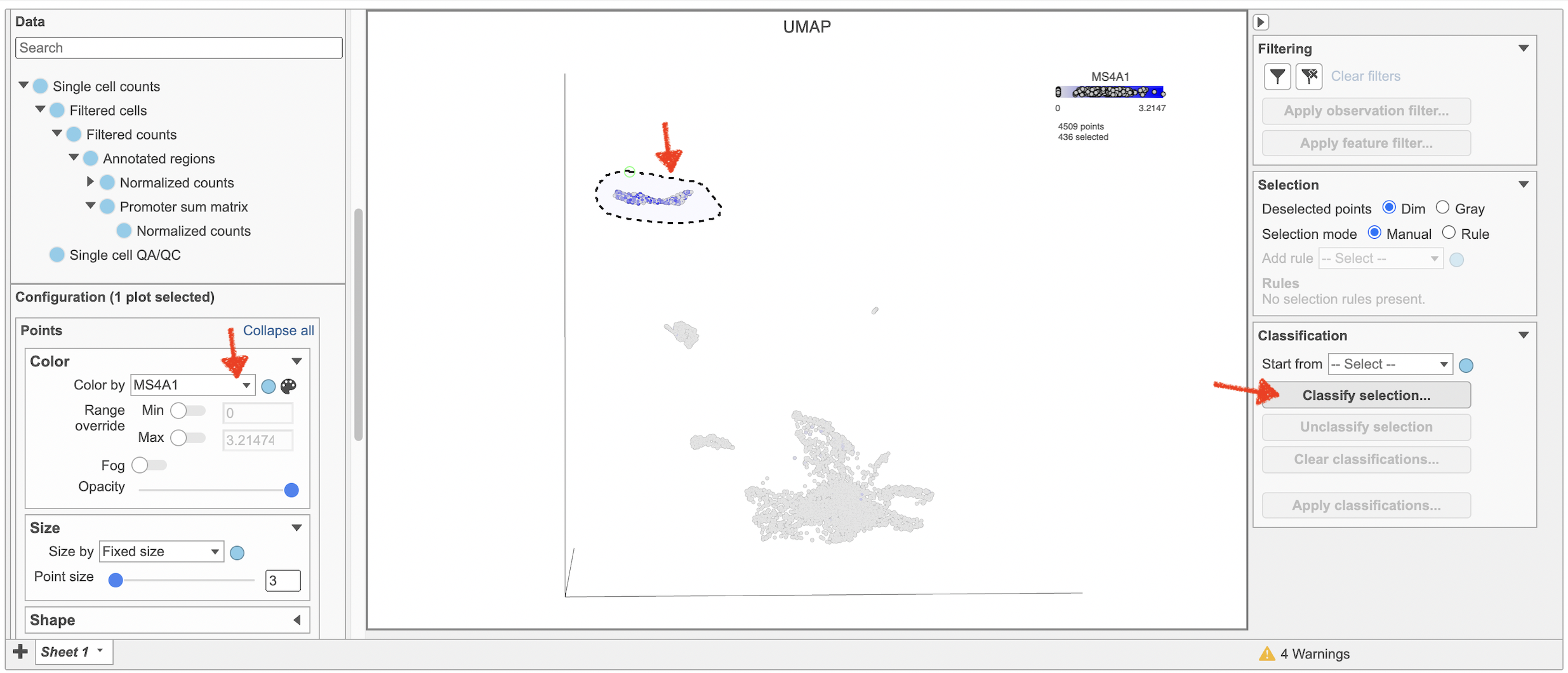 Image Removed Image Removed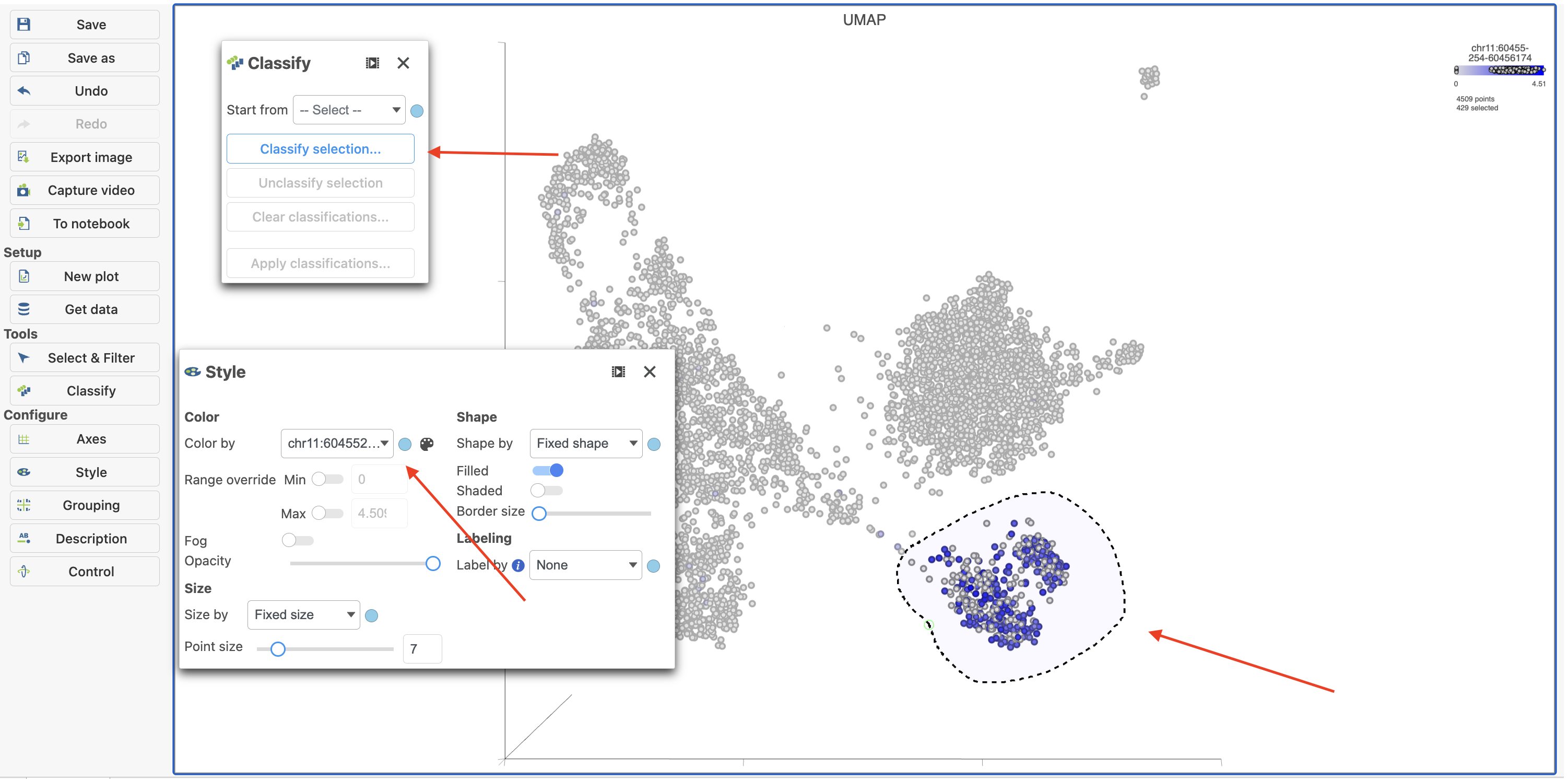 Image Added Image Added
|
Differential analysis
...
| Numbered figure captions |
|---|
| SubtitleText | Hurdle model for differential analysis in Flow. |
|---|
| AnchorName | Hurdle model |
|---|
|
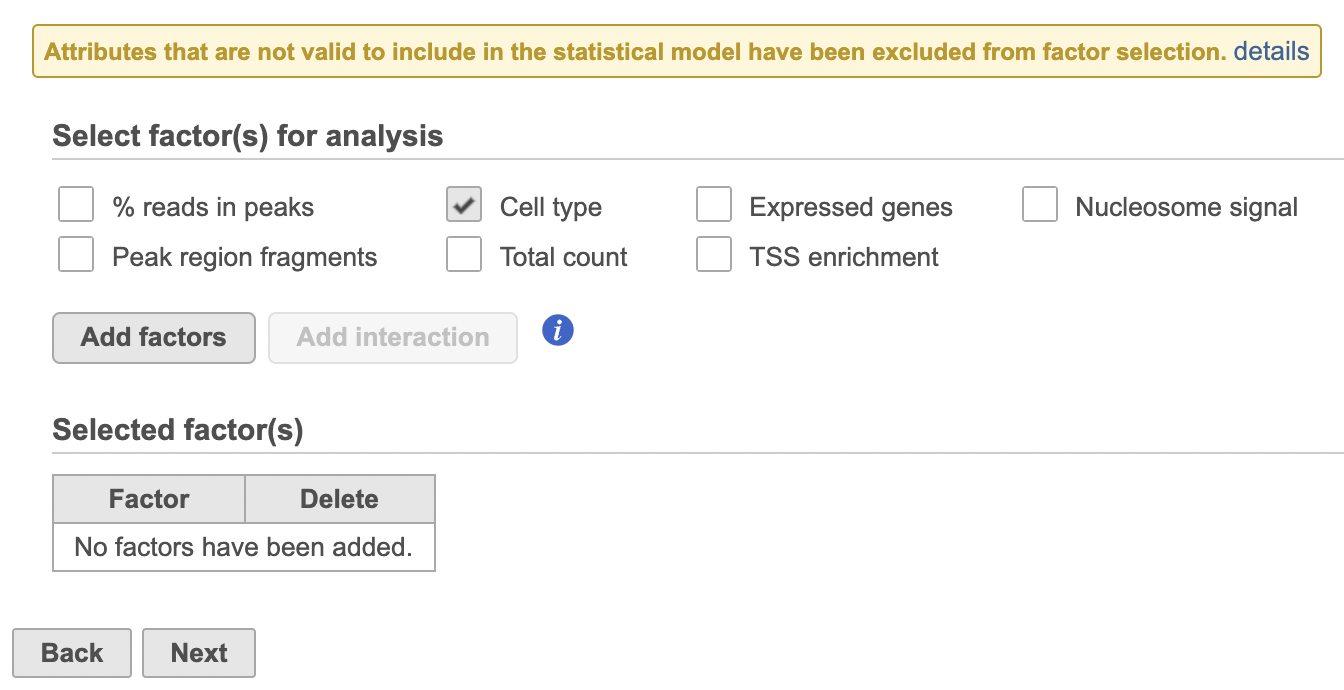 Image Removed Image Removed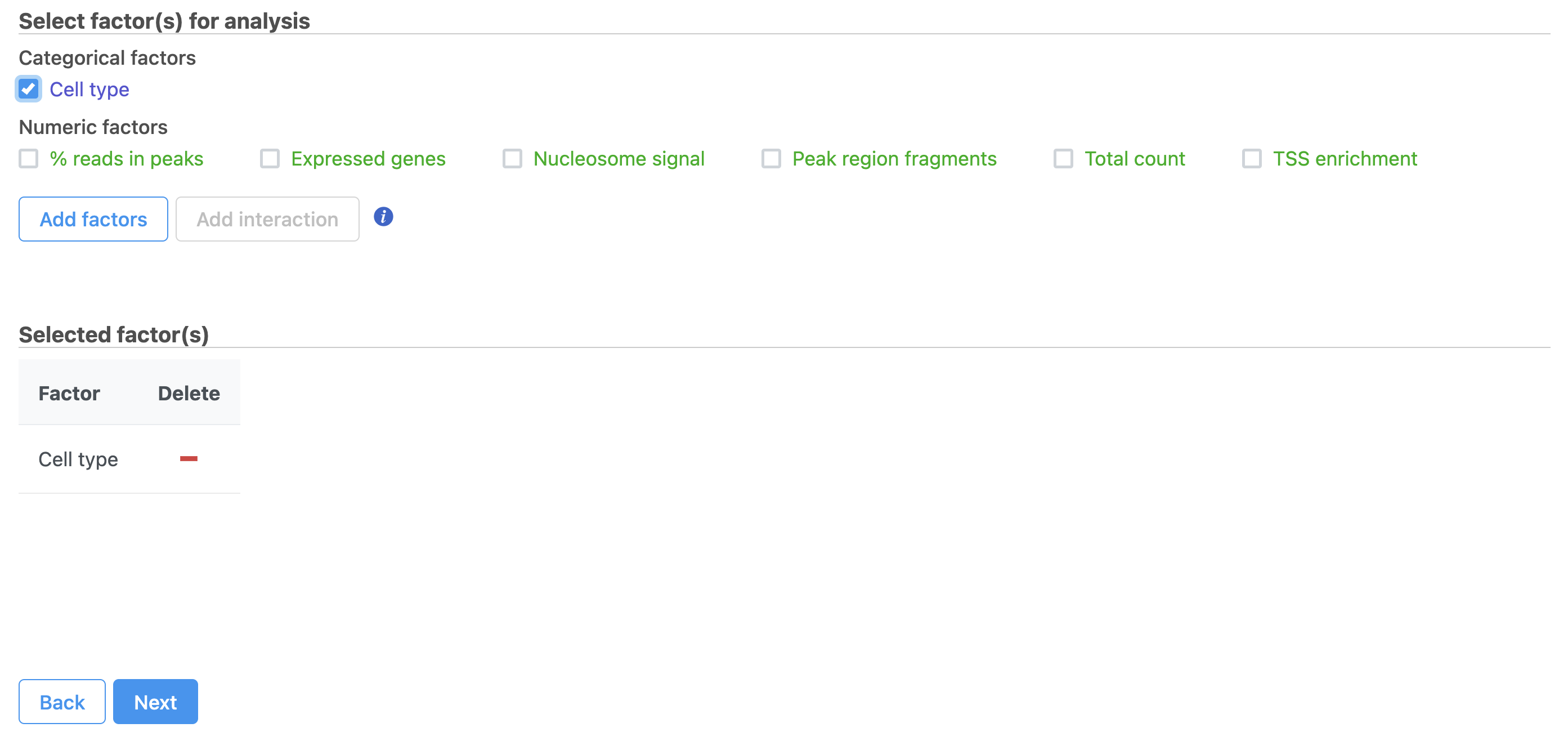 Image Added Image Added
|
- Click Next
- Define comparisons between factor or interaction levels (Figure 20)
- Click Add comparison to add the comparison to the Comparisons table.
- Click Finish to run the statistical test as default
| Numbered figure captions |
|---|
| SubtitleText | Define comparisons in Hurdle model. |
|---|
| AnchorName | Define comparisons |
|---|
|
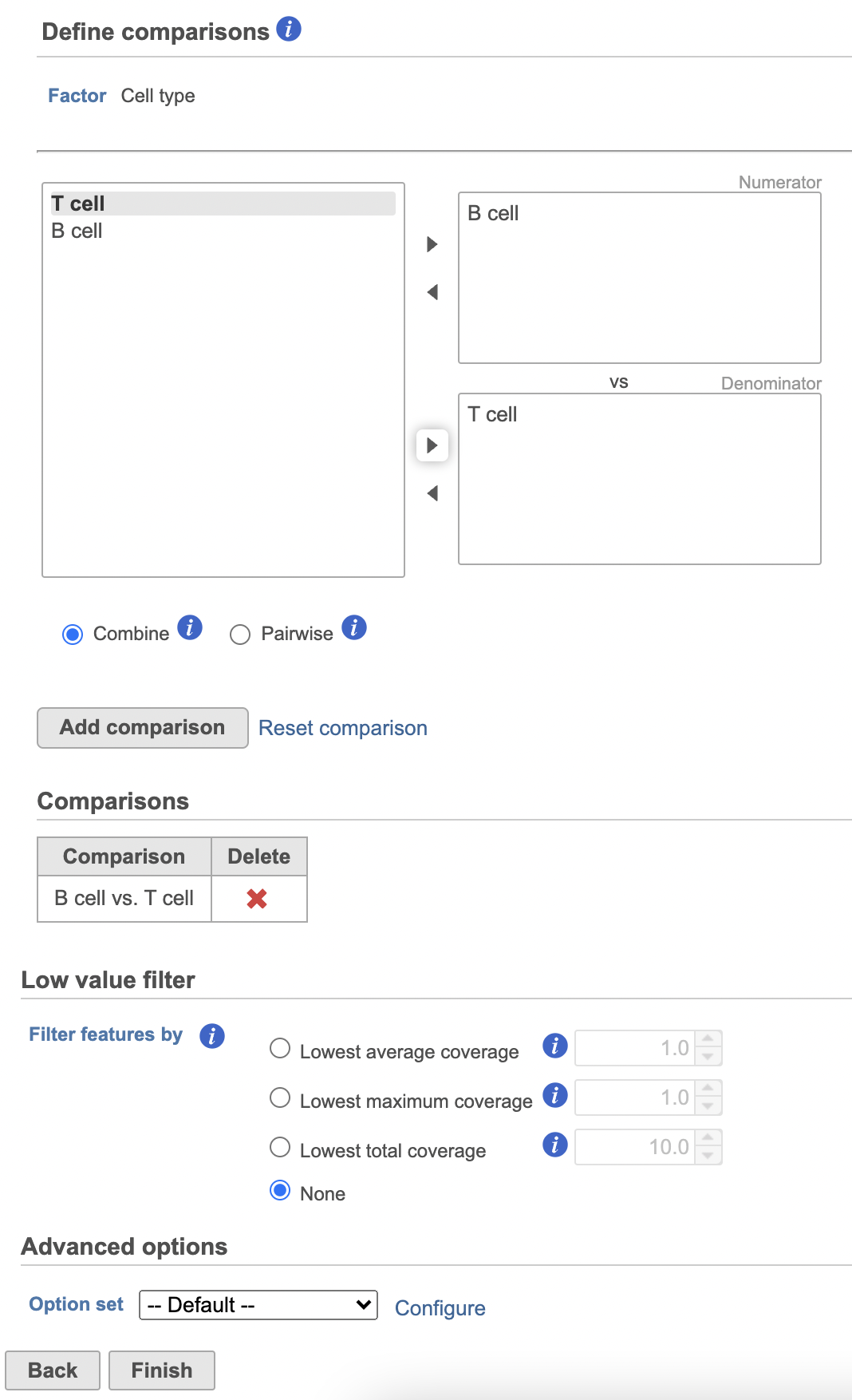 Image Removed Image Removed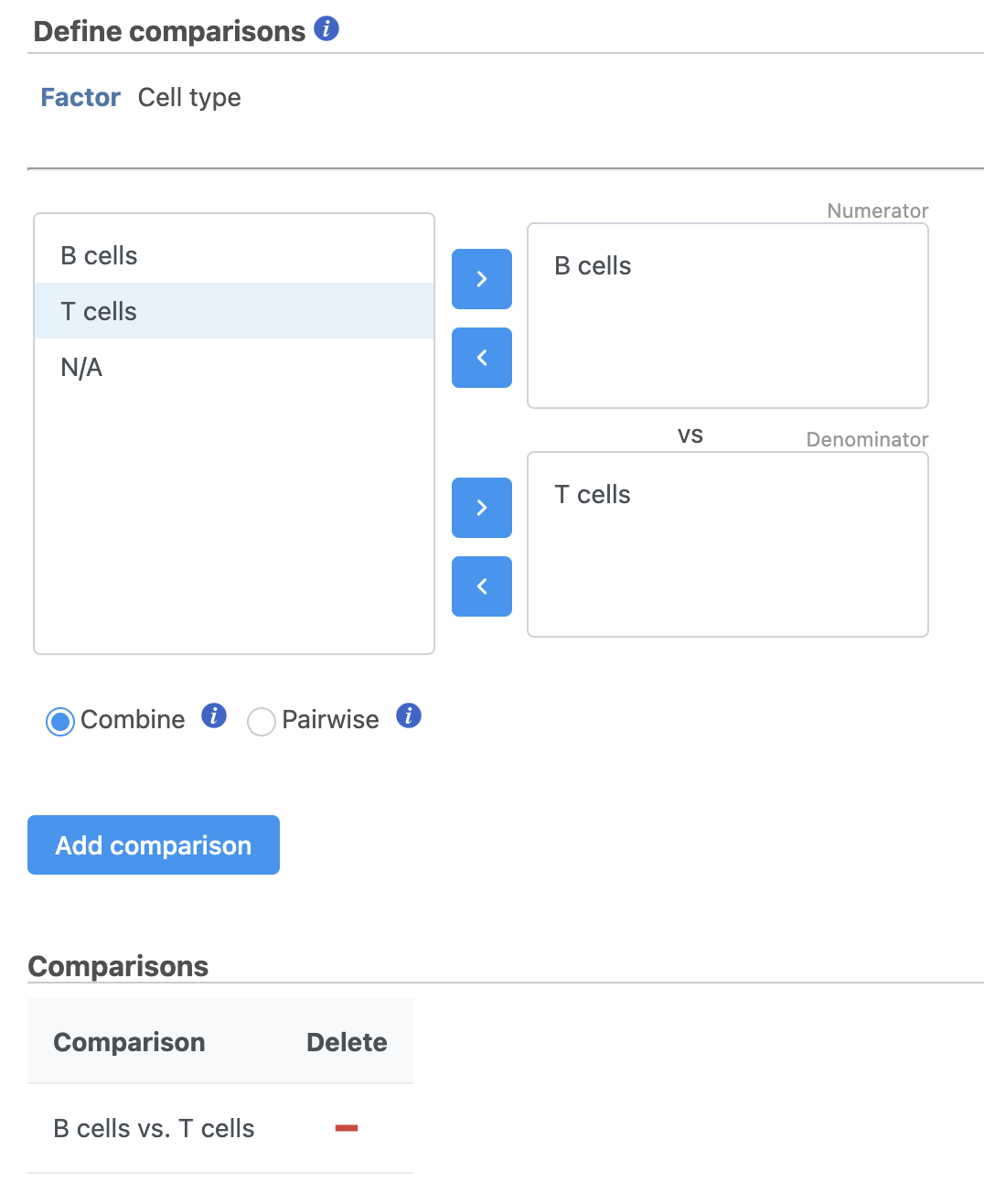 Image Added Image Added
|
Hurdle model produces a Feature list task node. The results table and options are the same as the GSA task report except the last two columns. The percentage of cells where the feature is detected (value is above the background threshold) in different groups (Pct(group1), Pct(group2)) are calculated and included in the Hurdle model report.
...
| Numbered figure captions |
|---|
| SubtitleText | Generate filtered node for differential analysis results in Flow. |
|---|
| AnchorName | Generate filtered node |
|---|
|
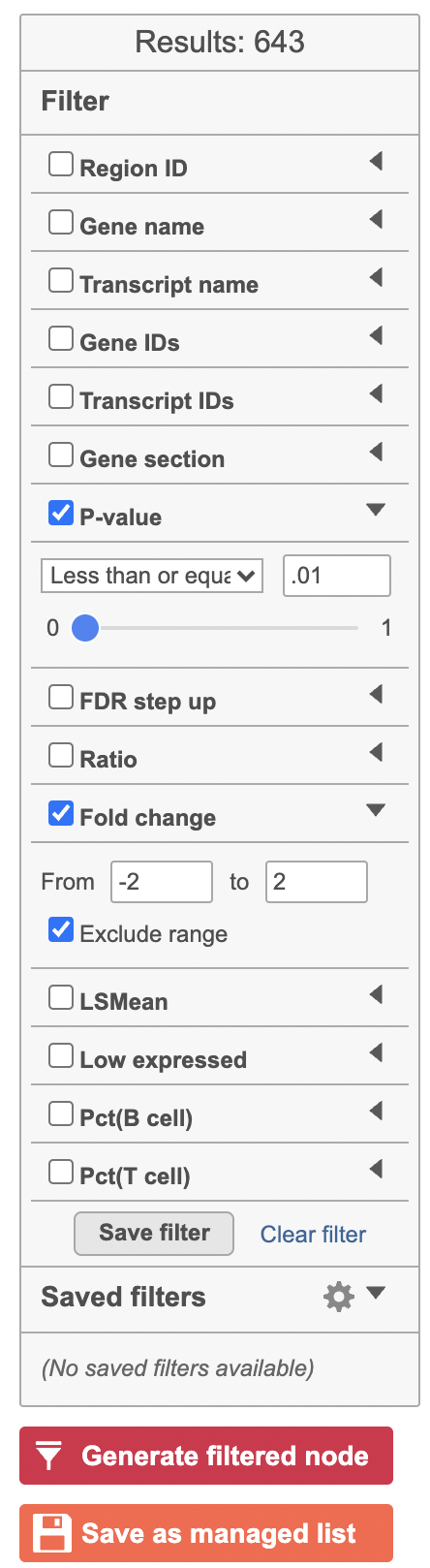 Image Removed Image Removed |
|
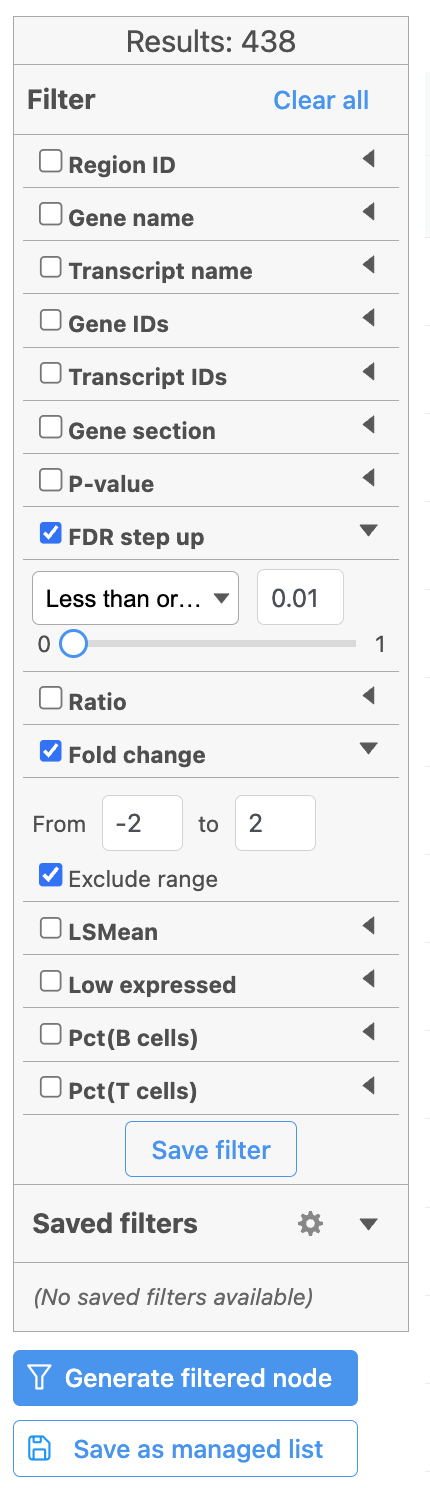 Image AddedOnce we have filtered a list of differentially expressed genes, we can visualize these genes by generating a heatmap, or perform the Gene set enrichment analysis and motif detection.
Image AddedOnce we have filtered a list of differentially expressed genes, we can visualize these genes by generating a heatmap, or perform the Gene set enrichment analysis and motif detection.
...

























