Page History
Data tracks section of the Select tracks dialog enables you to specify the tracks for visualization on the canvas. An overview of the available track types is provided in Figure 1. Note that not all tracks are visible at all times and that their presence depends on the whether specific data types are present in the project as well as the zoom level. The tracks can be customised and their appearance changed by using the control panel on the right.
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Alignments track Isoform proportion track Variants track Amino acids track Reads pileup track Probe intensities track Peaks track |
Alignments Track
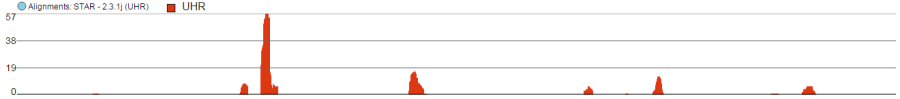
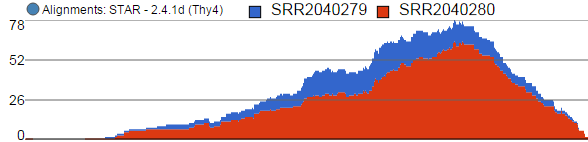
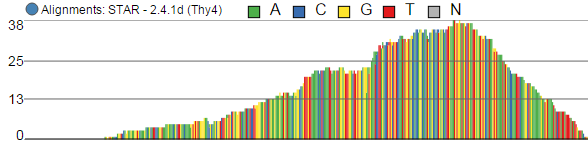
Alignments track displays a view of alignments present in .bam files in a stacked histogram fashion (similar to Partek® Genomics Suite®). The y-axis shows number of (raw) base calls per position. By default, reads are coloured by sample; the exception is invocation of the chromosome view on a variant table, when the reads are coloured by base calls. The difference is shown in Figure 2.
. Variations on the track are displayed below. They can be configured in the following ways:
- The histogram can be colored by sample or by base call. By default, it is colored by sample except when the chromosome viewer is invoked from a variant data table.
- When colouring reads by sample, the reads are stacked (on top of each other), i.e. in the example below there are more reads in the red sample than in the blue sample. This is an example of a Sum histogram type. This can also be configured to display overlays or averages.
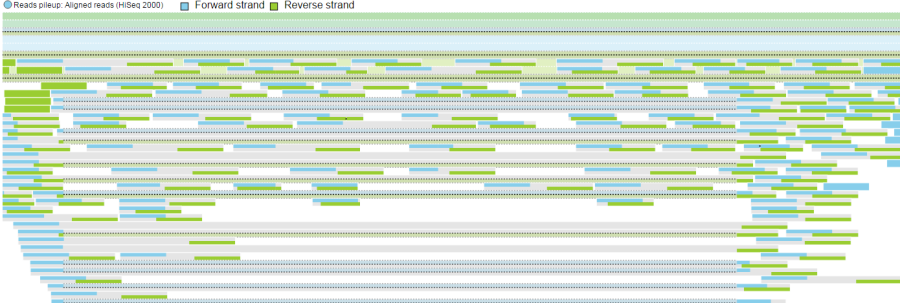
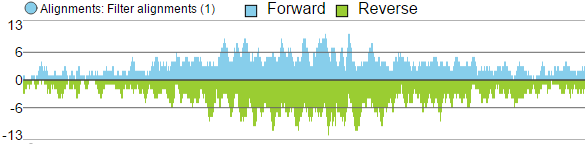
- Reads can be split into two tracks corresponding to the strand that they map to. This can be invoked by clicking the Split read histogram by strand checkbox.
For more information on configuring tracks see our page on Customizing the chromosome viewer
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Reads coloured by sample Reads coloured by base calls Reads split by strand |
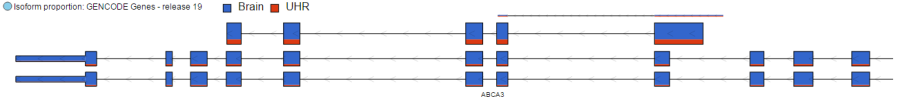
Isoform Proportion Track
The Isoform proportion track displays the reads mapped to transcripts and helps to visualize differential expression and alternative splicing, using standard symbols for exons (boxes) and introns (lines connecting the boxes). The size of each transcript is proportional to the number of reads that map to that transcript. The color indicates the samples to which the reads belong. Figure 3 shows a gene with two transcripts in RefSeq database; the top transcript is more abundant than the bottom transcript and is preferentially expressed in the "blue" condition (labeled as 0 uM). The bottom transcript, on the other hand, seems to be expressed at the same level across all three conditions (i.e. 0 uM, 5 uM, 10 uM). The number and structure of transcripts on the plot depend on the transcript model that was used for mapping.
...