Page History
...
A list of SNPs using dbSNP IDs can be imported as a text file and associated with an annotation file as described for a list of genes. The annotation file you use to annotate the SNPs should minimally contain the chromosome number and physical position of each locus.
Novel SNPs , or SNPs that are not found in your annotation source , must be imported as a region list. The only difference would be to For this, follow the procedure outlined in Starting with a list of genomic regions, but use the SNP name in place of a region name.
...
Starting with a list of SNPs that have been associated with genomic loci using an annotation file and assigned a species with genome build, you may can use Find Overlapping Genes to annotate these SNPs with the closest genes.
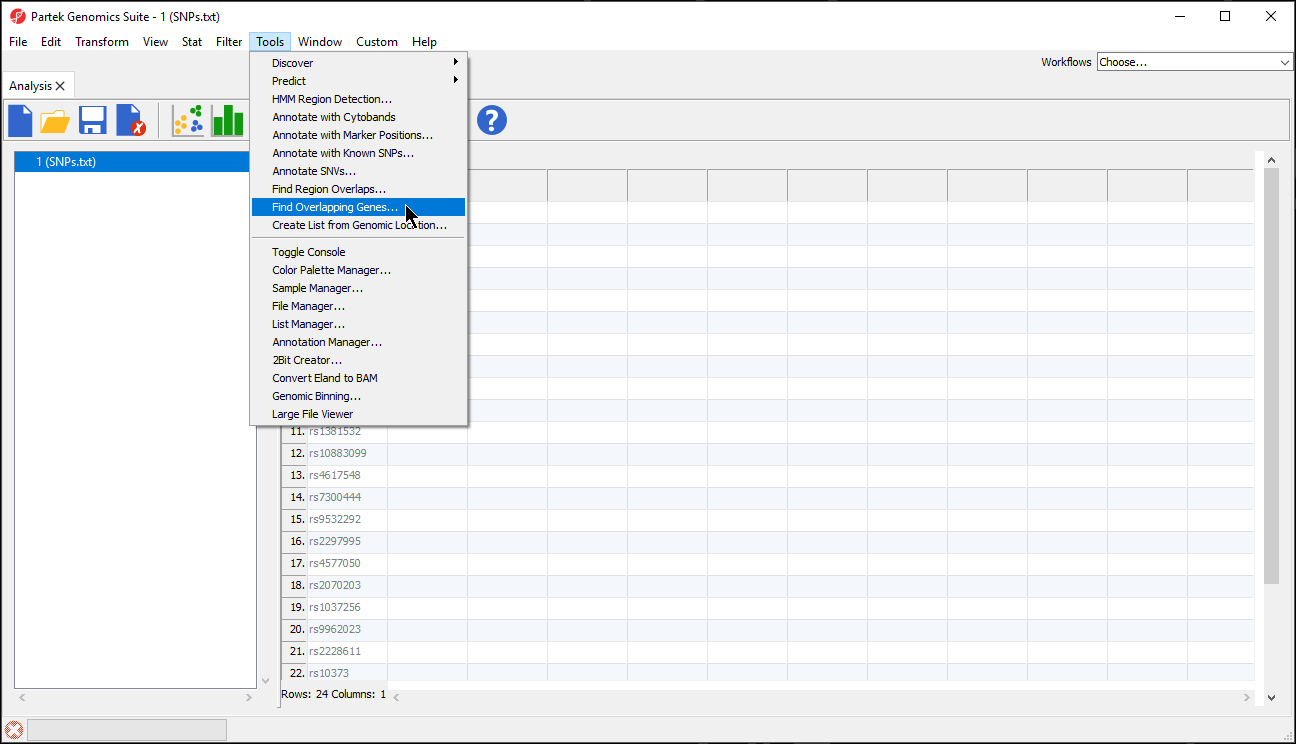
- Select Tools from the main toolbar
- Select Find Overlapping Genes (Figure 1)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...