A list of SNPs using dbSNP IDs can be imported as a text file and associated with an annotation file as described for a list of genes. The annotation file you use to annotate the SNPs should minimally contain the chromosome number and physical position of each locus.
Novel SNPs or SNPs that are not found in your annotation source must be imported as a region list. For this, follow the procedure outlined in Starting with a list of genomic regions, but use the SNP name in place of a region name.
Annotating SNPs with genes
Starting with a list of SNPs that have been associated with genomic loci using an annotation file and assigned a species with genome build, you can use Find Overlapping Genes to annotate these SNPs with the closest genes.
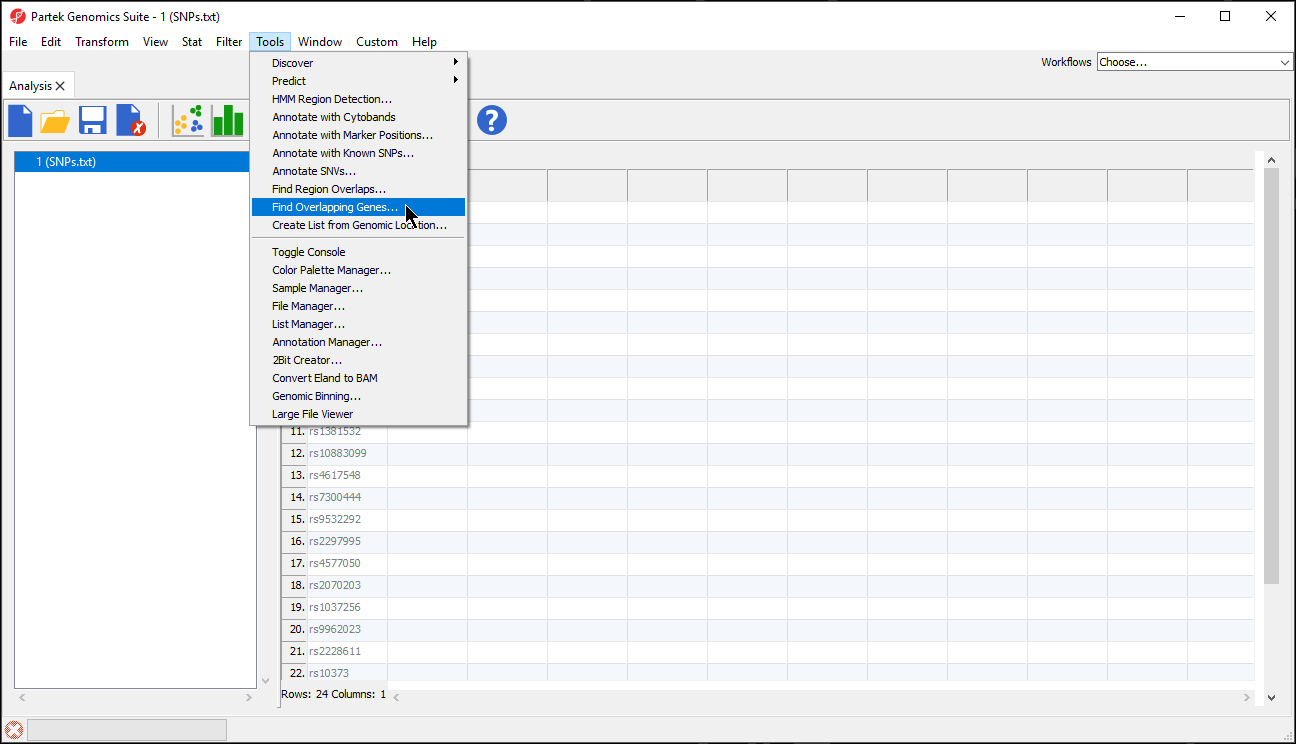
- Select Tools from the main toolbar
- Select Find Overlapping Genes (Figure 1)
- Select Add a New Column with the Gene Nearest to the Region from the method dialog
The Report Regions from the specified database dialog will open.
- Select your preferred database. Be sure to match the species and genome build of your SNP list
- Select OK
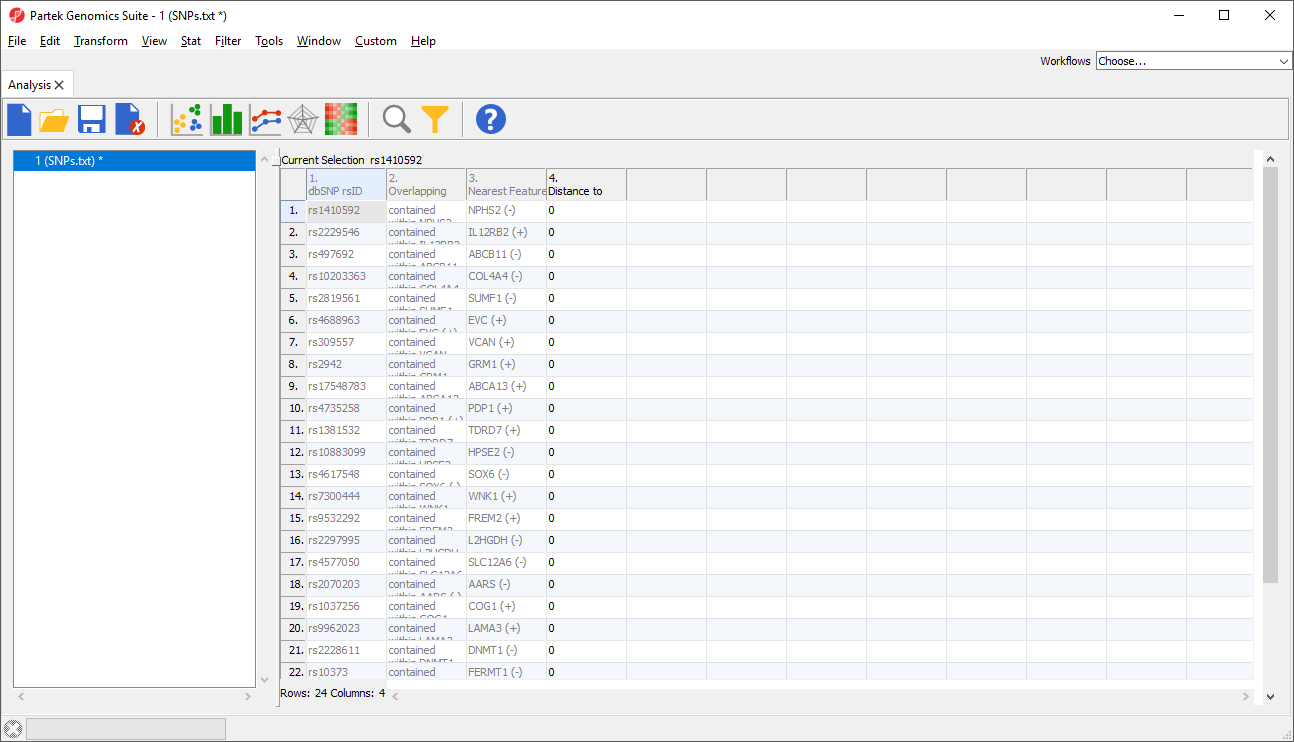
This will add 3 columns to the list of SNPs spreadsheet including Nearest Feature, which will indicate the nearest gene and strand (Figure 2).
To allow gene list operations such as GO Enrichment or Pathway Enrichment to be performed on the SNP list, we can set the Nearest Feature column as the gene symbol column for the spreadsheet.
- Right click the spreadsheet in the spreadsheet tree
- Select Properties from the pop-up menu
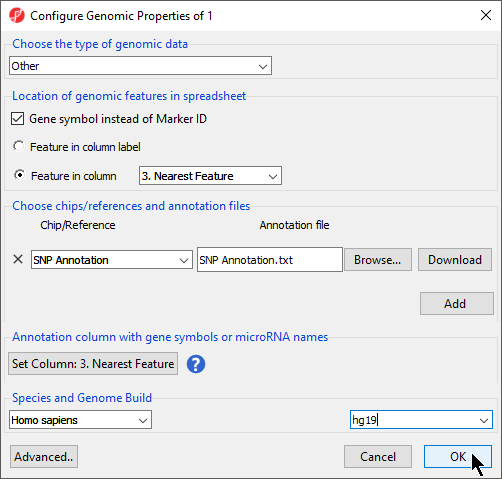
- Select Gene symbol instead of Marker ID
- Select Feature in column and select Nearest Feature (Figure 3)
- Select OK
Annotating a Partek Genomics Suite-generated SNP list with SNVs
If you have a SNP spreadsheet that was generated using Partek Genomics Suite (not imported as a .txt file), you can annotate the SNP list with gene, transcript, exon, and information about the predicted effect of the SNPs.
- Select Tools from the main command toolbar
- Select Annotate SNVs
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
35 | rates |