The significant CpG loci detected in the previous step actually form a methylation signature that differentiates , which we shall now visualise by a heat map.
- Select the (LCLs vs. B cells) spreadsheet in the spreadsheet pane on the left
- Select Cluster Based on Significant Genes from the Visualization panel of the Illumina BeadArray Methylation workflow
- Select Hierarchical Clustering for Specify Method (Figure 1)
| Numbered figure captions |
|---|
| SubtitleText | Selecting Heirarchical Clustering for clustering method |
|---|
| AnchorName | Selecting clustering method |
|---|
|
|
- Select OK
- Verify that (CpG of interest) is selected in the drop-down menu
- Select Standardize for Expression normalization (Figure 2)
| Numbered figure captions |
|---|
| SubtitleText | Selecting spreadsheet and normalization method for clustering |
|---|
| AnchorName | Selecting normalization for clustering |
|---|
|
|
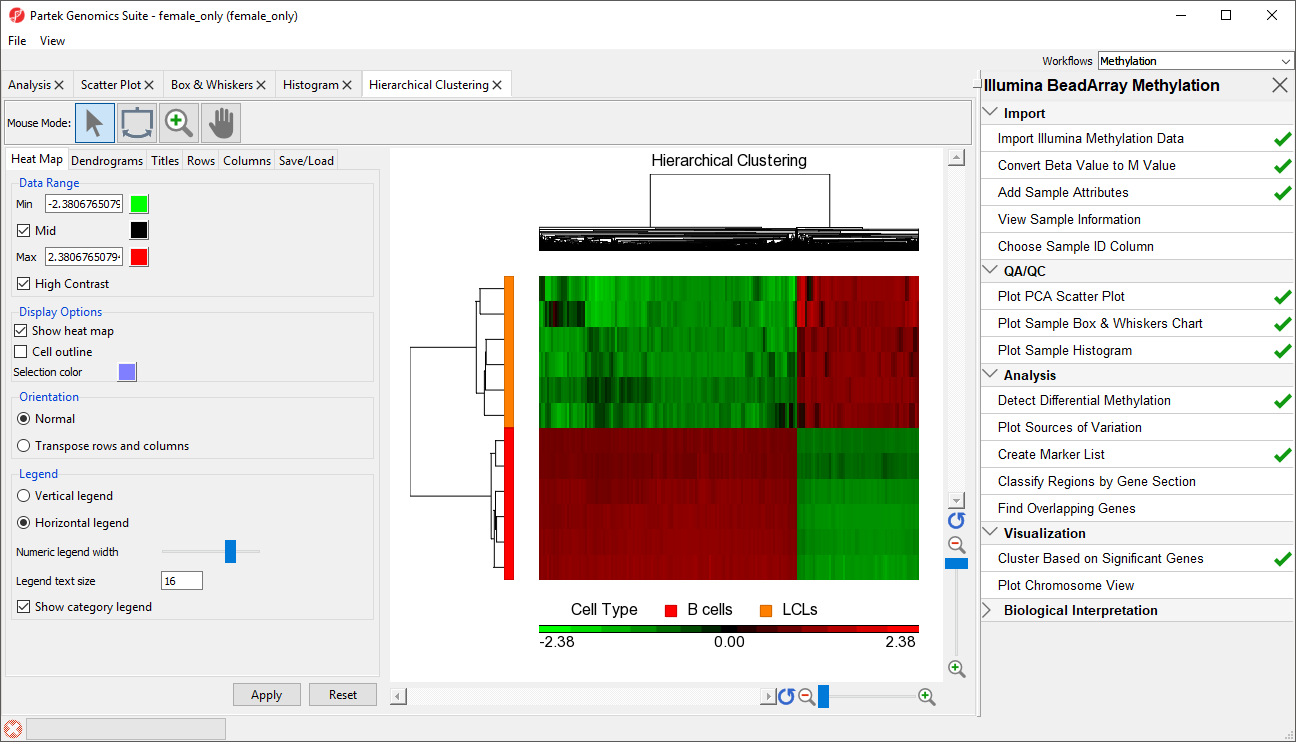
The heat map will be displayed on the Hierarchical Clustering tab (Figure 3).
| Numbered figure captions |
|---|
| SubtitleText | Hierarchical clustering with heat map invoked on a list of significant CpG loci |
|---|
| AnchorName | heat map |
|---|
|
|
The experimental groups are rows, while the CpG loci from the
(LCLs vs. B cells) spreadsheet are columns. Methylation levels are compared between the LCLs and B cells groups. CpG loci with higher methylation are colored red, CpG loci with lower methylation are colored green. B cells samples are colored red and LCLs samples are colored orange in the dendrogram on the the left-hand side of the heat map.


