Binding sites for the DNA-binding protein of interest are indicated by peaks of enriched sequencing read density. How are peaks calculated from reads in Partek Genomics Suite?
Using the effective fragment length calculated by Cross Strand-Correlation, each read is extended in the 3' direction by the effective fragment length and overlapping extended reads are merged into single peaks. For paired-end reads, the distance between paired reads is used as the fragment length and overlapping fragments are merged into peaks. For peak detection, the genome is divided into windows of a user-defined size and the number of fragments whose mid-points fall within each window is counted. A model for expected read density (a zero-truncated negative binomial) is used to determine which peaks are significantly enriched over a user-defined false discovery rate (FDR). See the ChIP-Seq white paper for more information on the peak-finding algorithm and tips for setting the Fragment extension and window sizes.
- Select spreadsheet 1 (ChIP-Seq) from the spreadsheet tree
- Select Detect peaks from the Peak Analysis section of the ChIP-Seq workflow
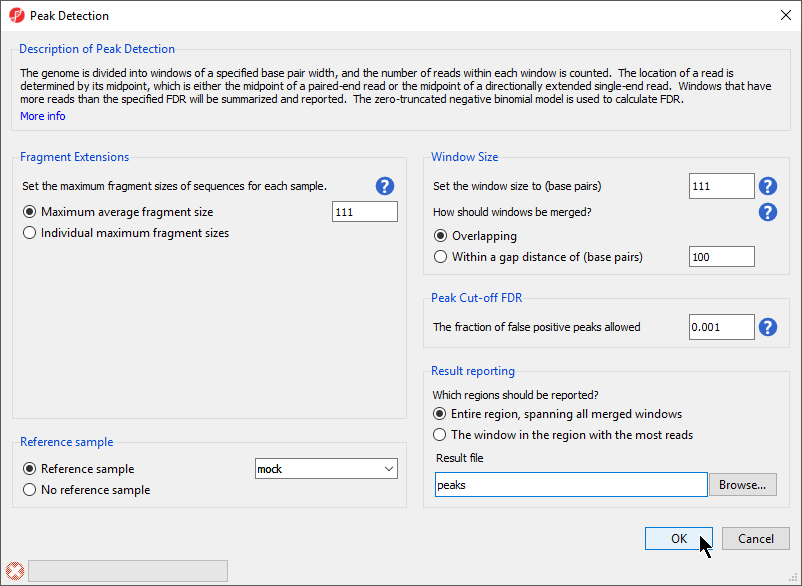
The Peak Detection dialog will open. Configure the dialog as shown (Figure 1).
- Select Maximum average fragment size for Fragment Extensions
- Set Maximum average fragment size to 111
Your choice for Maximum average fragment size is based on your experimental design. If you have used an antibody that binds DNA as the control antibody such as an IgG control, you could use different fragment lengths for each sample based on its effective fragment length by selecting the Individual maximum fragment sizes option. Here, we have chosen the effective fragment length of 111 base pairs calculated using Cross Strand-Correlation.
- Select Reference sample from Reference sample
- Select mock from the Reference sample drop-down menu
- Set Set the window size to (base pairs) to 111
The peak detection algorithm divides the genome into windows to find windows with enriched for reads based on an FDR cut-off value. Here, we have chosen to match the window and individual maximum fragment sizes.
- Select Overlapping for How should windows be merged?
- Set The fraction of false positive peaks allowed to 0.001
The Peak Cut-off FDR determines the cut-off for calling peaks. Setting a lower value demands greater differences between mock and chip samples for a peak to be called; a false discovery rate of 0.001 anticipates 1 false positive per 1000 peaks called.
- Select Entire region, spanning all merged windows for Which regions should be reported?
Optimal peak detection settings are dependent on your experimental design and data so fine tuning may be required. Because transcription factor binding sites tend to have localized and sharp clusters of reads, the window size used during the analysis of a transcription factor study can be left relatively small, approximately the same as the average fragment length, and the option to allow for gaps between enriched windows does not need to used. Region in the window with most reads could also be selected to report a more narrow region for each peak call. Conversely, histone modification peaks tend to be subtle and diffuse. To analyze histone modification ChIP-seq data, larger window sizes, combining neighboring windows into larger windows using Within a gap distance of, and reporting entire regions using Entire region, spanning all merged windows might be appropriate.
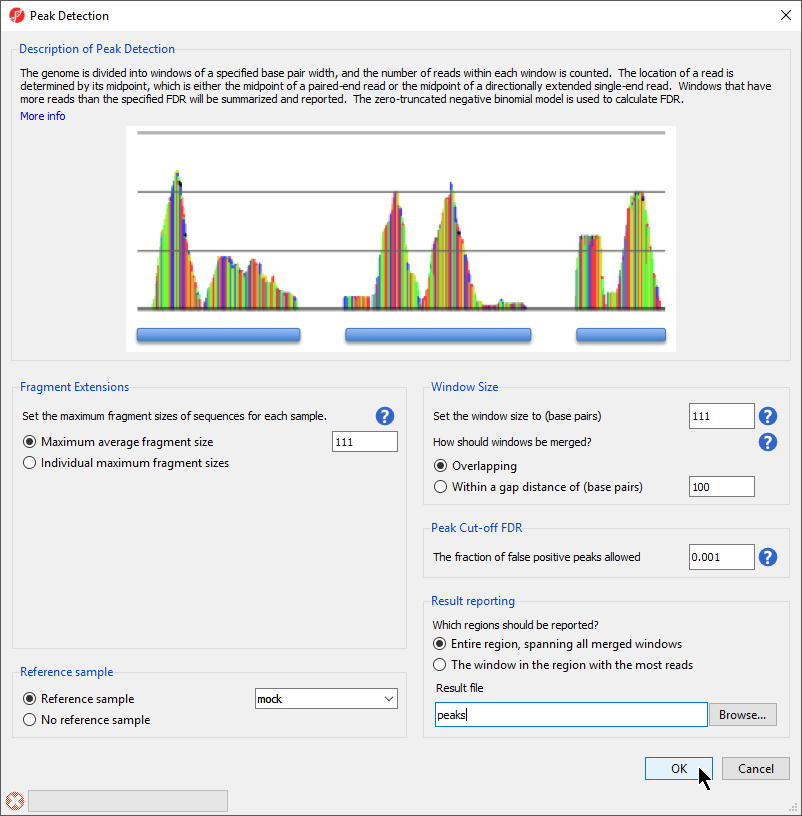
A convenient way to visualize the relationship between window size and gap size is to select the More info link at the top of the Peak Detection dialog box. A simulated read count histogram will open below the Description of Peak Detection section (Figure 2). The blue bars underneath the histogram will reflect how regions are detected and reported using your current Peak Detection settings. Try changing the How should windows be merged or Which regions should be reported? options to visualize their effects on peak detection.
- Select OK to run the peak detection algorithm with your chosen settings
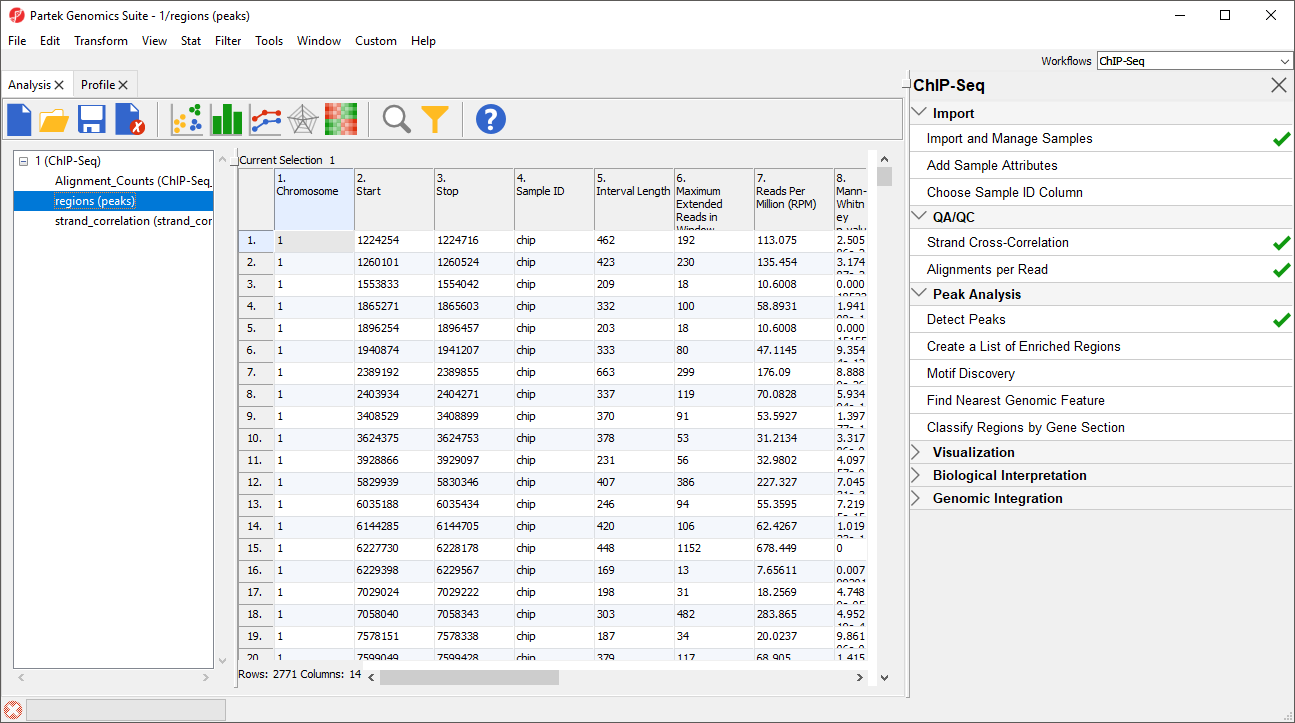
Peak Detection generates a new child spreadsheet, regions (peaks) (Figure 3).
This spreadsheet is sorted by chromosome number and genomic location. Each row represents one genomic region of peak enrichment. The columns are:
Column 1. Chromosome gives the chromosome location of region
Column 2. Start gives the start of region (inclusive)
Column 3. Stop gives the end of region (exclusive)
Column 4. Sample ID gives the sample containing the enriched region
Column 5. Interval length gives the length of the region, Start - Stop, in base pairs
Column 6. Maximum Extended Reads in Window gives the greatest number of extended reads in any of the windows of a region
Column 7. Reads per Million (RPM) divides the total number of aligned reads in the sample (in millions). This helps you compare peaks across samples, especially when there is a large difference in the number of aligned reads between samples.
Column 8. Mann-Whitney p-value identifies the separation between forward and reverse peaks for single-end reads using the Mann-Whitney U-test. Lower p-values indicate better separation. This p-value can be used if there was no control sample or to eliminate regions called due to PCR bias.
Columns 9-10. Total reads in region gives the total number of non-extended reads for each sample in the given genomic region. One column for each sample.
Column 11. p-value(Sample ID. vs. mock) compares the sample specified in column 4 to the reference sample for this genomic region using a one-tailed binomial test. A low p-value means there are significantly more reads in the sample specified in column 4 than in the mock sample. This column is only included if a reference sample is specified.
Column 12. scaled fold change (Sample ID vs. mock) compares intensity of signal between the sample specified in column 4 to the reference sample in the given genomic region. The fold-change is scaled by a ratio of the number of reads for each sample (IP vs. control) on a per-chromosome basis. Scaled fold changes >1 indicate more enrichment in the IP-sample than in the control sample. This column is only included if a reference sample is specified.
Columns 13 -14. <Sample ID> overlap percent gives the fraction of the given genomic region that overlaps a called peak region from the indicated sample. For example, the values of 100% in column 13 and 0% in column 14 indicate regions detected in the chip sample, but not in the mock sample. Similarly, regions with the value of 100% in column 14 were detected in the mock sample.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
36 | rates |