The Combine alignments task can potentially maximize the number of reads that map to a region. When unaligned reads resulting from one aligner are then aligned using a different one, the two resulting alignments can combined together for downstream analysis within Partek Flow. Note that this can only be performed if they were aligned to the same reference genome.
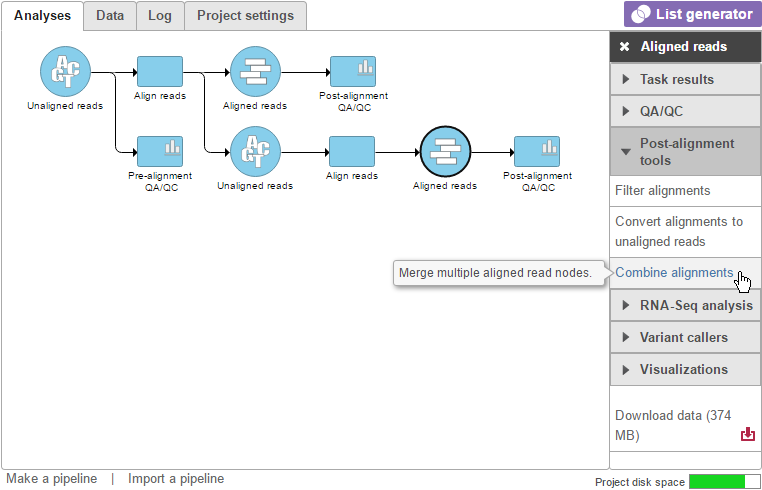
To invoke this task, click an Aligned reads data node and select Combine alignments (Figure 1).
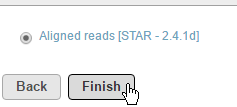
A list of compatible alignments will appear. The color of the text signifies the layer the alignment corresponds to (Figure 2). Select the alignment you would like to combine and click Finish.
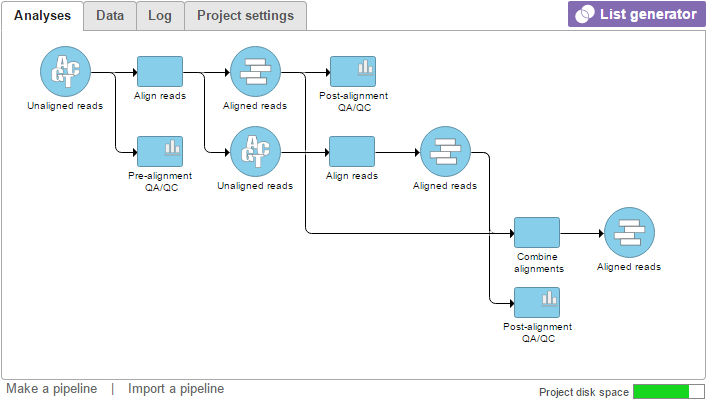
The resulting Aligned reads data node can now be used for downstream analysis (Figure 3).
Note that this task combines the files in the data node within Partek Flow but does not merge the BAM files. Downloading an aligned reads data node from a Combine alignments task will result to multiple BAM files per sample.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
43 | rates |
1 Comment
Melissa del Rosario
author: cignacio