Running a Chromosome View Task from a Data Node
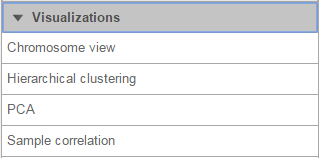
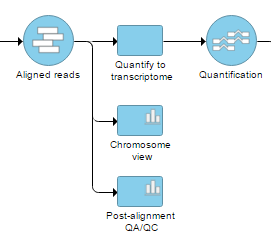
On the Analysis tab, selecting a data node containing aligned reads, variants, gene or transcript counts, or feature lists, shows Chromosome view in the Visualization section of the context-sensitive menu (Figure 1).
If Partek Flow has no information on the genome build, you will need to provide the species and genome build in a subsequent dialog (not shown). Otherwise, chromosome view will come up directly.
Browsing directly to a location directly from a Task Report
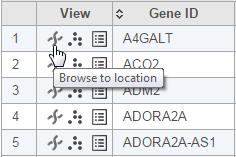
Another way to get the Chromosome view is through a Task report. You can launch the viewer by selecting the chromosome icon in the View column (Figure 3) of GSA or View Variants reports. In that case, the Chromosome view will browse directly to the selected genomic location (i.e. a transcript or a variant, depending on the pipeline).
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |