This task can be invoke from count matrix data node or clustering task report, it performs t-Tests on selected attribute, comparing every group vs other for each subgroup and select the up-regulated genes as biomarker.
Computer biomarker dialog
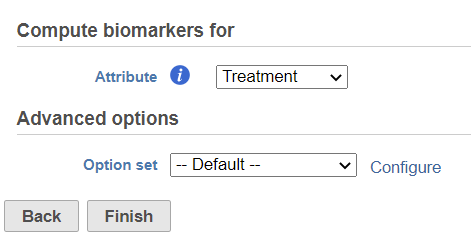
In the dialog, select attribute. The available attributes are categorical attribute can be seen on the data node which includes project level attribute and data node local annotation--e.g. graph-based cluster result (Figure 1). If the task is run on graph-based clustering output data node, the calculation is upstream data node which contains feature count–typically the input data node of PCA.
Click on the Configure of Advanced option to change the criteria of output features (Figure 2).
By default, the result outputs features that are up-regulated at least 1.5 (in linear scale) fold change for each subgroup comparing to others. The result is displayed in a table with each column is a subgroup name, each row is a feature.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
2 | rates |