Associating a Spreadsheet with an Annotation File
For Partek Genomics Suite to recognize an annotation spreadsheet, it must meet three requirements. First, there must be a column header row in the annotation file. Second, there must be a column in the annotation file that matches the probe identifiers in your data spreadsheet. Third, any text above the column header row must start with #.
We will illustrate annotation using imported .txt data files from an Illumina HumanHT-12 v4.0 Gene Expression BeadChip array and the HumanHT-12 v4.0 Whole-Genome Manifest File (TXT Format) from Illumina.
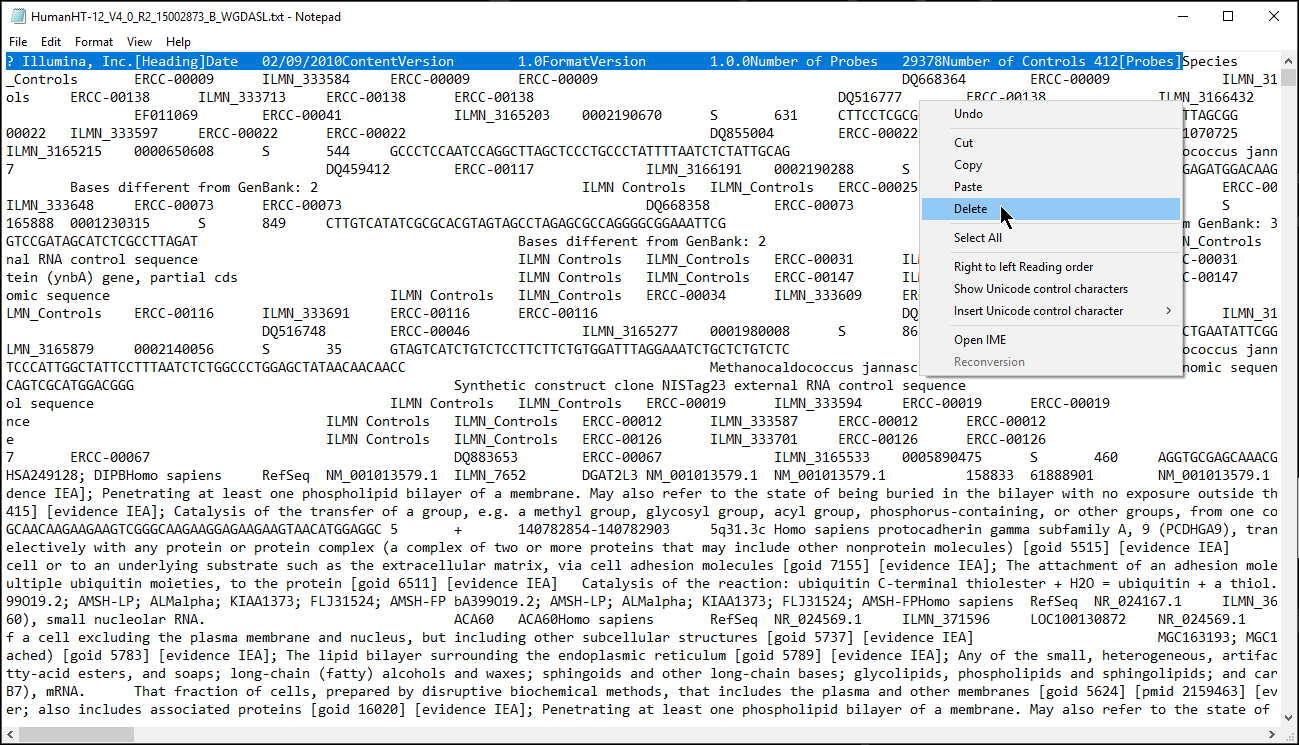
- Open the annotation file with a text editor such as Notepad
- Verify that a column in the annotation file matches the probe identifiers in your data spreadsheet
- Remove the text before the first column header (Figure 1) or add # to each text box and save the annotation file
If it is difficult to visualize where the column headers start, it might help to import the annotation file into a spreadsheet viewing program such as Microsoft Excel or Google Sheets.
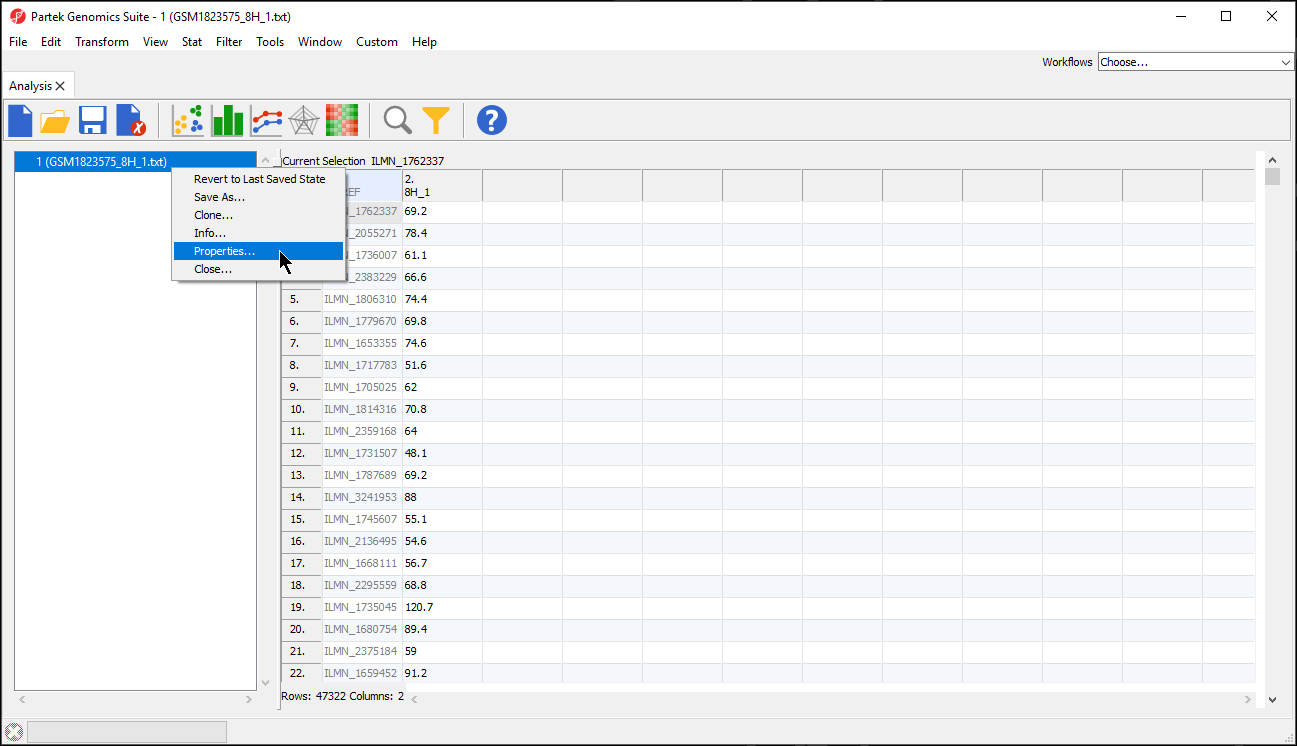
- Right click the spreadsheet you want to annotate in the spreadsheet tree panel
- Select Properties from the pop-up menu (Figure 2)
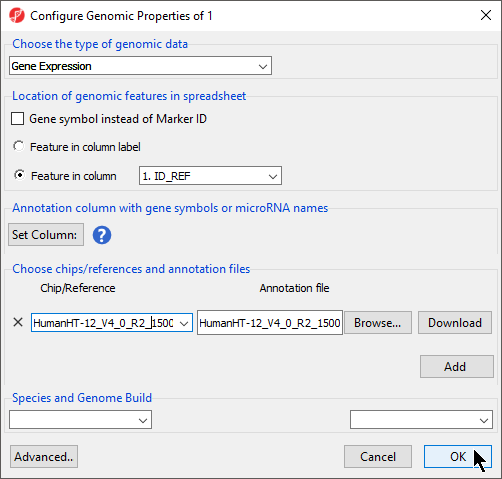
The Configure Genomic Properties dialog will now open. Select the appropriate option for Choose the type of genomic data; here we have chosen Gene Expression.
- Select Browse... from Choose chips/references and annotation files
- Select your annotation spreadsheet file using the file selection interface
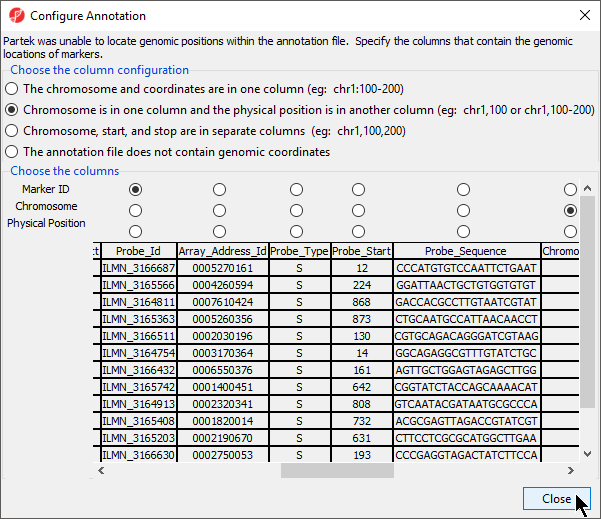
If the genomic position information from the annotation file cannot be automatically parsed, the Configure Annotation dialog will launch. This dialog allows you to choose which columns in the annotation file give the identity and genomic location of the features in your data spreadsheet. There are four options depending on if and how chromosome coordinates are described in the annotation file.
- Select Chromosome is in one column and the physical position is in another column (eg: chr1, 100 or chr1, 100-200) in our example
The Choose the columns section displays the annotation file spreadsheet with options to choose which columns are the Marker ID,Chromosome, and Physical Position (Figure 3).
- Select the column that matches the feature IDs in your data spreadsheet for Marker ID
We have chosen Probe_Id for Marker ID.
- Select the column(s) that matches the chromosome location data
We have chosen Chromosome for Chromosome and Probe_Coordinates for Physical Position.
- Select Close to return to the Configure Genomic Properties
The Chip/Reference text field will be populated with the annotation file name. You can edit this text field this if you wish.
- Select the species and genome from the Species and Genome Build section of the dialog; we have chosen Homo sapiens and hg19 (Figure 4)
- Select OK apply the annotation file to your data spreadsheet
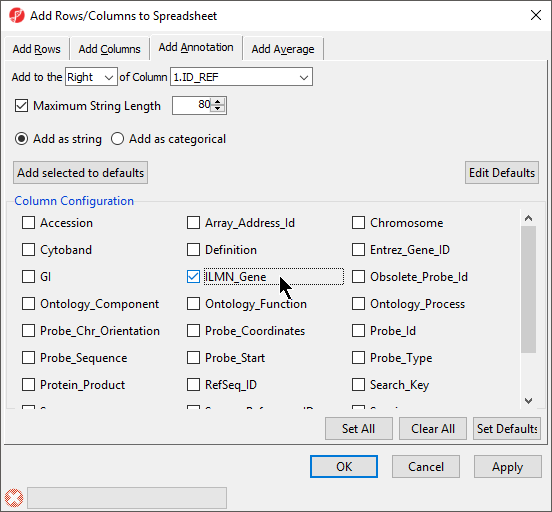
To verify that the annotation has been added, we can try to add annotation information to the spreadsheet.
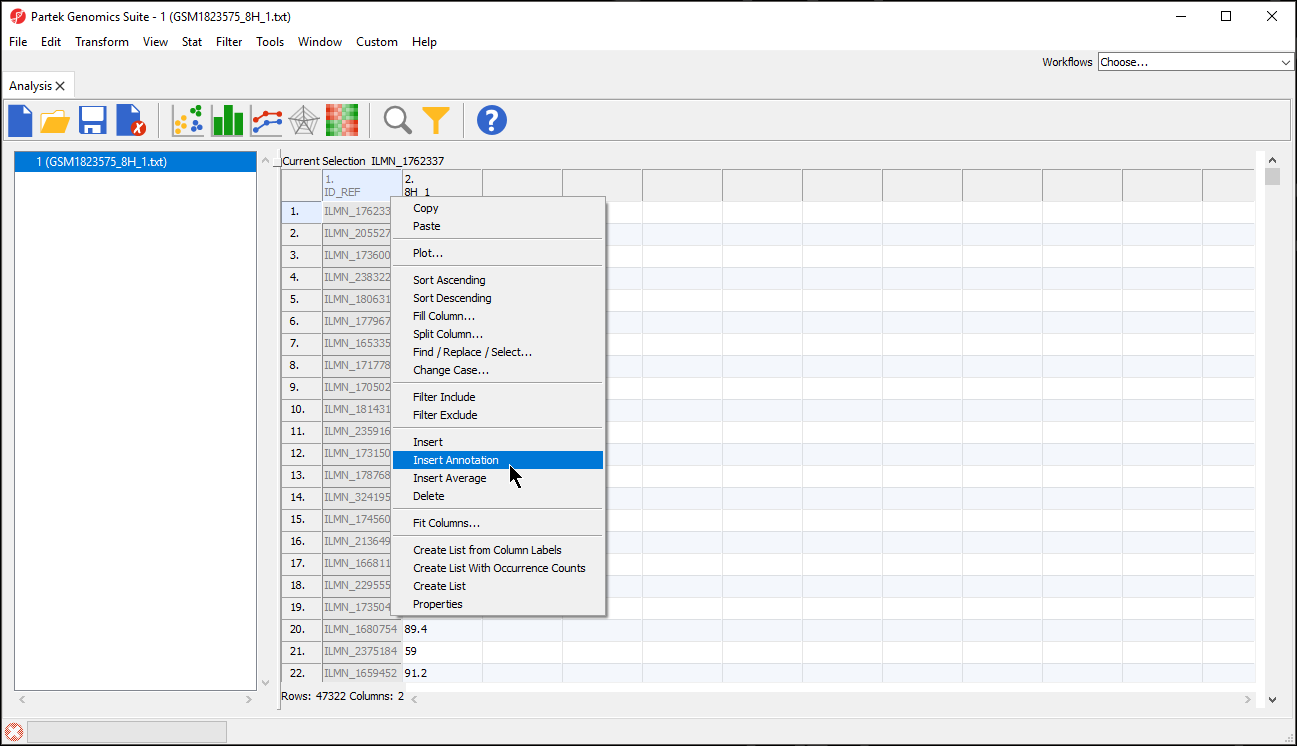
- Right-click on a column in the annotated data file spreadsheet
- Select Insert Annotation from the pop-up menu (Figure 5)
Building an Annotation File
If you are using an Illumina, Affymetrix, or an Agilent chip, you can obtain the appropriate annotation file from the chip manufacturer. If you have a custom chip or want to use a customized annotation file, you can create an annotation file that will allow you to add annotations to your data spreadsheet and invoke a Probe Set HTML report. Your annotation file must meet the following criteria:
- The annotation file must have a header (a line which contains a column label for each field)
- Any comments before the header must start with # or the header will not be recognized
- The fields of the annotation file must be tab or comma delimited
- A column in the annotation file must correspond to the feature ID column of your data spreadsheet
To invoke a genome view of your data, your annotation file must also have one or more columns that contain the genomic location in a format that Partek can recognize.
The annotation file must also contain a column that has the chromosome and base pair location (start and stop or physical position). Cytoband and/or strand can also be included. The table below provides possible column labels, a description of the format for that field, and an example.
Note: In this table the examples are for a gene on the top strand of chromosome 3; on the p arm in cytoband 14.2 starting at 69,871,322 base pairs and ending at 70,100,176
| Column label | Description of format | Example |
|---|---|---|
| chromosome | a chromosome label | 3 |
| start | an integer, the start position (in base pairs) of the feature | 69871322 |
| stop | an integer, the stop position (in base pairs) of the feature | 70100176 |
| genomic_coordinates | chromosome:start-stop | 3:69871322-70100176 |
| strand | + for top, - for bottom | + |
| physical position | an integer, the position (in base pairs) of the feature | 70100176 |
Here are a few examples of the first two rows of annotation files:
- Using Agilent format
| ProbeID | GeneName | GenomicCoordinates | Cytoband |
|---|---|---|---|
| A_44_P1025812 | TC521361 | chr12:2546883-2546824 | rn|12p12 |
- Using Affymetrix SNPs format
| Probe Set ID | Chromosome | Physical Position | Strand | Cytoband |
|---|---|---|---|---|
| SNP_A-1512540 | 9 | 22205296 | - | p21.3 |
- Using Affymetrix exons format
| probeset_id | seqname | strand | start | stop |
|---|---|---|---|---|
| 2315588 | chr1 | + | 1155398 | 1155624 |
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |