To detect differential methylation between CpG loci in different experimental groups, we can perform an ANOVA test. For this tutorial, we will perform a simple one-way ANOVA to compare naive-state hPSCs treated with shRNAs targeting NANOG to those targeting POU5F1.
- Select Defect Differential Methylation from the Analysis section of the Illumina BeadArray Methylation workflow
- Select 3. shRNA treatment from the Experimental Factor(s) panel
- Select Add Factor > to move 3. shRNA treatment to the ANOVA Factor(s) panel (Figure 1)
- Select Contrasts...
- Select Yes for Data is already log transformed? because M-values are based on logit transformation
- Select shPOU5F1
- Select Add Contrast Level > for Group 1 to add shPOU5F1 to Group 1, which will be renamed shPOU5F1
- Repeat steps to add shNANOG to Group 2
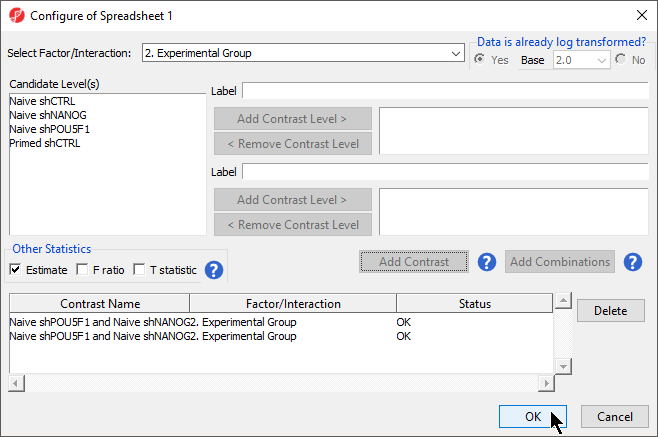
- Select the Estimate box in the Other Statistics section of the Configure dialog (Figure 2)
By default, the fold-change value for each contrast will be calculated. Selecting Estimate will also include the difference in methylation levels between the groups at each CpG site in the output. These values will be needed later in the tutorial when we filter the differentially methylated loci.
- Select Add Combination
- Select OK to close the Configuration dialog
- Select OK to close the ANOVA dialog and run the ANOVA
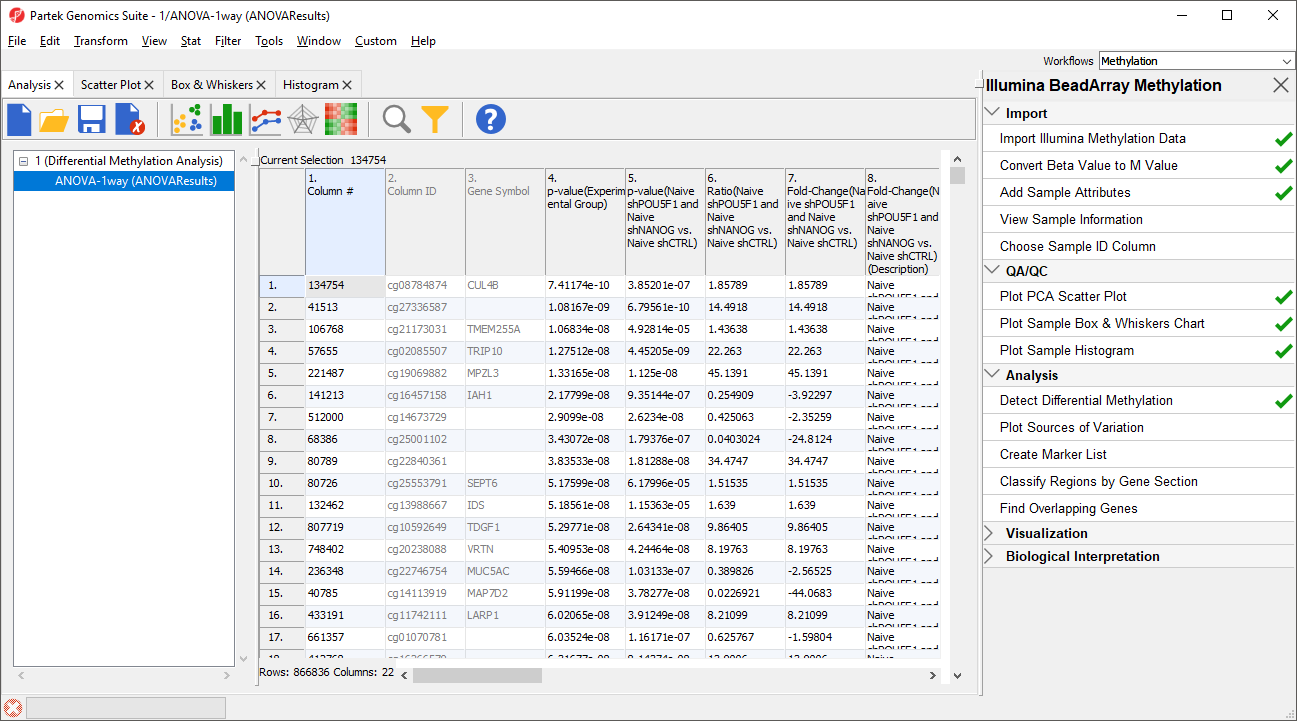
The results will appear as ANOVA-1way (ANOVAResults), a child spreadsheet of 1 (Differential Methylation Analysis). Each row of the spreadsheet represents a single CpG locus (identified by Column ID).
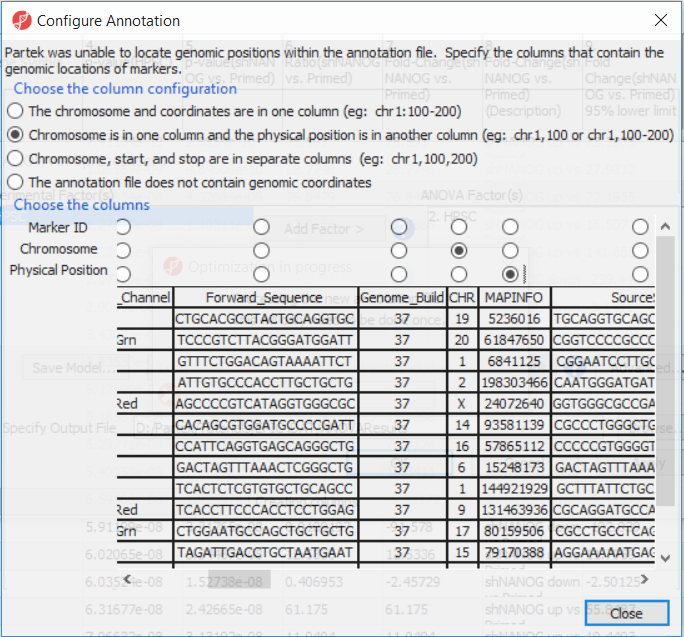
The first time you use MethylationEPIC array, the manifest file needs to be configured and the window like the one in Figure 4 will pop up. First select the second option (Chromosome is in one column and the physical location is in another column). Marker ID is on the first column (Ilmn ID), Chromosome is on the column CHR, while the Physical Position is on the MAPINFO column. Set according to Figure 4 and select Close. This enable Partek Genomics Suite to parse out probe annotation from the manifest file.
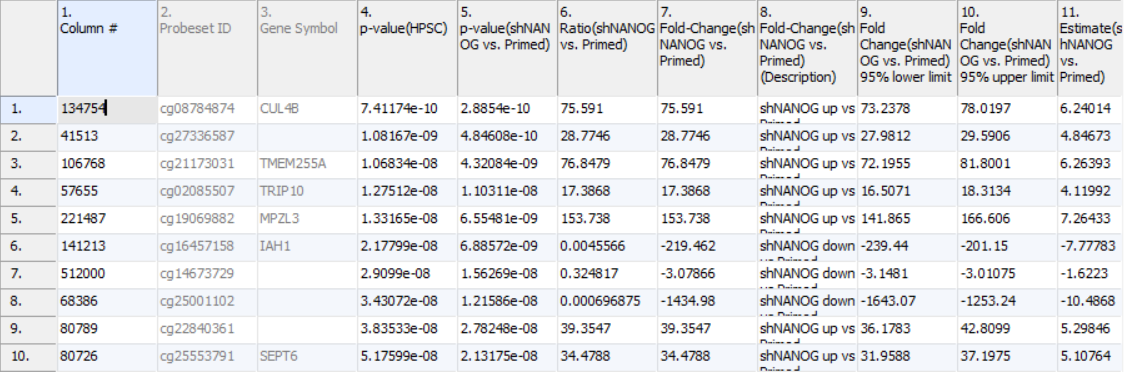
The result of 1-way ANOVA is shown in Figure 5. Each row of the table represents a single CpG locus (identified by Probeset ID column). The remaining columns contain the following information:
Column 3. Gene Symbol: the gene overlapping the probe as specified in the Illumina manifest file
Column 4. p-value(HPSC): overall p-value for the specified factor (in parenthesis). A low p-value indicates that there is a difference in methylation between the levels of this attribute (i.e. study groups). The contrast p-values should then be used to evaluate individual group comparisons. If more than one factor is included in the model, p-value will be reported for each.
Next, for each contrast included in the model, a block of seven columns will be added, as follows:
Column 5. p-value(shNANOG vs. Primed): p-value for the given contrast (in parenthesis). A low p-value indicates a difference in methylation between the groups included in the contrast (here: shNANOG and Primed).
Column 6. Ratio(shNANOG vs. Primed): ratio of average methylation level in one over the other the other contrasted group (shNANOG and Primed, respectively). Ratio is reported in linear space.
Column 7. Fold Change(shNANOG vs. Primed): fold-change in one over the other contrasted group (shNANOG and Primed, respectively). Fold-change is reported in linear space.
Column 8. Fold Change(shNANOG vs. Primed) (Description): if fold-change > 1, it means hypermethylation in the first group (e.g. shNANOG up vs Primed), if fold-change < -1, it means hypomethylation in the first group (e.g. shNANOG down vs Primed), relative to the second group (Primed). This column enables quick filtering
Columns 9. & 10. Lower and upper (respectively) limits of 95% confidence interval of the fold-change
Column 11. Estimate(shNANOG vs. Primed): difference between means of two groups (i.e. shNANOG and Primed) (this column is optional and depends on the way contrasts were set up)
Columns 12. - 18. correspond to columns 5. - 11.
Columns 19.+ Statistical output
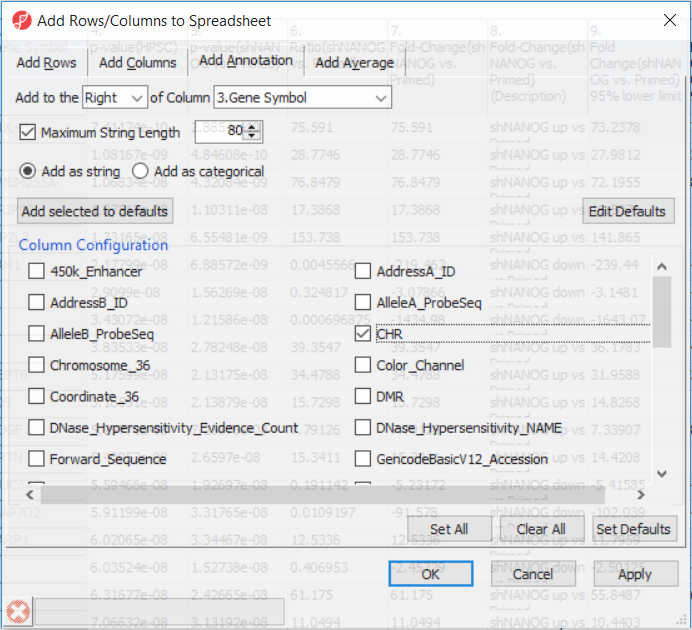
Going forward, analysis of differentially methylated loci typically includes removal of the probes on X and Y chromosomes (to avoid the problems with inactivation of one X chromosome). To annotate the ANOVA spreadsheet with the information required for filtering, right-click on the Gene Symbol column, select Insert Annotation, tick-mark the CHR filed (Figure 6) and push OK. A new column will be appended to the spreadsheet.
To enable filtering right-click on the header of the CHR column > Properties and set the type to categorical (and OK). Now, activate the Interactive Filter tool () . If needed use the drop-down list to point to the CHR column. The column chart represents the number of appearances of each chromosome in the spreadsheet (i.e. the number of probes per chromosome). To remove the probes on the X and the Y chromosome left click on the two right-most columns (the pop up balloon will show you the chromosome label) and the columns will be grayed out (Figure 7).
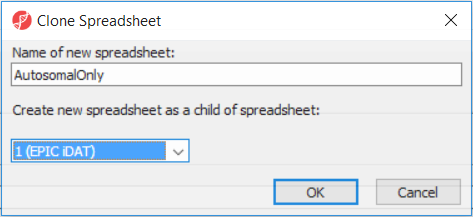
After removal of the probes on the sex chromosomes, let us extract all the autosomal probes to a new spreadsheet. Right click on the ANOVA 1-way spreadsheet > Clone... Set the Name of new spreadsheet to AutosomalOnly and make it a child of the top-level spreadsheet (EPIC iDAT) (Figure 8). Push OK. The newly created autosomalonly spreadsheet will be a new starting point for all the downstream steps.
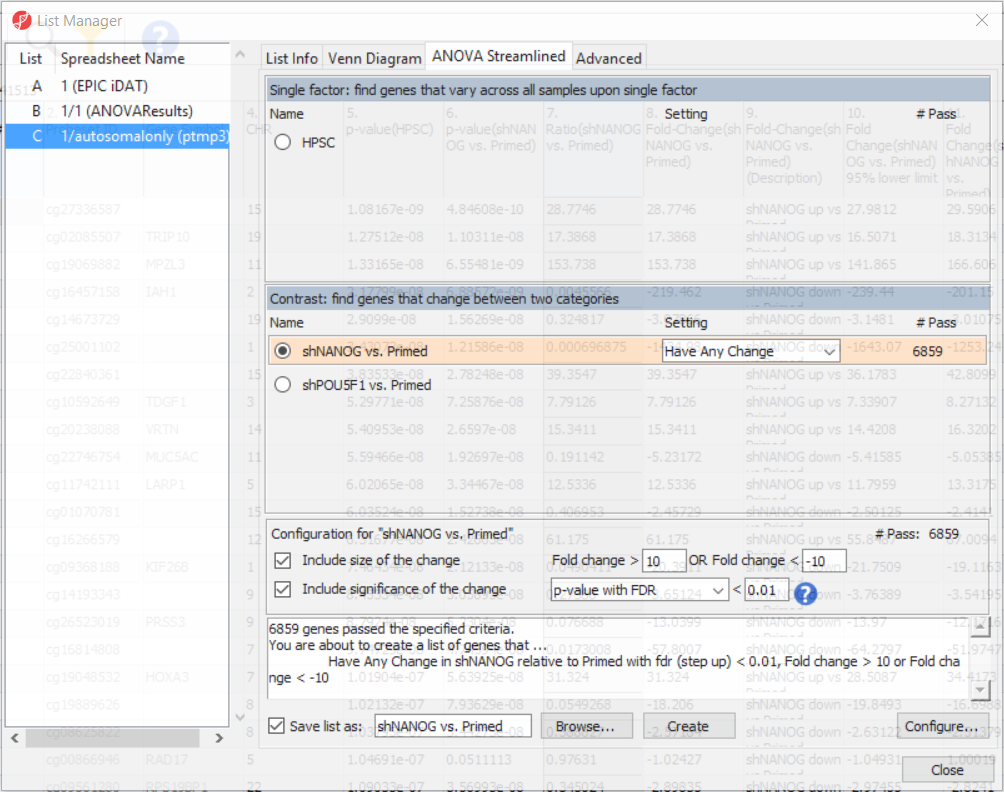
To come up with a list of differentially methylated loci, proceed to the workflow and select Create Marker List. That will open the List Manager functionality and you will be on the ANOVA Streamlined tab. Factors in the model are listed in the top section, contrasts specified in the model are in the middle, while the filter settings are at the bottom.
Select shNANOG vs Primed contrast to see the current filter configuration. By default, both fold-change and p-value with false discovery rate (FDR) correction are applied, with the number of CpG loci passing the filter given as # Pass. For this tutorial, set the fold-change to > 10 and < -10, and reduce the p-value with FDR down to 0.01 (Figure 9). Then push the Create button to save the list of significant loci under the default name (shNANOG vs. Primed). Repeat the procedure for the shPOU5F1 vs Primed, using the same cut offs.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |