SC transform task performs the variance stabilizing normalization proposed in [1]. The task's interface follows that of SCTransform() function in R [2].
We recommend performing the normalization on a single cell raw count data node. Select SCTransfrom task in Normalization and scaling section on the pop-up menu to invoke the dialog (Figure 1).
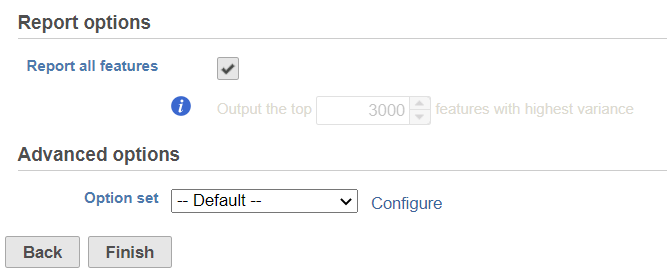
By default, it will generate report on all the input features. Unchecking the Report all features, user can limit the results to a certain number of features with highest variance.
In Advanced options, users can the click Configure to change the default settings (Figure 2).
Features for parameter estimation: Specify number of features to use in estimation of parameters; 0 means to use all input features.
Center results: When set to Yes, center all the transformed features to have zero mean expression.
Clip results: If this is set to No, outliers might have big effect and the transformed data can be very large for some features, usually the ones with few non-zero counts. When set to Yes, the range to clip the transformed data is between -sqrt(n/30) and sqrt(n/30), where n is the number of cells.
Random seed: use the same random seed to reproduce the results.
Data has been log transformed with base: specify the input data is logged or not.
The data in the output node is a matrix of normalized values (residuals) that by default has the same size as the input data set. The range of normalized values is roughly between -4 and 4.
References
- Christoph Hafemeister, Rahul Satija Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. https://doi.org/10.1101/576827
- SCTransform() documentation https://www.rdocumentation.org/packages/Seurat/versions/3.1.4/topics/SCTransform
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |