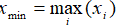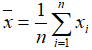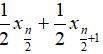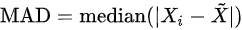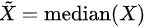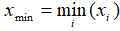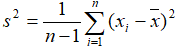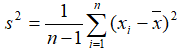Descriptive statistics task can be invoked on matrix data node e.g. gene counts, normalized counts data node in bulk RNA seq analysis pipeline or single cell counts data node etc. It calculates measures of central tendency and variability on observations or features of the matrix data.
Running Descriptive statistics
- Click on a counts data node
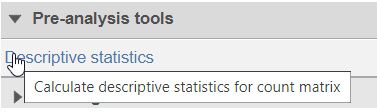
- Choose Descriptive Statistics in Pre-analysis tools section of the toolbox (Figure 1)
Figure 1. Descriptive statistics menu
This will invoke the dialog (Figure 2), select the calculation will be performed on samples (or cells on single cell count data node) or features
Figure 2. Select to calculate descriptive statistics on samples or cells
The available statistics are listed on the left panel, suppose "x1, x2, ..., xn"represent an array of numbers
- Coefficient of variation (CV): s represent the standard deviation
- Geometric mean: g=
- Max:
- Mean:
- Median: when n is odd, median is , when n is even, median is
- Median absolute deviation: , where
- Min:
- Non zero count: number of observations that is not zero
- Q1: 25th percentile
- Q3: 75th percentile
- Range: xmax - x min
- Standard deviation: where
- Sum:
- Variance:
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |
Overview
Content Tools