Join us for an event September 26!
How to Streamline RNA-Seq analysis and increase productivity—point, click, and done
Introduction
The Choose taxonomic level task allows you to generate a count matrix summarizing the number of reads that have been classified by Kraken for each taxon, at a given taxonomic level. The counts give a measure of the relative abundance of each taxon, which can be used for downstream analysis and visualization as if it were RNA-Seq count data.
Running the Choose Taxonomic Level Task
The task can be performed on a Taxonomic data node, which is the output from a Kraken task.
- Click a Taxonomic data node
- Choose Choose taxonomic level from the Metagenomic section of the toolbox
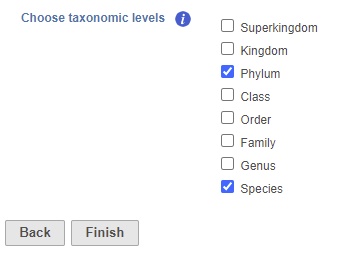
- Check one or more taxonomic levels. The options are Superkingdom, Kingdom, Phylum, Class, Order, Family, Genus, or Species (Figure 1). A separate output data node will be generated for each one that is selected
- Click Finish
Figure 1. Choose taxonomic level task set up page. Check one or more boxes
Download a count matrix
Downstream Analysis
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
0 | rates |
Overview
Content Tools