| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
Importing single cell data
Partek Flow supports the import of filtered gene-barcode matrices generated by 10x Genomics' Cell Ranger pipeline. The following document will illustrate how to perform this import.
| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| minLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
Below is a video summarizing the import of these files:
...
Configure all the relevant sample metadata, including sample name and the annotation that was used to generate the matrices, and click Finish when completed. Note that all matrices must have been generated using the same reference genome and annotation to be imported into the same project.
Importing spatial data
Importing Xenium Output Bundle
Raw output data generated by the 10x Genomics' Xenium Onboard Analysis pipeline consists of decoded transcript counts and morphology images. The raw output and other standard output files derived from them are compiled into a zipped file called Xenium Output Bundle.
To import the Xenium Output Bundle into Partek Flow, create a new project and click Add data, then select Import 10x Genomics Xenium under Single cell > Spatial, click Next.
| Numbered figure captions |
|---|
| SubtitleText | Importing Xenium spatial data |
|---|
| AnchorName | Import_xenium |
|---|
|
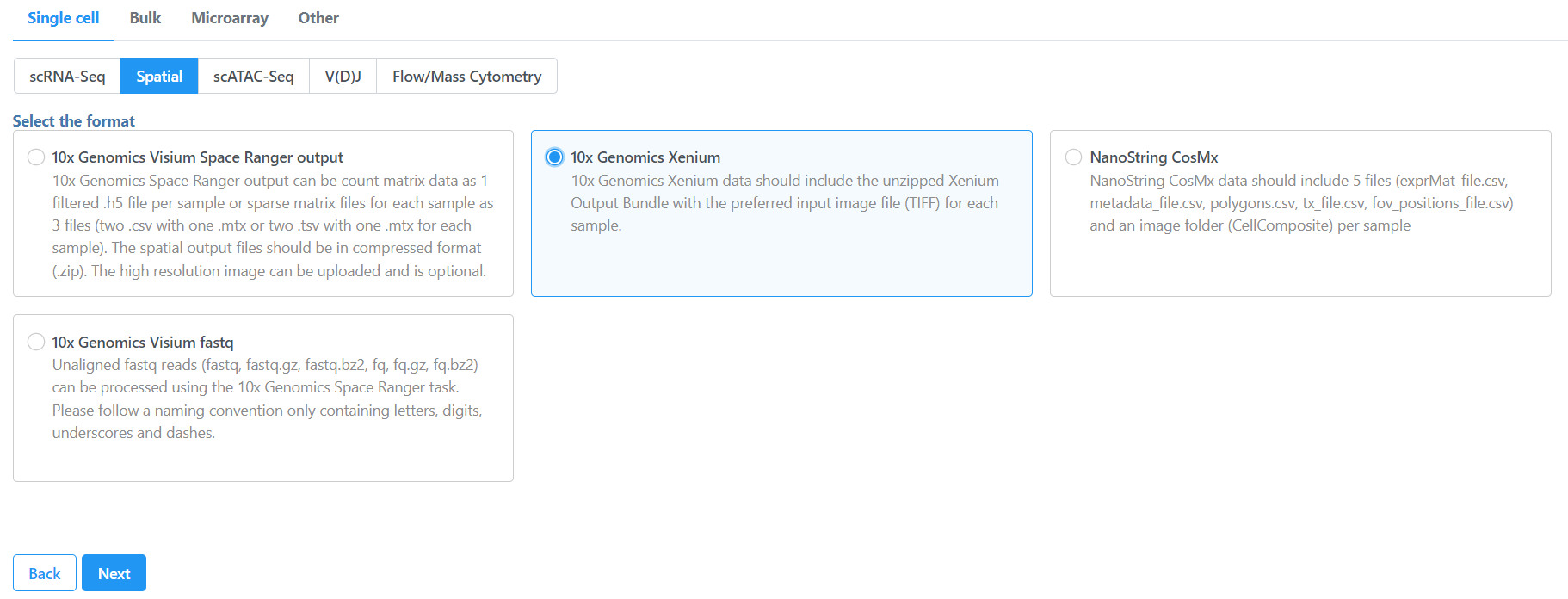 Image Added Image Added
|
Samples can be added using the Add sample button. Each sample should be given a name and a folder containing the required 6 files: cell_feature_matrix.h5, cells.csv.gz, cell_boundaries.csv.gz, nucleus_boundaries.csv.gz, transcripts.csv.gz, morphology_focus.ome.tif should be uploaded per sample using the Browse button. The required 6 files should be all included in the Xenium Output Bundle folder.
| Numbered figure captions |
|---|
| SubtitleText | Import Xenium Output Bundle folder for each sample |
|---|
| AnchorName | Import_Xenium output bundle folder |
|---|
|
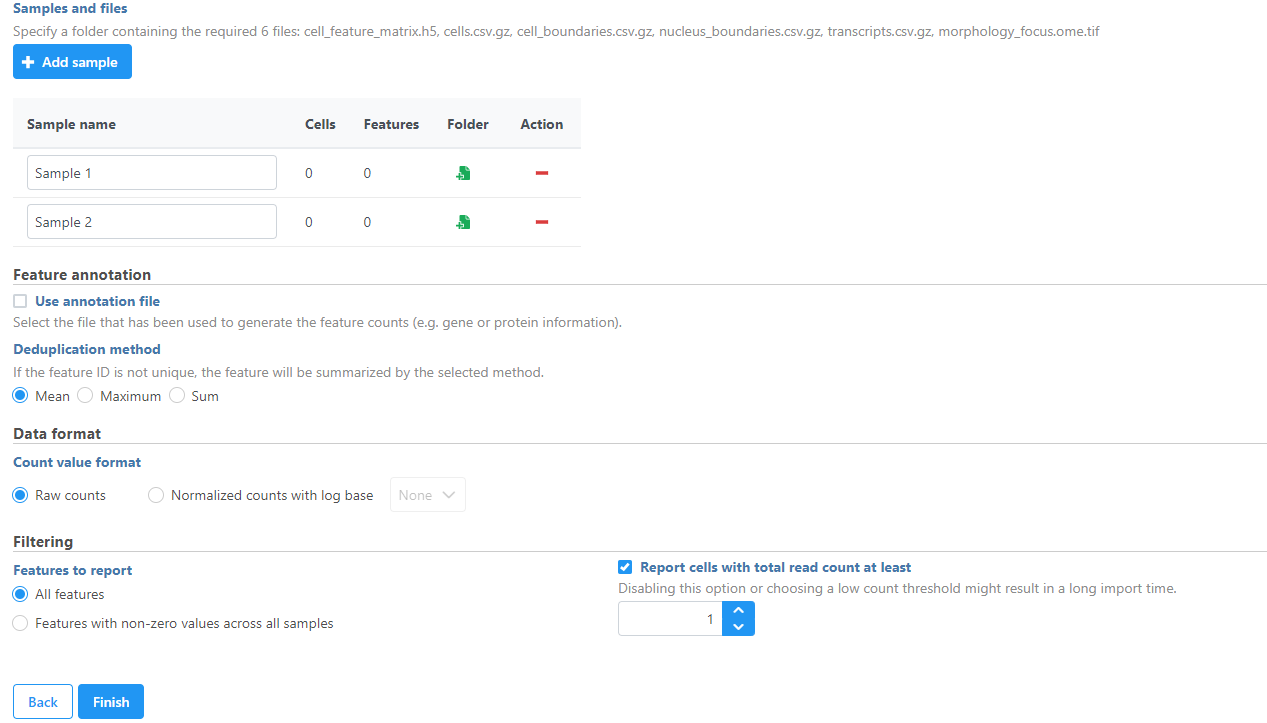 Image Added Image Added
|
If you have not already, transfer the files to the server to be accessed when you click Browse. Follow the directions here to add files to the server. You will need to decompress the Xenium Output Bundle zip file before they are uploaded to the server. After decompression, you can drag and drop the entire folder into the Transfer files dialog, all individual files in the folder will be listed in the Transfer files dialog after drag & drop, with no folder structure. The folder structure will be restored after upload is completed.
| Numbered figure captions |
|---|
| SubtitleText | Drag & drop unzipped Xenium Output Bundle folder into Transfer files dialog |
|---|
| AnchorName | Drag_and_drop_Xenium_output_bundle_folder |
|---|
|
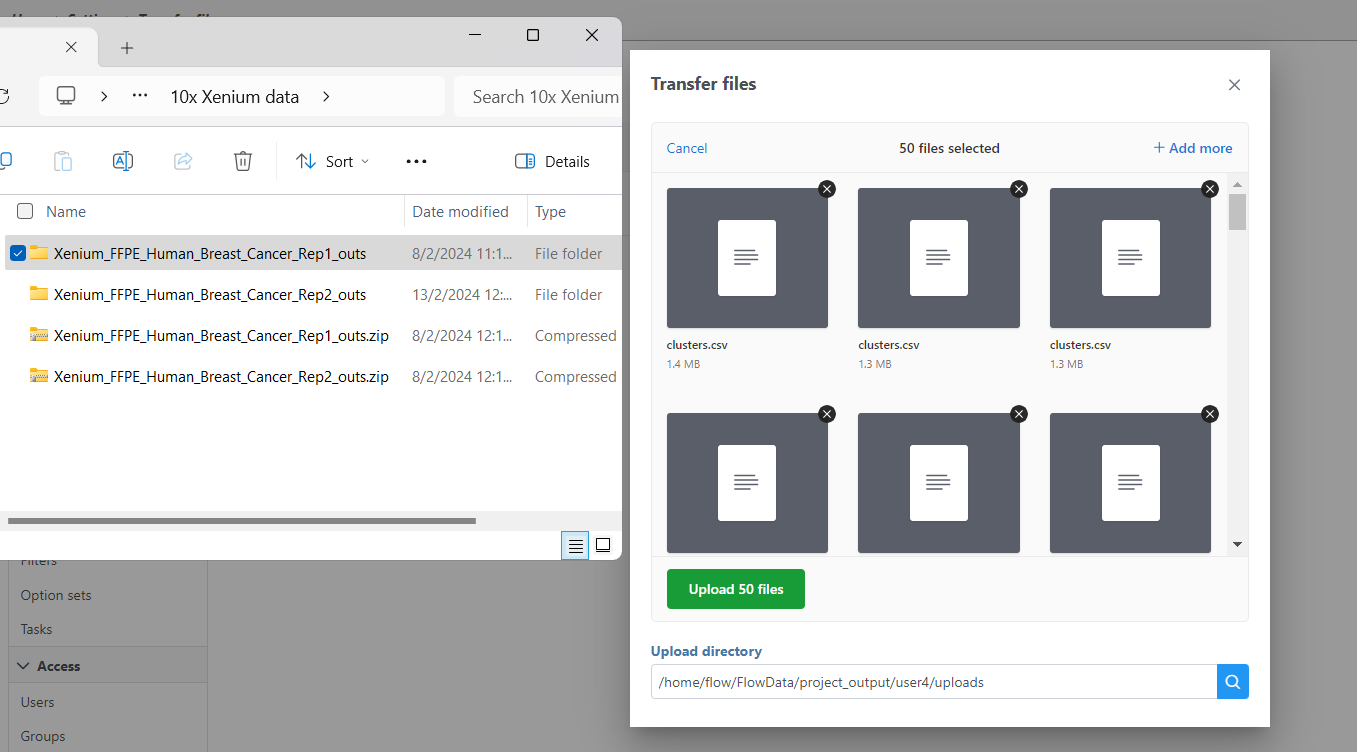 Image Added Image Added
|
Once you have uploaded the folder into the server, you can continue to select the folder for each sample from Browse. Once the folder is selected, the Cells and Features values will auto-populate. You can choose an annotation file that matches what was used to generate the feature count. Then, click Finish to start importing the data into your project.
| Numbered figure captions |
|---|
| SubtitleText | Add Xenium Output Bundle and select annotation |
|---|
| AnchorName | Add_Xenium_output_bundle_folder |
|---|
|
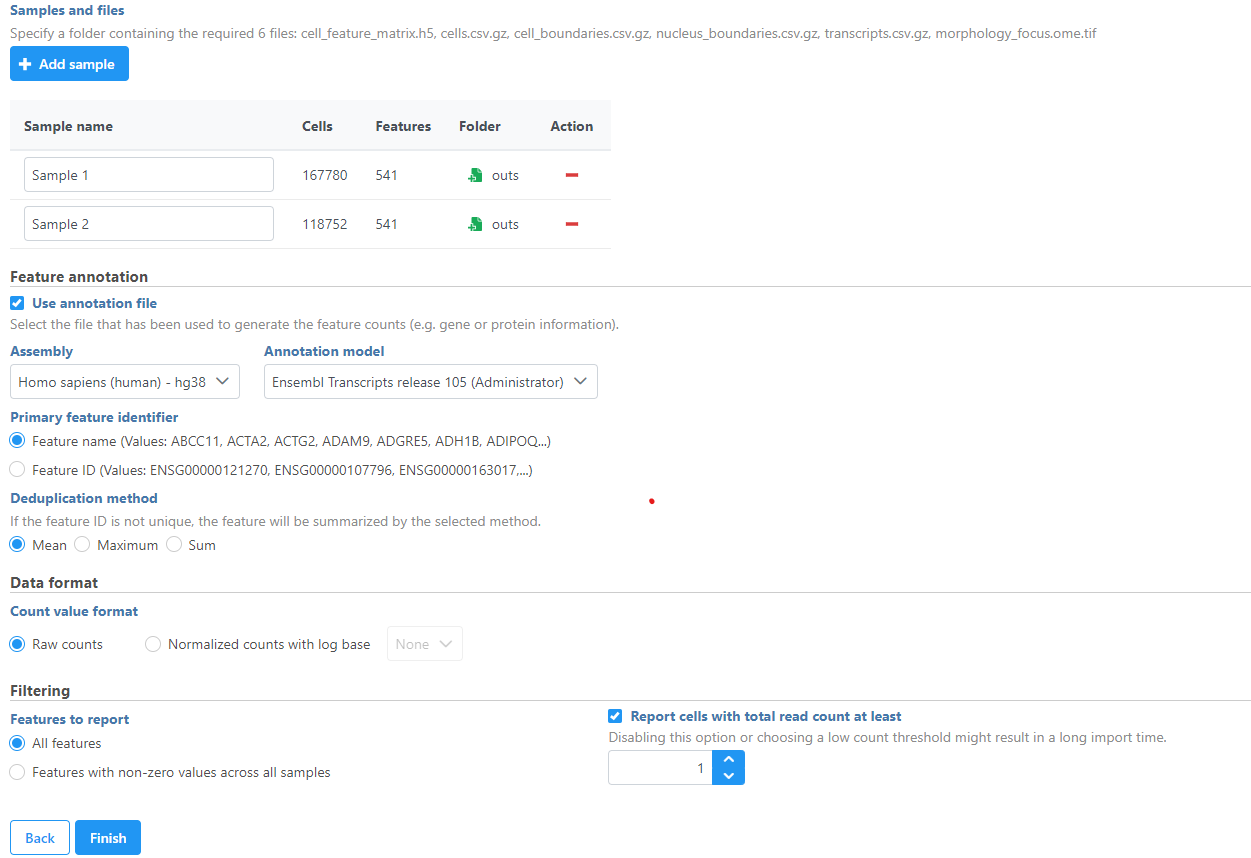 Image Added Image Added
|
...



