...
The Choose taxonomic level task allows you to generate task generates a count matrix summarizing the number of reads that have been classified by Kraken for each taxon, at a given taxonomic level. The counts give a measure of the relative abundance of each taxon, which can be used for downstream analysis and visualization as if it were RNA-Seq count data.
...
- Click a Taxonomic data node
- Choose Choose taxonomic level from the Metagenomic section of the toolbox
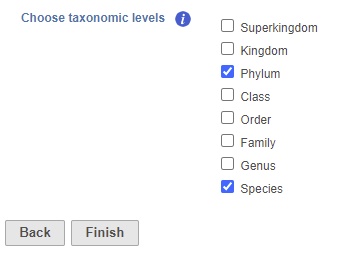
- Check one or more taxonomic levels. The options are Superkingdom, Kingdom, Phylum, Class, Order, Family, Genus, or Species (Figure 1). A separate output data node will be generated for each one that is selected selected (Figure 2)
- Click Finish
| Numbered figure captions |
|---|
| SubtitleText | Choose taxonomic level task set up page. Check one or more boxes |
|---|
| AnchorName | Choose taxonomic level task set up |
|---|
|
|
| Numbered figure captions |
|---|
| SubtitleText | One output data node is produced for each taxonomic level chosen |
|---|
| AnchorName | Choose taxonomic level output |
|---|
|
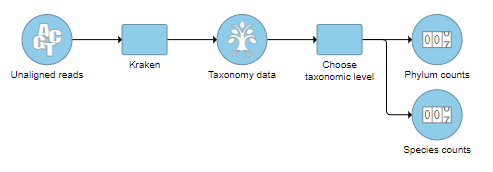 Image Added Image Added
|
Download a count matrix
To export the count matrix for a taxonomic level, select the output data node and choose Download data from the toolbox. You can choose to put the features on the columns or rows (Figure 3). The 'features' in this context are the taxa. For example, if a Phylum data node is downloaded, the features will be different phyla. The download will be a tab-delimited text file with read counts for each sample (Figure 4).
| Numbered figure captions |
|---|
| SubtitleText | Download the matrix of read counts for each taxon per sample |
|---|
| AnchorName | Download count matrix |
|---|
|
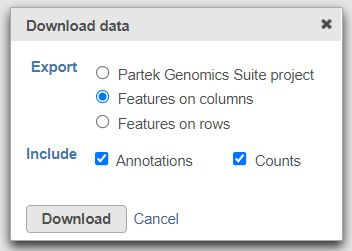 Image Added Image Added
|
| Numbered figure captions |
|---|
| SubtitleText | Example of Phylum-level count matrix with features (phyla) on columns. Column 1 is the sample name. Columns 2 & 3 are sample attributes. Columns 4+ are different phyla. The counts re the number of reads that have been classified for each phylum, for each sample |
|---|
| AnchorName | Phylum-level count matrix |
|---|
|
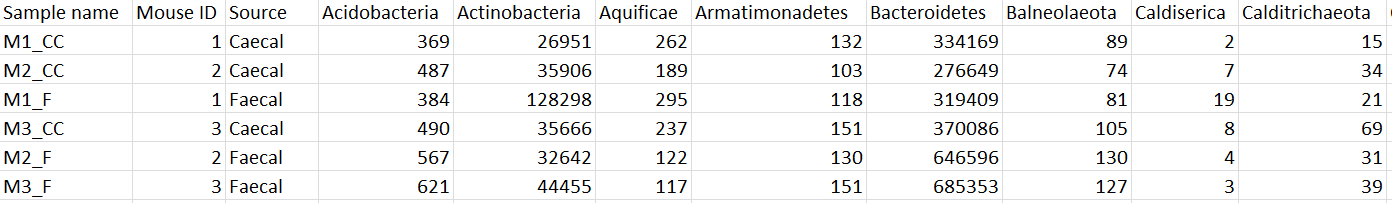 Image Added Image Added
|
Downstream Analysis
...



