Page History
...
Comparing gene expression between cell types
A common goal in single cell analysis is to identify genes that distinguish a cell type. To do this, you can use the differential analysis tools in Partek Flow. I will show how to use the Gene Specific Analysis (GSA) test in Partek Flow, which on its default settings is equivalent to limma-trend, a statistical test has been shown to be highly effective for differential analysis of single cell RNA-Seq data (REFERENCE).
- Click the Classified groups data node
- Click Differential analysis in the toolbox
- Click GSA
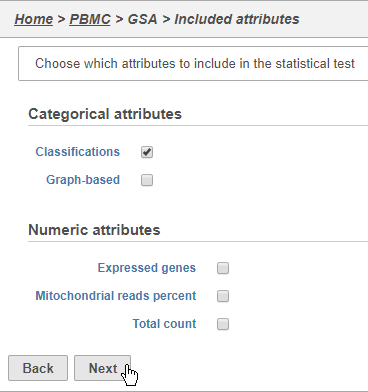
The first page of the configuration dialog asks what attributes you want to include in the statistical test. Here, we only want to consider the Classifications, but in a more complex experiment, you could also include experimental conditions applied to different samples.
- Click Classifications
- Click Next (Figure 30)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
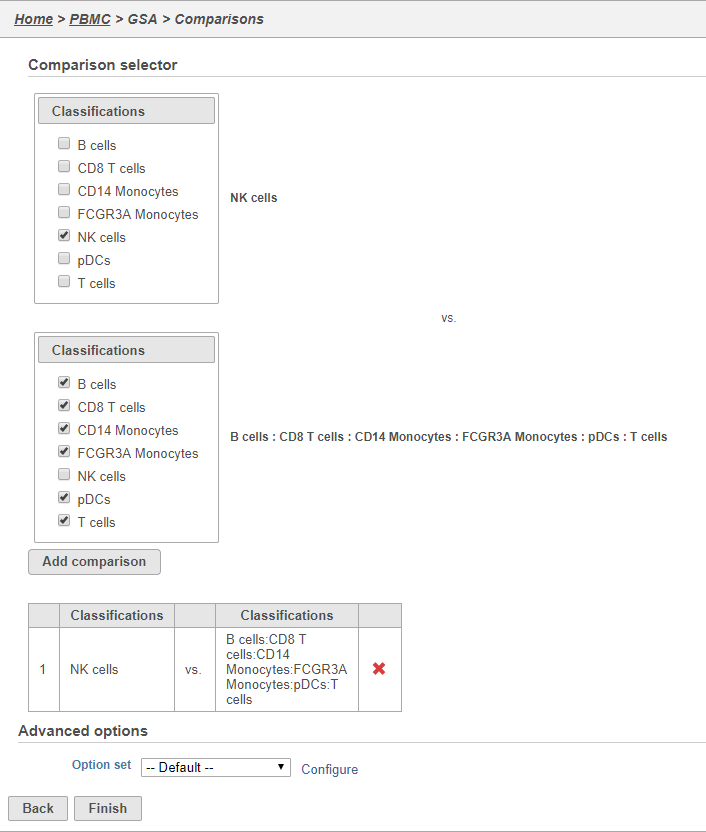
Here, we will make a comparison between NK cells and all the other cell types to identify genes that distinguish NK cells. However, you could also use this tool to identify genes that differ between two cell types or genes that differ in the same cell type between experimental conditions.
- Click NK cells in the top panel
The top panel is the numerator for fold-change calculations so the experimental or test group should be selected in the top panel.
- Click all the other classifications in the bottom panel
The bottom panel is the denominator for fold-change calculations so the control group should be selected in the bottom panel.
- Click Add comparison
This adds the comparison to the statistical test.
- Click Finish to run the GSA task (Figure 32)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Double-click the Feature list data node to open the GSA task report
| Additional assistance |
|---|
|
| Rate Macro | ||
|---|---|---|
|