Partek Flow supports the import of filtered gene-barcode matrices generated by 10X Genomics' Cell Ranger pipeline. The following document will illustrate how to perform this import.
| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| minLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
Below is a video summarizing the import of these files:
| Multimedia |
|---|
| name | 10X_matrix_import.mp4 |
|---|
| width | 80% |
|---|
|
| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| minLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
Importing matrices into Partek Flow
To import the matrices into Partek Flow, create a new project and click Import data>Import single cell data.
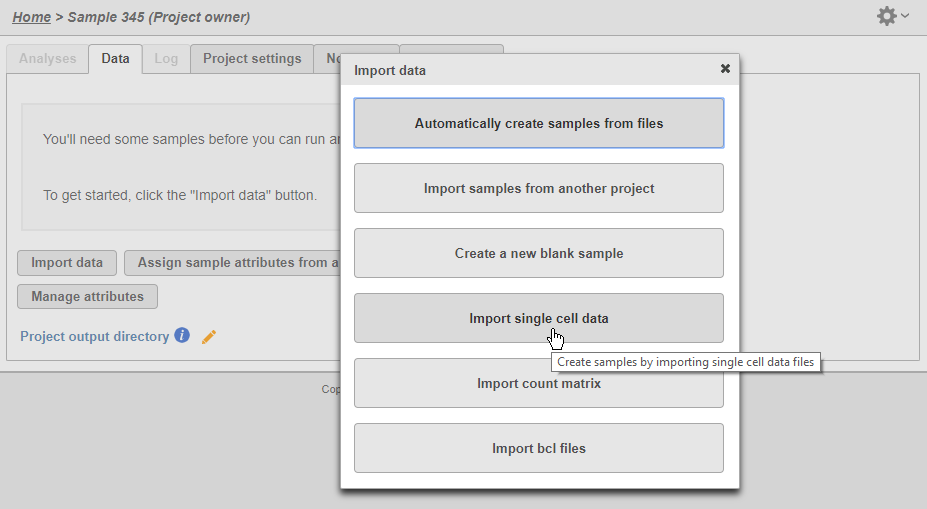 Image Removed
Image Removed
| Numbered figure captions |
|---|
| SubtitleText | Importing single cell data |
|---|
| AnchorName | Import_single_cell_data |
|---|
|
 Image Added Image Added
|
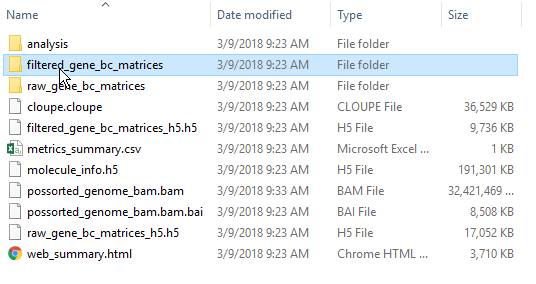
Navigate to the folder containing your filtered matrices. By default, the Cell Ranger pipeline output will have a folder called filtered_gene_bc_matrices.
| Numbered figure captions |
|---|
| SubtitleText | Filtered matrix folder from Cell Ranger pipeline |
|---|
| AnchorName | filtered_gene_bc_matrices |
|---|
|
|
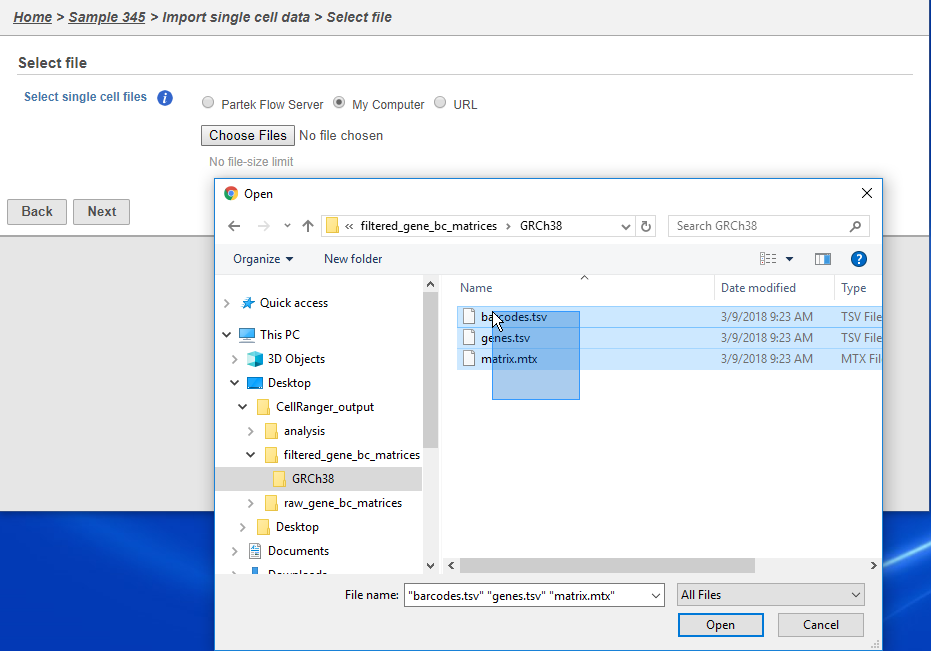
There are folders nested within the matrix folder, typically representing the reference genome it was aligned to. Navigate to the lowest subfolder, this should contain three files:
...
Select all 3 files for importing into Partek Flow and click Next
| Numbered figure captions |
|---|
| SubtitleText | Select all 3 files for importing |
|---|
| AnchorName | Select_filtered_gene_bc_matrices |
|---|
|
|
Specify the annotation file used when running the pipeline. If desired, the sample name can also be modified.
| Numbered figure captions |
|---|
| SubtitleText | Configuring matrix metadata |
|---|
| AnchorName | Configure_SC_matrix_metadata |
|---|
|

|
Click Finish when you have completed configuration.This will queue the import task.
...
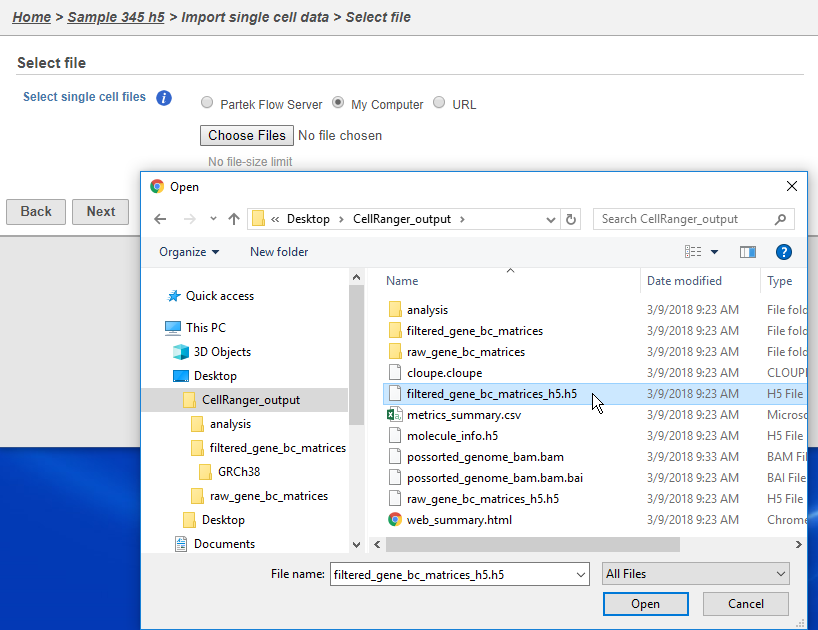
The Cell Ranger pipeline can also generate the same filtered gene barcode matrix in h5 format. This gives you the ability to select just one file per matrix. To import an h5 matrix, simple select the h5 file when selecting a file for import and click Next. This will take you to the same configuration page listed above.
| Numbered figure captions |
|---|
| SubtitleText | Importing matrix in h5 format |
|---|
| AnchorName | H5_matrix_import |
|---|
|
|
This feature is also useful for importing multiple samples in batch. Simply put all h5 files from your experiment on a single folder, navigate to that and select all the matrices you would like to import.
| Numbered figure captions |
|---|
| SubtitleText | Import multiple matrices in batch by selecting all h5 files |
|---|
| AnchorName | Select_multiple_h5 |
|---|
|

|
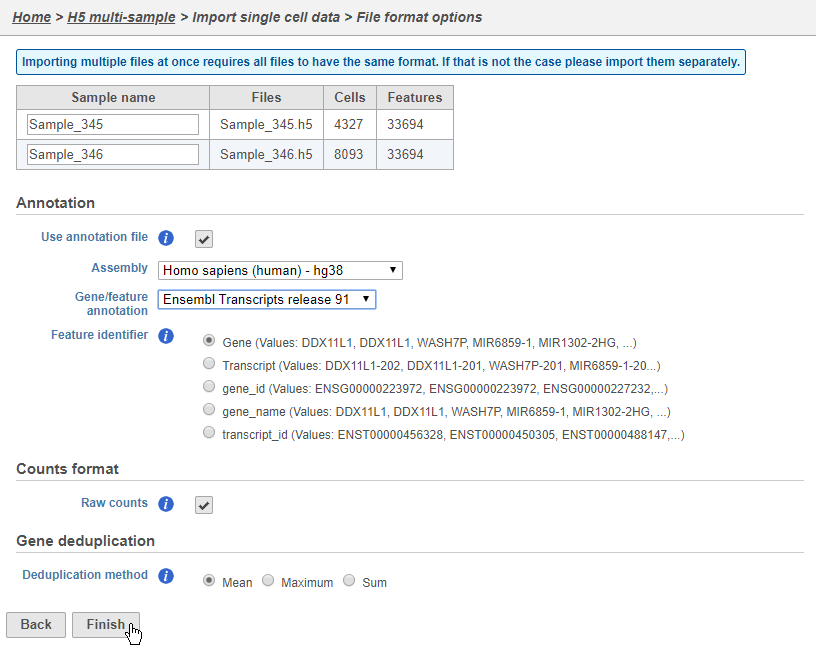
Configure all the relevant sample metadata, including sample name and the annotation that was used to generate the matrices, click Finish when completed. Note that all matrices must have been generated using the same reference genome and annotation to be imported into the same project.
| Numbered figure captions |
|---|
| SubtitleText | Configure matrix information for all h5 matrices to be imported |
|---|
| AnchorName | Configure_multiple_h5_import |
|---|
|
|
...





