Page History
...
- Select Create Marker List from the Analysis section of the Illumina BeadArray Methylation workflow
- Select 1/anova-1way_autosomal _only (ANOVA-1way_autosomal_only) as the source spreadsheet from the left-hand panel
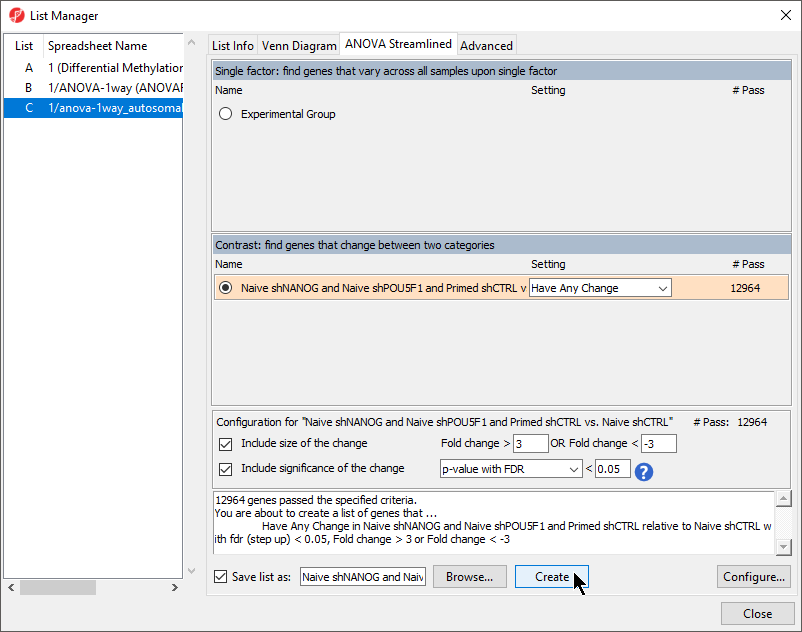
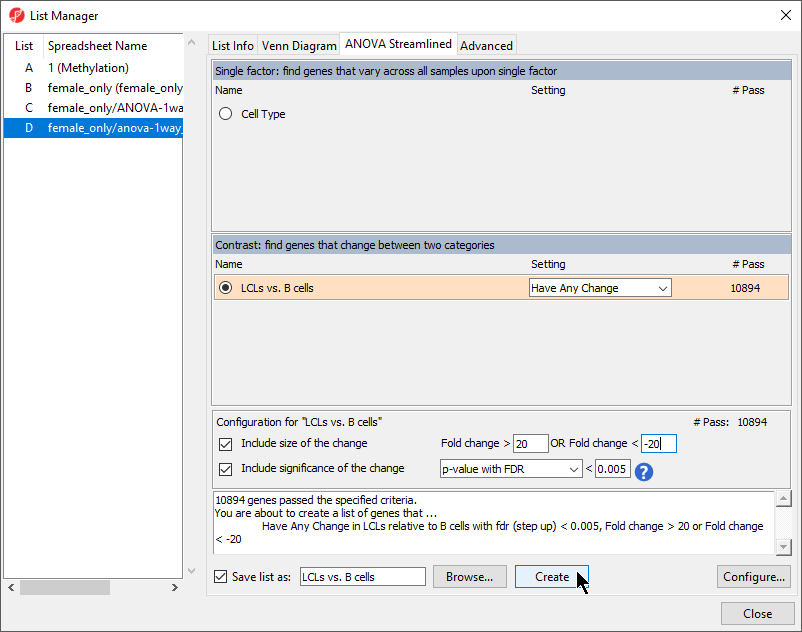
- Select Naive shPOU5F1 and Naive shNANOG and Primed shCTRL LCLs vs. Naive shCTRLB cells from the Contrast: find genes that change between two categories section of the ANOVA Streamlined tab of the List Manager dialog
- Set Fold change > to 3 20
- Set OR Fold change < to -320
- Set p-value with FDR to 0.005
- Set file name as CpG of interestLCLs vs. B cells
- Leave the rest of the option set to defaults (Figure 1)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select Create
- Select Close to exit the List Manager dialog
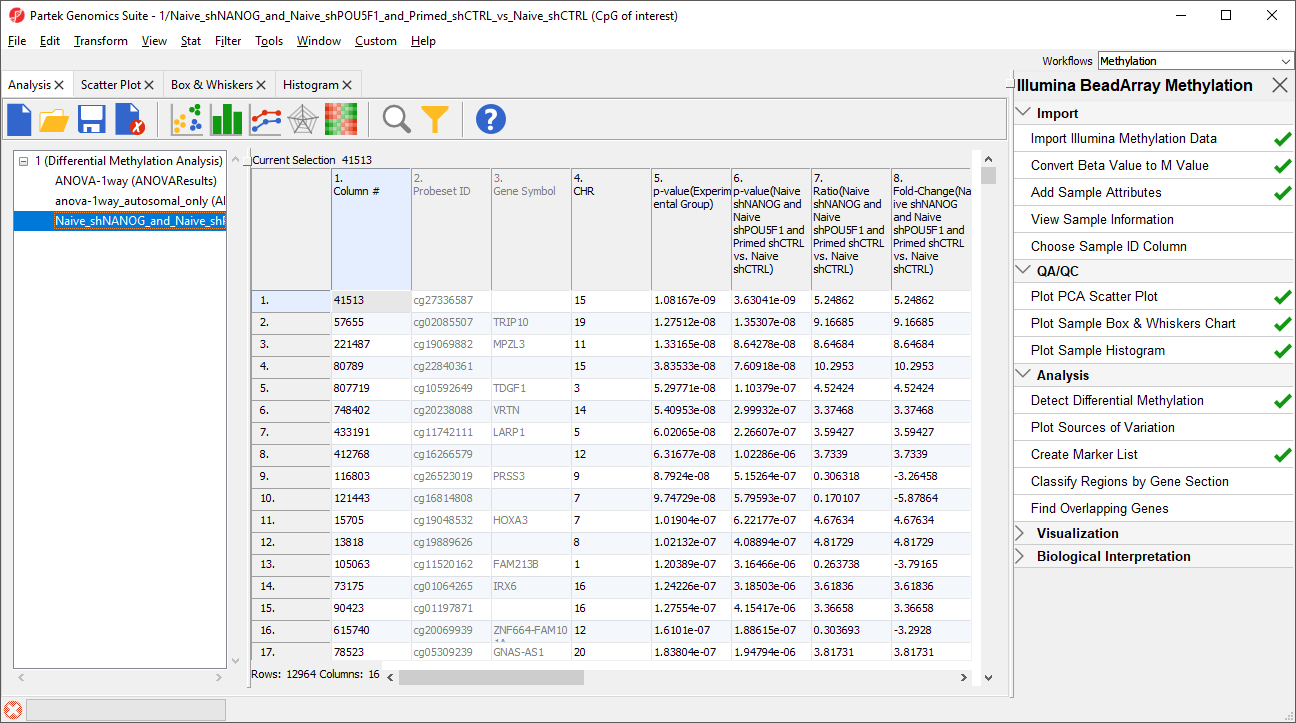
The new spreadsheet Naive_shPOU5F1_and Naive_shNANOG_and_Primed_shCTRL_vs_Naive_shCTRL (CpG of interestspreadsheet LCLs vs. B cells (LCLs vs. B cells) will open in the Analysis tab (Figure 2).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
|
...
Overview
Content Tools