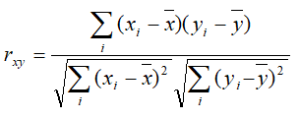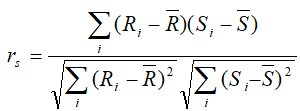Page History
| Table of Contents | ||||||
|---|---|---|---|---|---|---|
|
...
- Click the counts data node
- Click the Correlation Statistics section in the toolbox
- Click Correlation analysis
- Select the factors and interactions to include in the statistical test
...
- Correlation
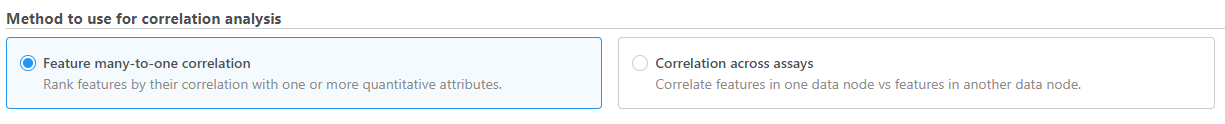
- Choose the method to use for correlation analysis (Figure 1)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Feature many-to-one correlation
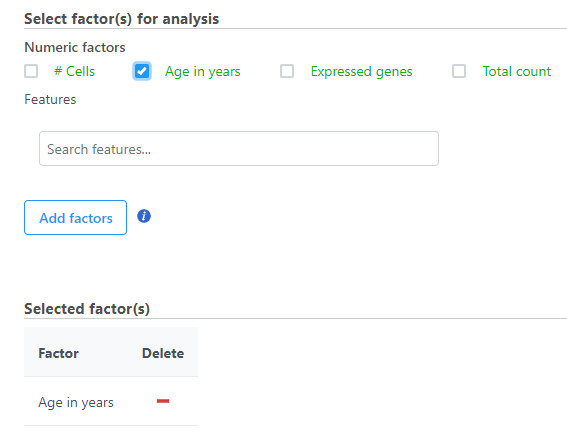
When multiple numeric factors are added, the correlation analysis will perform each factor with a feature in the data node independently. If you are interested in particular features, use the Search features box to add one or more.
- Select the factors and interactions to include in the statistical test (Figure 2).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Click Next
- It is optional to apply a lowest coverage filter or configure the advanced settings
- Click Finish to run
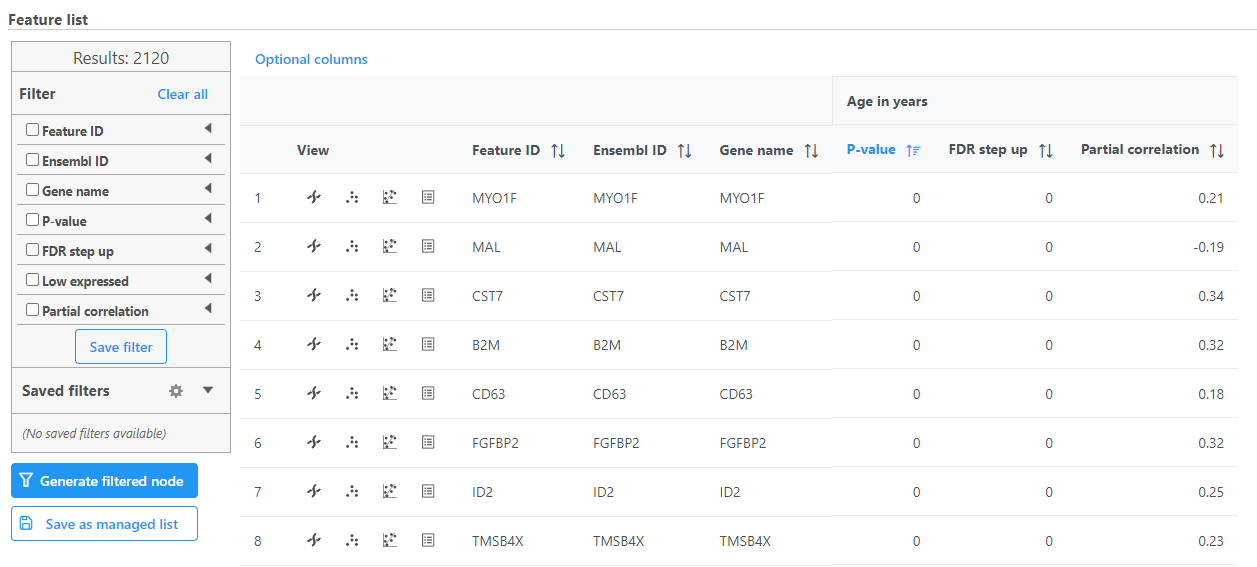
Correlation analysis produces a Feature list a Correlation data node. The ; double-click to open the task report (Figure 13) which is similar to the ANOVA/LIMMA-trend/LIMMA-voom and GSA task reports and includes a table with features on rows and statistical results on columns.
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Each numeric attribute includes p-value, adjusted p-value columns (FDR step up and/or Storey q-value if included), and a partial correlation value. Each interaction will have p-value and adjusted p-value columns (FDR step up and/or Storey q-value if included).
Each feature includes chromosome view, dot plot, correlation plot, and extra details buttons in the View column.
Correlation analysis advanced options
Low value filter
Low-value filter allows you to specify criteria to exclude features that do not meet the requirements for the calculation. If there is a filter feature task performed in the upstream analysis, the default of this filter is set to None, otherwise, the default is Lowest average coverage is set to 1.
...
Sets the type of correlation used to calculate the correlation coefficient and p-value. Options are Pearson (linear), Spearman (rank), Kendall (tau). Default is Pearson (linear).
Correlation across assays
Correlation across assays should be used to perform correlation analysis across different modalities (e.g. ATAC-Seq enriched regions vs. RNA-Seq expression) for multiomics data analysis.
- Select the data node to be compared to the node that the task has been invoked from using the Select data node button
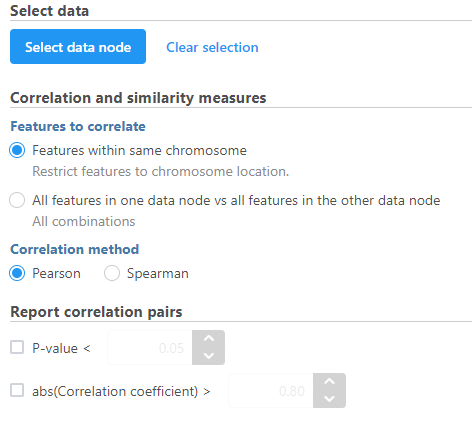
- Modify any parameters (Figure 4)
- Click Finish
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Correlation across assays analysis options
Correlation and similarity measures
Features within same chromosome: this option will restrict feature comparison to the chromosome location
All features in one data node vs all features in the other data node: this option will perform the comparison using all combinations without location constraint
Pearson: linear correlation:
Spearman: rank correlation:
Report correlation pairs
P-value: select a cut-off value for significance and only those pairs that meet the criteria will be reported
abs(Correlation coefficient): select a cutoff for reporting the absolute value of the correlation coefficient (represented by the symbol r) where a perfect relationship is 1 and no relationship is 0
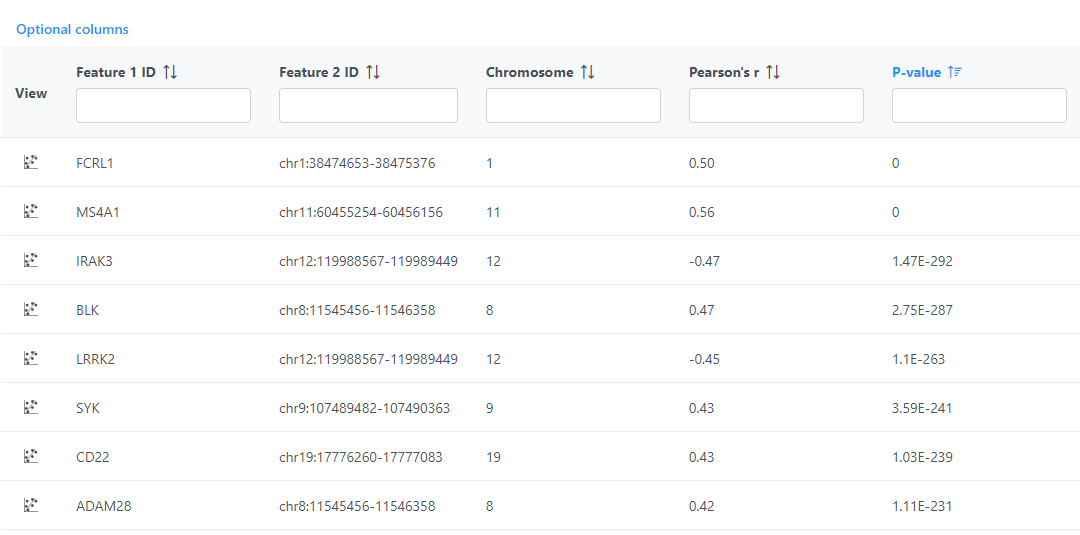
Correlation across assays produces a Correlation pair list data node; double-click to open the table (Figure 5). The table can be sorted and filtered using the column titles.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Click View correlation plot to open the correlation plot for each comparison. Scroll to the bottom of the table to download the full table report.
| Additional assistance |
|---|
| Rate Macro | ||
|---|---|---|
|
...