...
Because we have classified our cells, we can now filter based on those classifications. This can be used to focus on a single cell type for re-clustering and sub-classification or to exclude cells that are not of interest for downstream analysis.
- Click the Classified result Merged counts data node
- Click Filtering
- Click Filter groupscells
- Set to exclude Cell type is Doublets using the drop-down menus
- Click ANDOR
- Set the second filter to exclude Cell type is N/A using the drop-down menus
- Click Finish to apply the filter (Figure ?1)
| Numbered figure captions |
|---|
| SubtitleText | Set up the Filter groups task to exlcude Doublets and cells that are not classified |
|---|
| AnchorName | Filter groups |
|---|
|
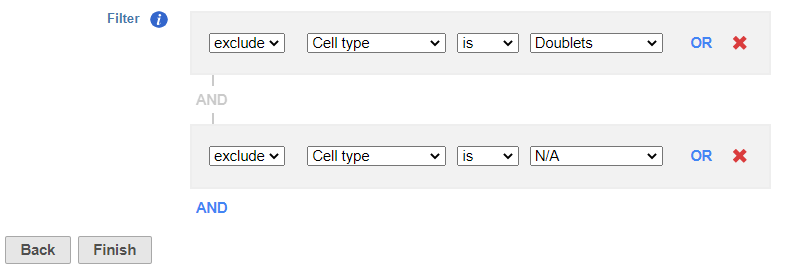 Image Removed Image Removed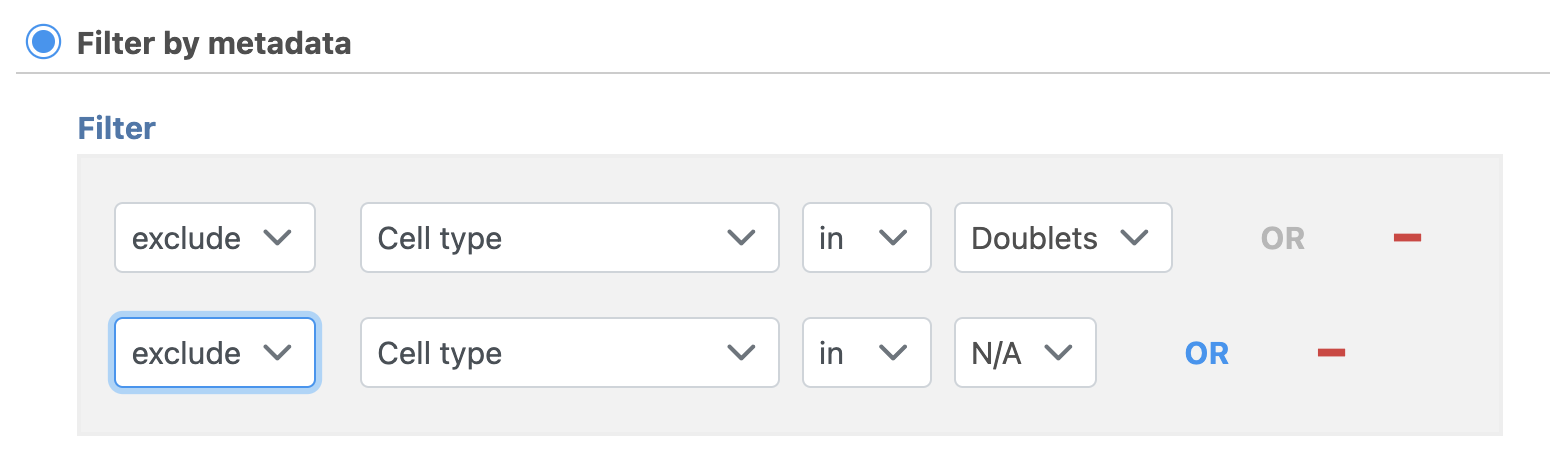 Image Added Image Added
|
This produces a Filtered counts data node (Figure ?2).
| Numbered figure captions |
|---|
| SubtitleText | Filter groups output |
|---|
| AnchorName | Filtered counts |
|---|
|
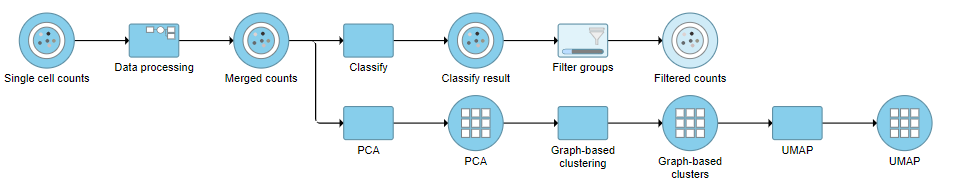 Image Removed Image Removed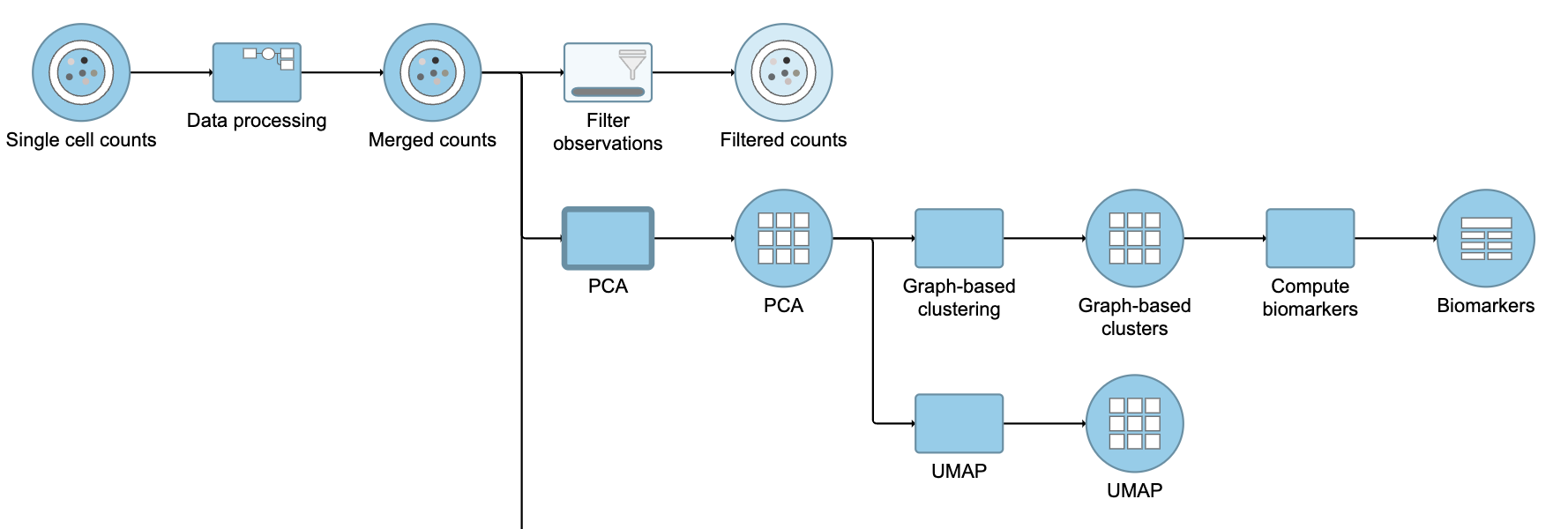 Image Added Image Added
|
Re-split the Matrix
- Click the Classified groups Filtered counts data node
- Click Pre-analysis tools
- Click Split matrixby feature type
This will produce two data nodes, one for each data type (Figure ?3). The split data nodes will both retain cell classification information.
...
| Numbered figure captions |
|---|
| SubtitleText | It is possible to re-split the merged matrix once again |
|---|
| AnchorName | Re-split the matrix |
|---|
|
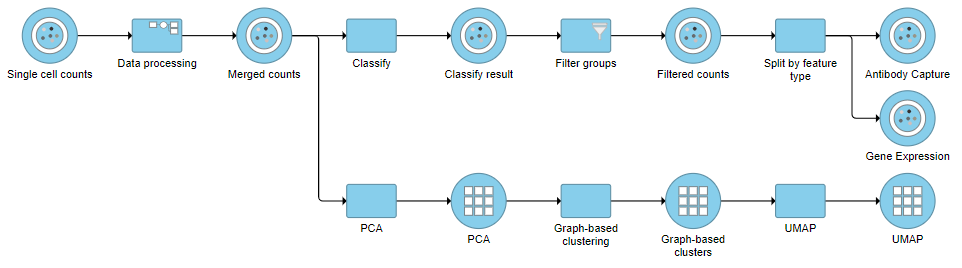 Image Removed Image Removed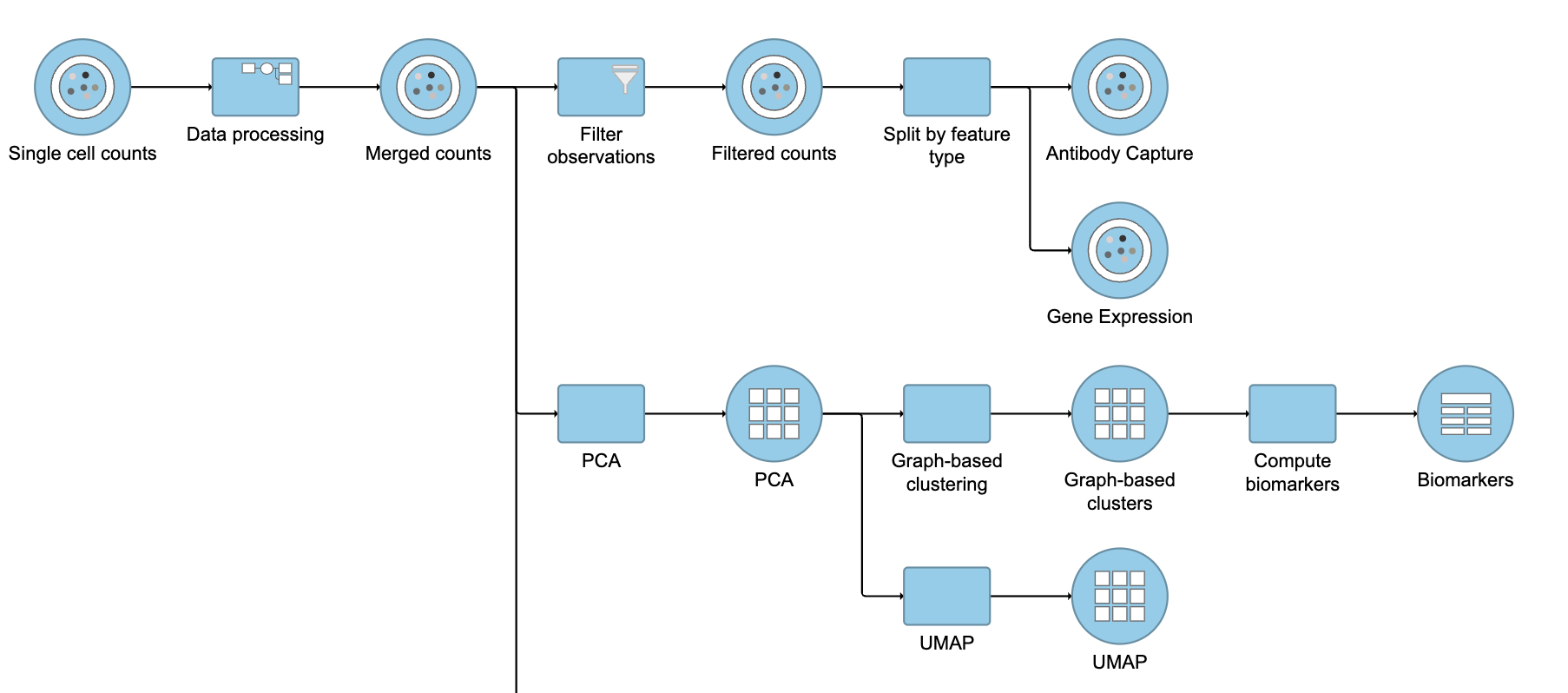 Image Added Image Added
|
Differential Analysis and Visualization - Protein Data
...
- Click the Antibody Capture data node
- Click Statistics
- Click Differential analysis
- Click GSAANOVA then click Next
The first step is to choose which attributes we want to consider in the statistical test.
- Check Click Cell type to include it in the statistical test
- Click Add factor
- Click Next
Next, we will set up the comparison we want to make. Here, we will compare the Activated and Mature B cells.
- Check Drag Activated B cells in the top panel
- Check Drag Mature B cells in the bottom panel
- Click Add comparison
...
- Click Finish to run the statistical test (Figure ?4)
| Numbered figure captions |
|---|
| SubtitleText | Setting up a comparison for differentially expressed proteins |
|---|
| AnchorName | CITE-Seq GSA task set up |
|---|
|
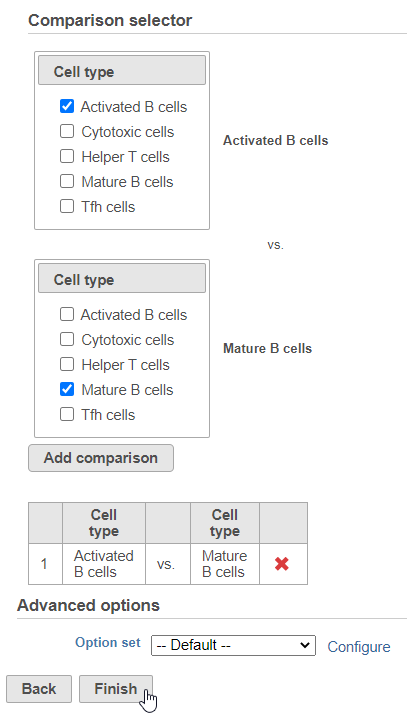 Image Removed Image Removed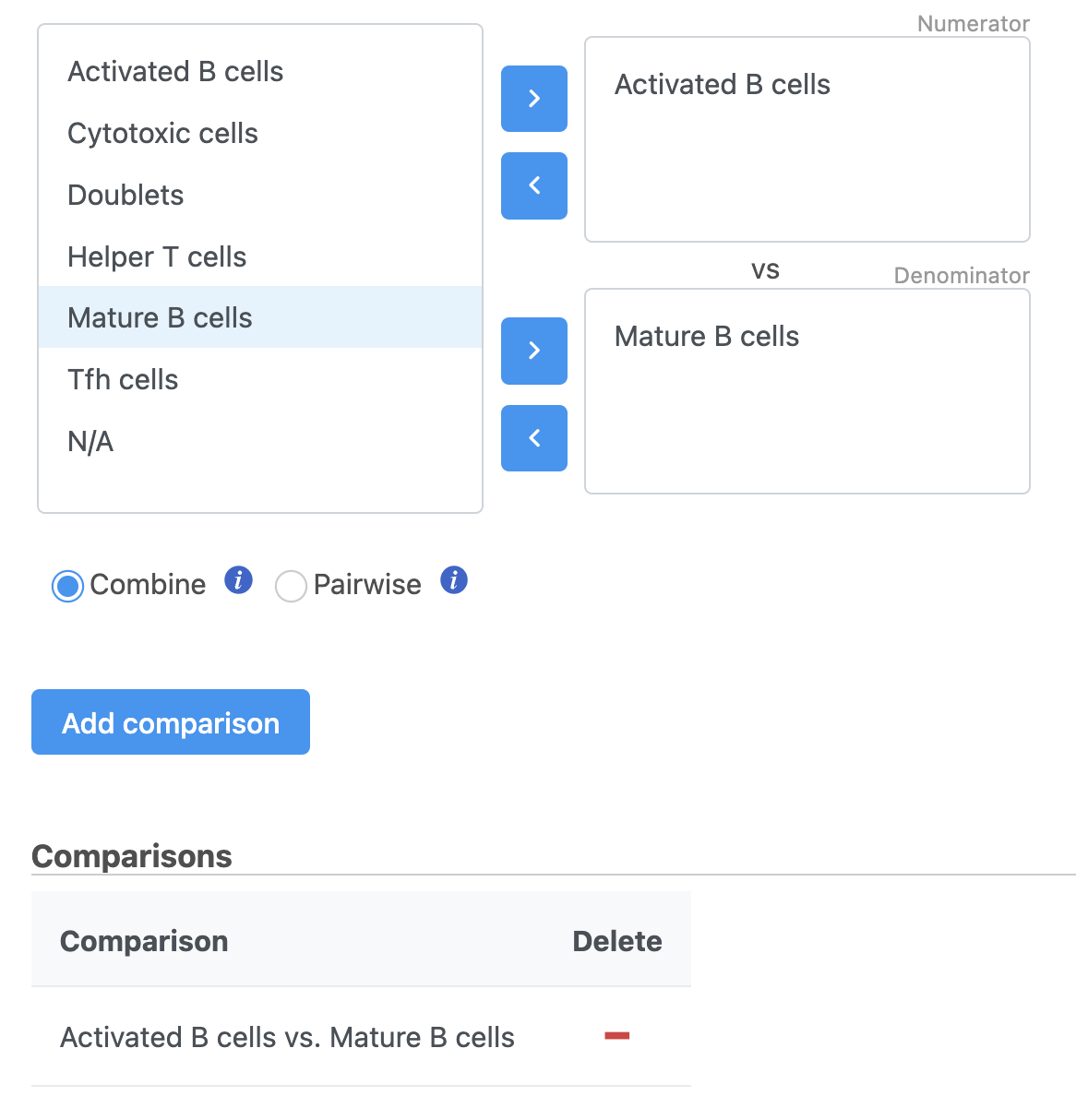 Image Added Image Added
|
The GSA ANOVA task produces a GSA an ANOVA data node.
- Double-click the GSA ANOVA data node to open the task report
The report lists each feature tested, giving p-value, false discovery rate adjusted p-value (FDR step up), and fold change values for each comparison (Figure ?5).
| Numbered figure captions |
|---|
| SubtitleText | GSA report for protein expression data |
|---|
| AnchorName | GSA protein result |
|---|
|
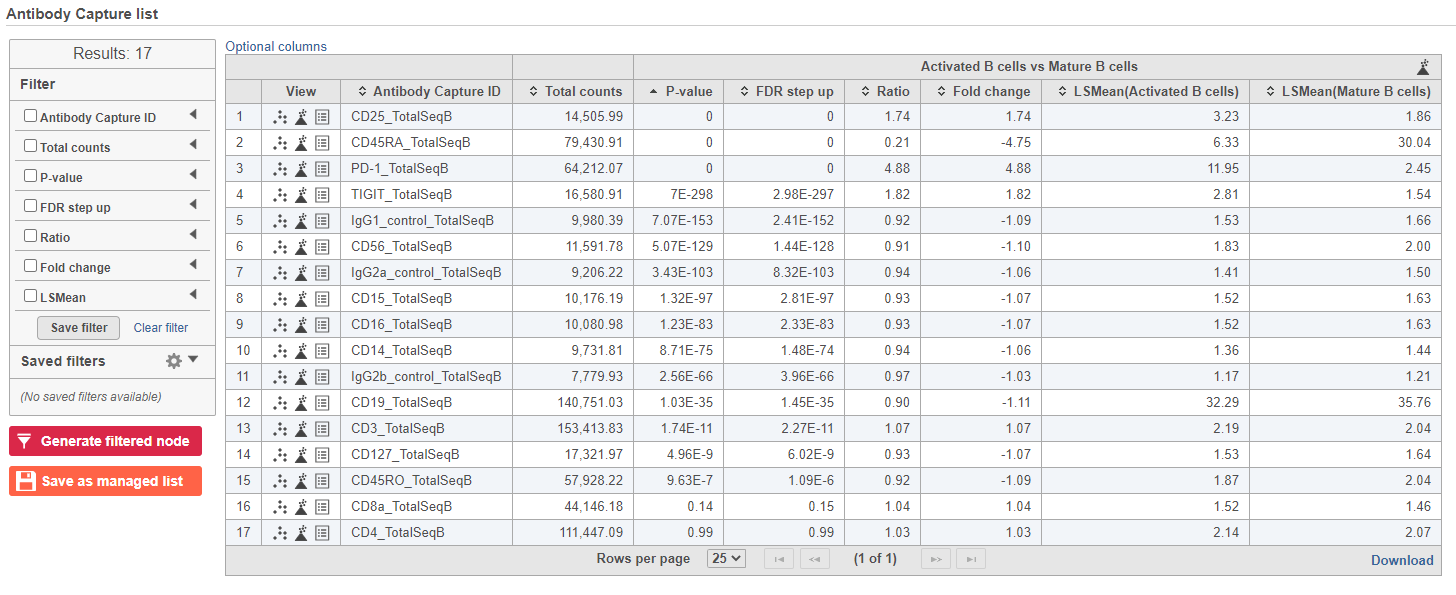 Image Removed Image Removed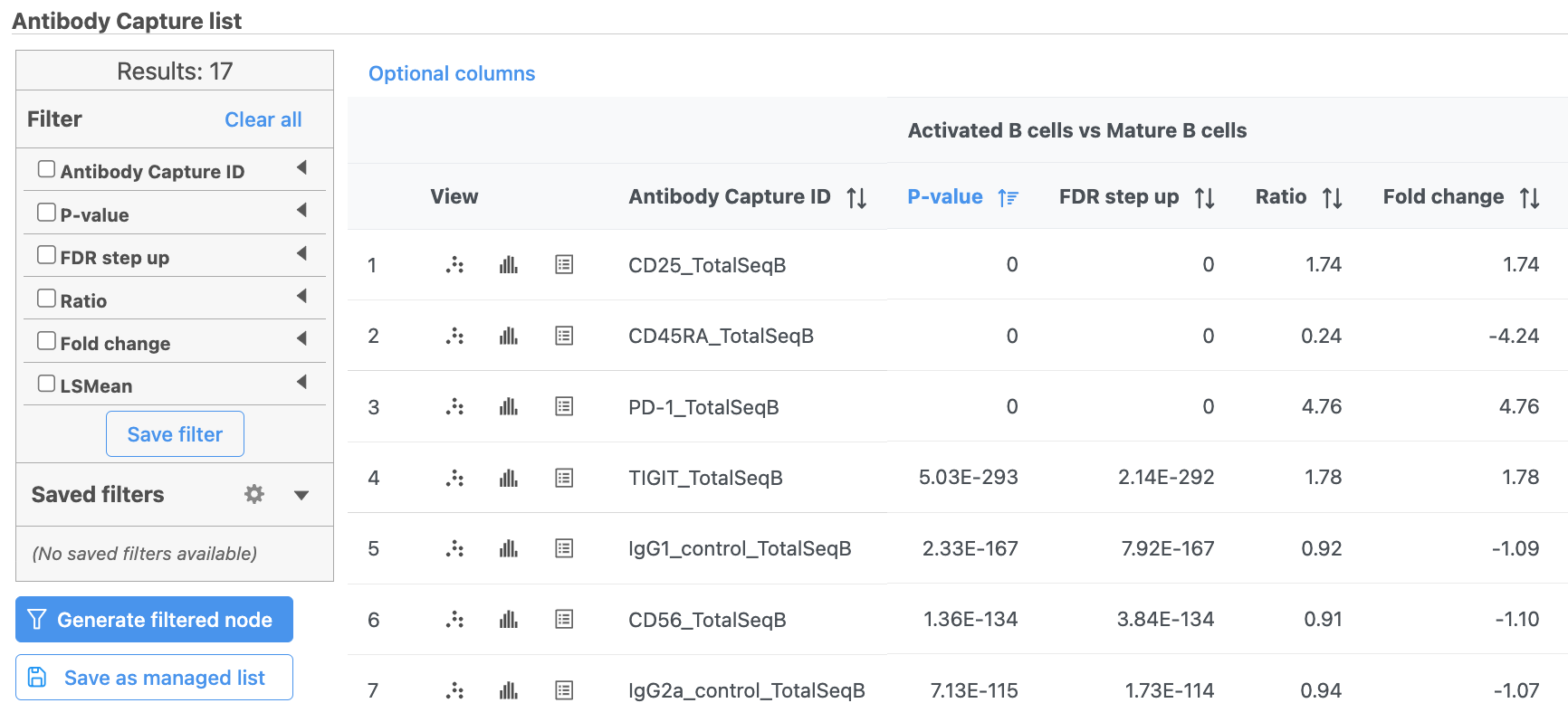 Image Added Image Added
|
In addition to the listed information, we can access dot and violin plots for each gene or protein from this table.
...
This opens a dot plot in a new data viewer session, showing CD45A expression for cells in each of the classifications (Figure ?)6). First, we exclude Doublets and N/A cells from the plot:
- Open Select and filter, select Criteria
- Drag "Cell type" from the legend title to the Add criteria box
- Uncheck Doublets and N/A
- Click to include selected points
| Numbered figure captions |
|---|
| SubtitleText | CD45RA dot plot for all cells |
|---|
| AnchorName | CD45RA dot plot |
|---|
|
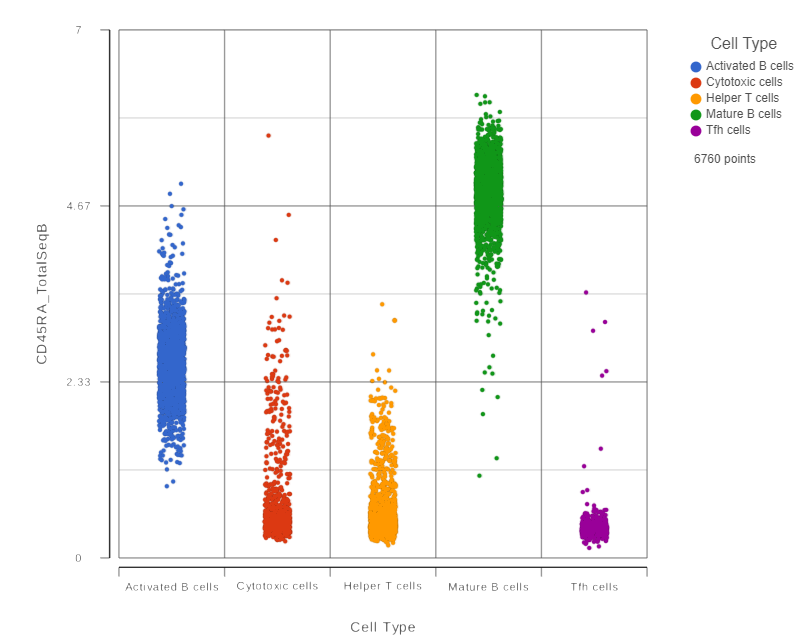 Image Removed Image Removed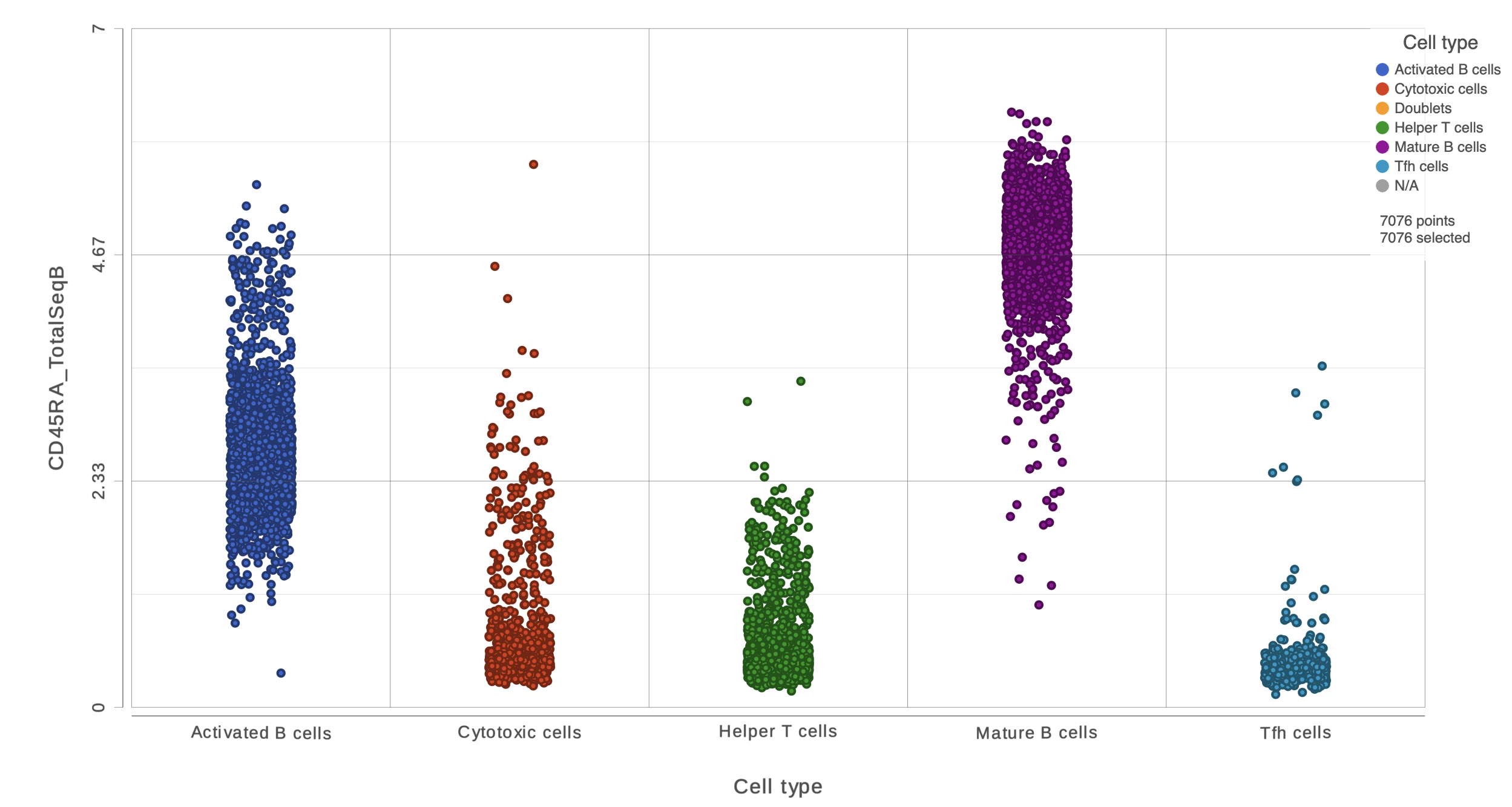 Image Added Image Added
|
We can use the Configuration panel on the left to edit this plot.
- Expand Open the Summary card Style icon
- Switch on Violins under Summary
- Switch on Overlay Overlay under Summary
- Switch on Colored
- Expand the Data card
- Use Colored under Summary
- Select the Graph-based clustering node in the Color by section
- Color by Graph-based clusters under Color and use the slider to increase decrease the Jitter Opacity
- Expand Open the Color card Axes icon
- Select the Graph-based clustering node in the X axis section
- Change the X axis data to Graph-based clusters
- Use the slider to decrease the Opacity increase the Jitter on the X axis (Figure ?7)
| Numbered figure captions |
|---|
| SubtitleText | Use Configure the Configuration panel to configure the dot plot using the tools on the left |
|---|
| AnchorName | Configure dot plot |
|---|
|
 Image Removed Image Removed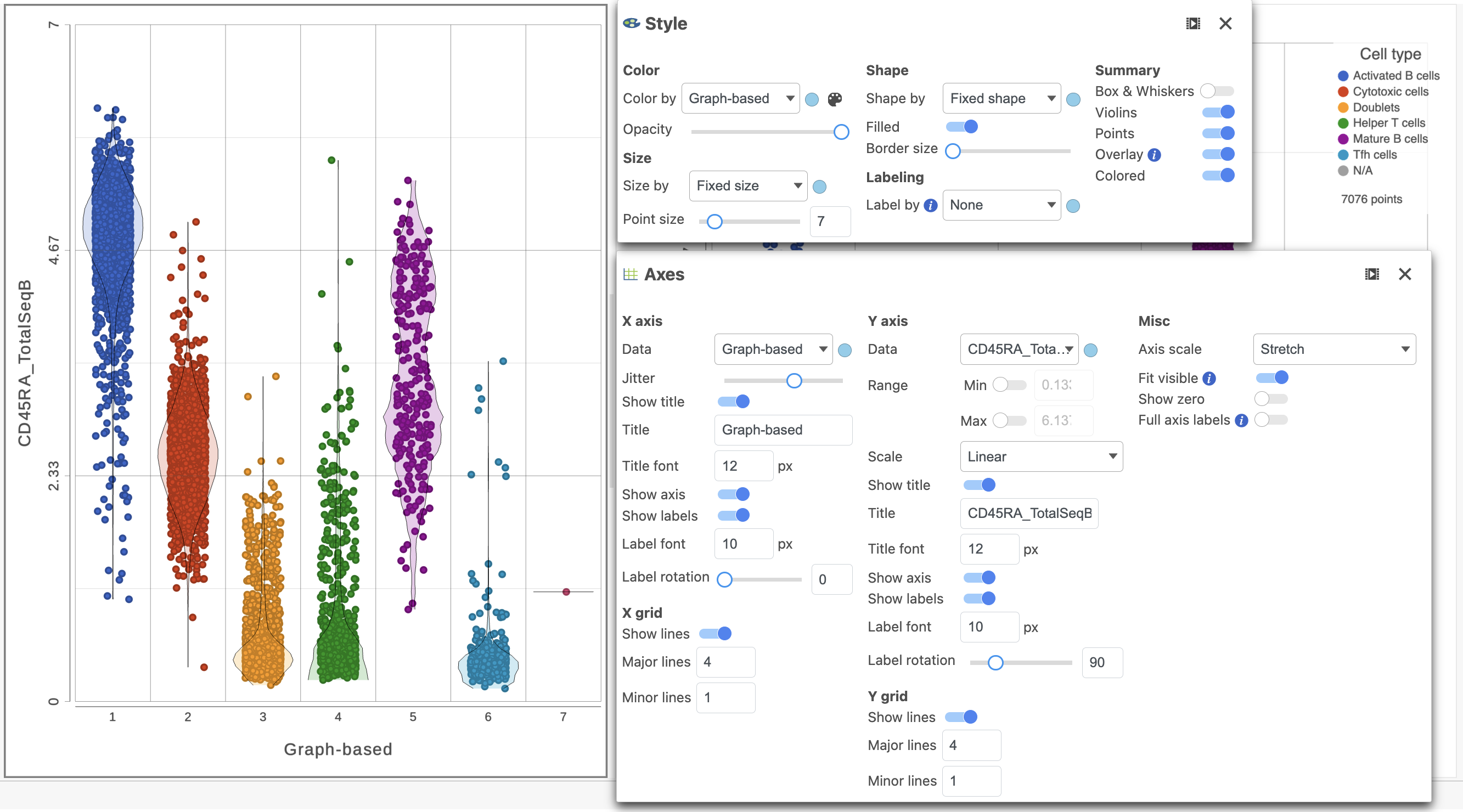 Image Added Image Added
|
- Click the project name to return to the Analyses tab
To visualize all of the proteins at the same time, we can make a hierarchical clustering heat map.
- Click the GSA ANOVA data node
- Click Exploratory analysis in the toolbox
- Click Hierarchical clustering/heat mapCheck Samples at the top to cluster the cells in addition to featuresheatmap
- In the Cell order section, choose Graph-based clusters from the Assign order drop-down list
- Click Finish to run with the other default settings
- Double-click the Hierarchical clustering task node to open the heat map (Figure ?heatmap
The heatmap can easily be customized using the tools on the left.
- Open the Axes icon
- Switch off Show Row labels
- Increase the Font to 16 (Figure 8)
| Numbered figure captions |
|---|
| SubtitleText | Heatmap showing expression of protein markers before configurationaltered Axes labels |
|---|
| AnchorName | Heatmap of proteins |
|---|
|
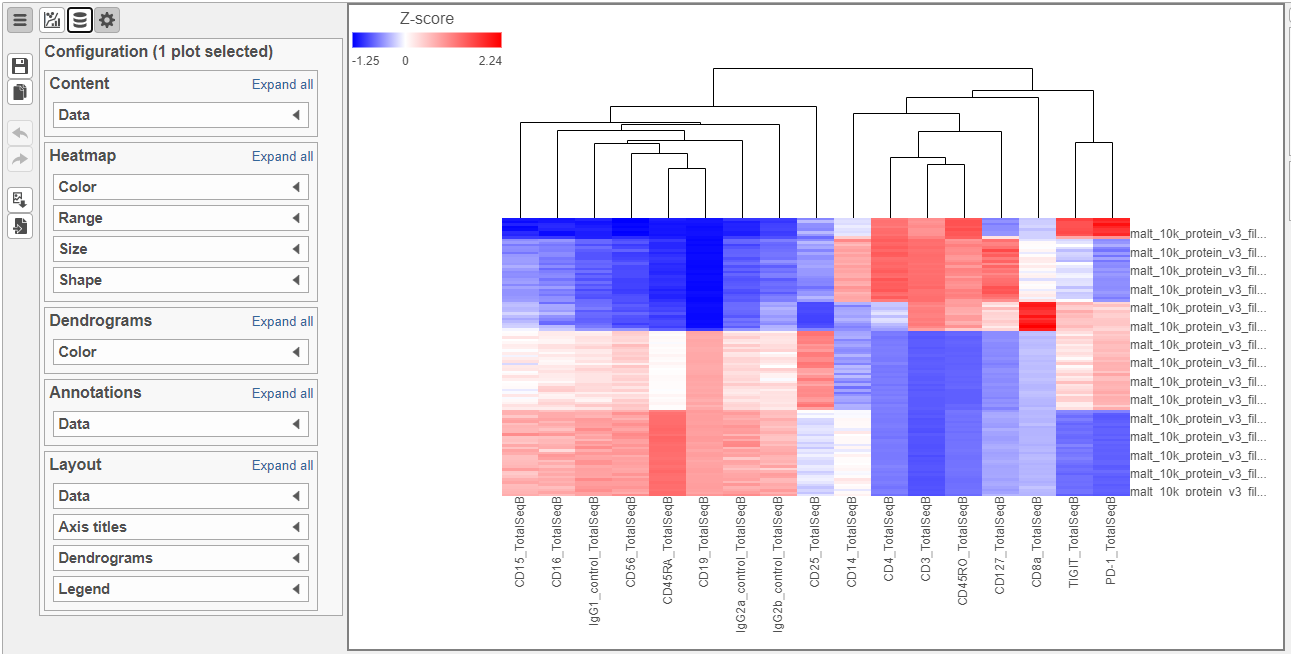 Image Removed Image Removed
|
The heat map can easily be customized to illustrate our results.
- Click
 Image Removed to transpose the heat map
Image Removed to transpose the heat map - Set High to 2.6 to match the low range
- Set the Sample dendrogram to By sample attribute Cell type
- Set Attributes to Cell type
- Click
 Image Removed and set Rotation to 0
Image Removed and set Rotation to 0 - Uncheck Samples under Show labels
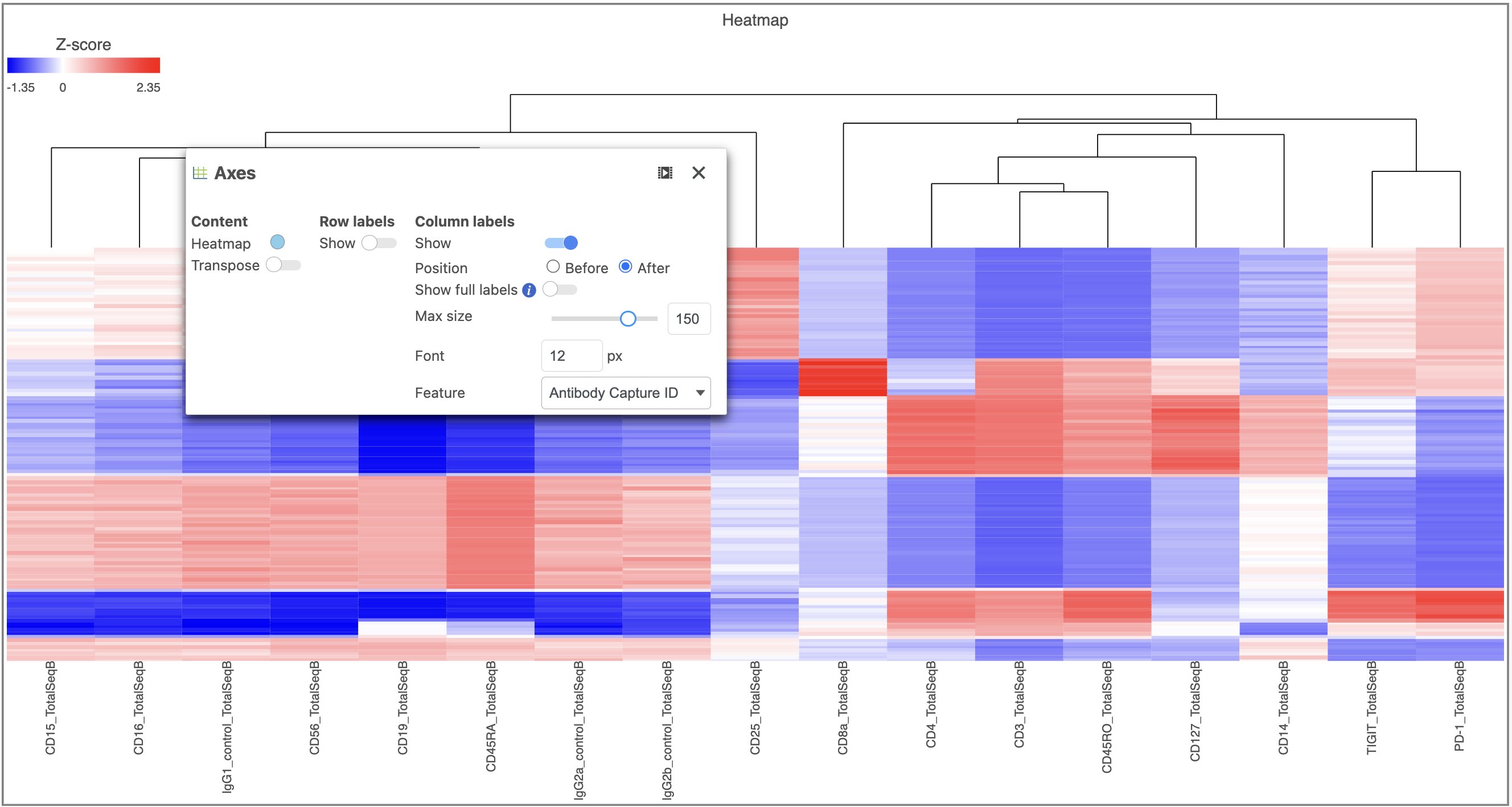 Image Added Image Added
|
- Activate the Transpose switch which will switch the Row and Column labels, so now the Row labels will be shown (Figure 9)
| Numbered figure captions |
|---|
| SubtitleText | Transpose the Heatmap to switch the columns and rows |
|---|
| AnchorName | Layout card |
|---|
|
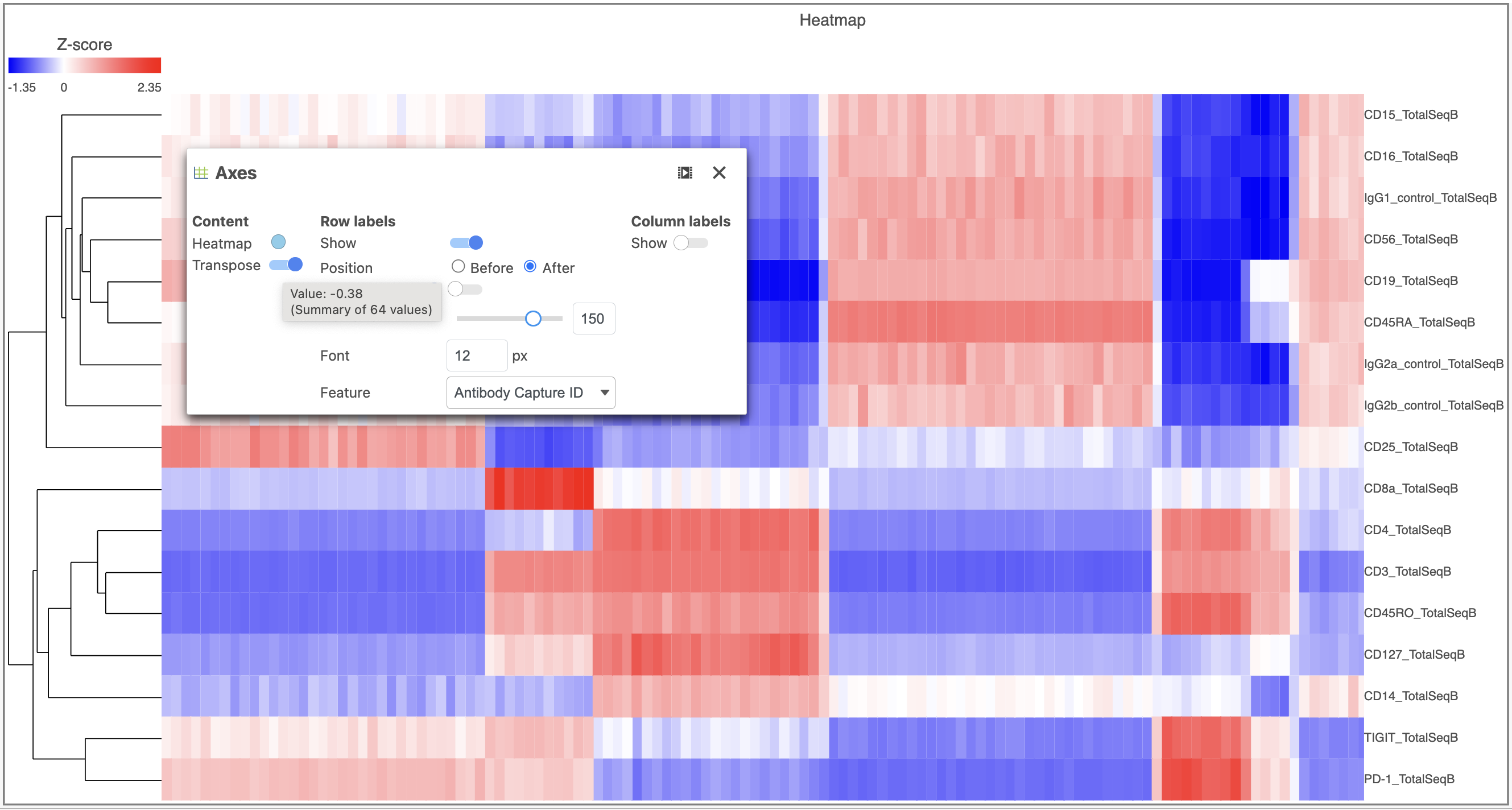 Image Added Image Added
|
- Open the Dendrograms icon
- Choose Row color By cluster and change Row clusters to 4
- Change Row dendrogram size to 80 (Figure 10)
| Numbered figure captions |
|---|
| SubtitleText | Configure the Dendrograms settings |
|---|
| AnchorName | Annotations card |
|---|
|
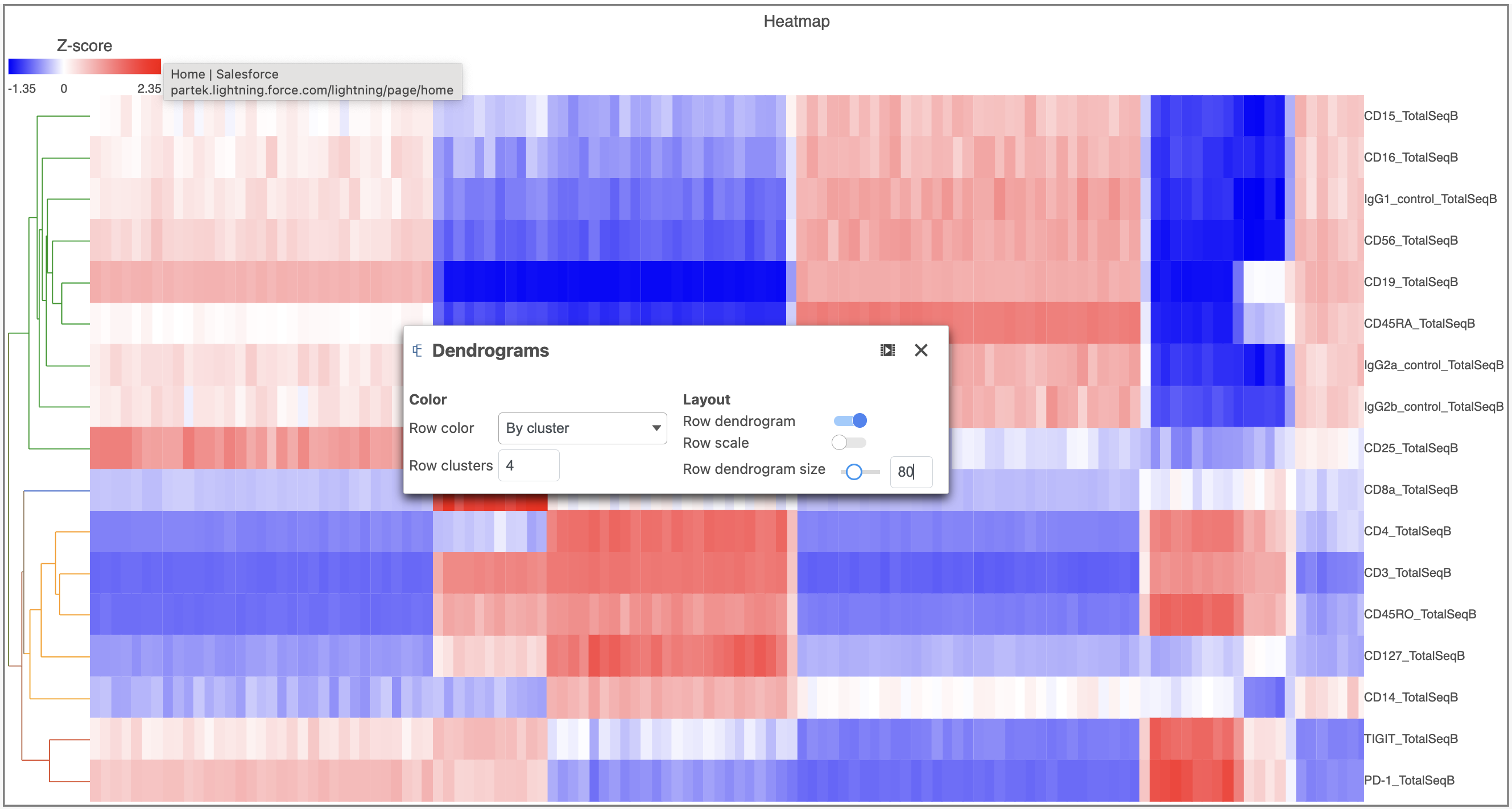 Image Added Image Added
|
- In the Heatmap icon
- Navigate to Range under Color
- Set the Min and Max to -1.2 and 1.2, respectively
- Change the Shape to Circle (Figure 11)
| Numbered figure captions |
|---|
| SubtitleText | Configure the Heatmap icon |
|---|
| AnchorName | Heatmap card |
|---|
|
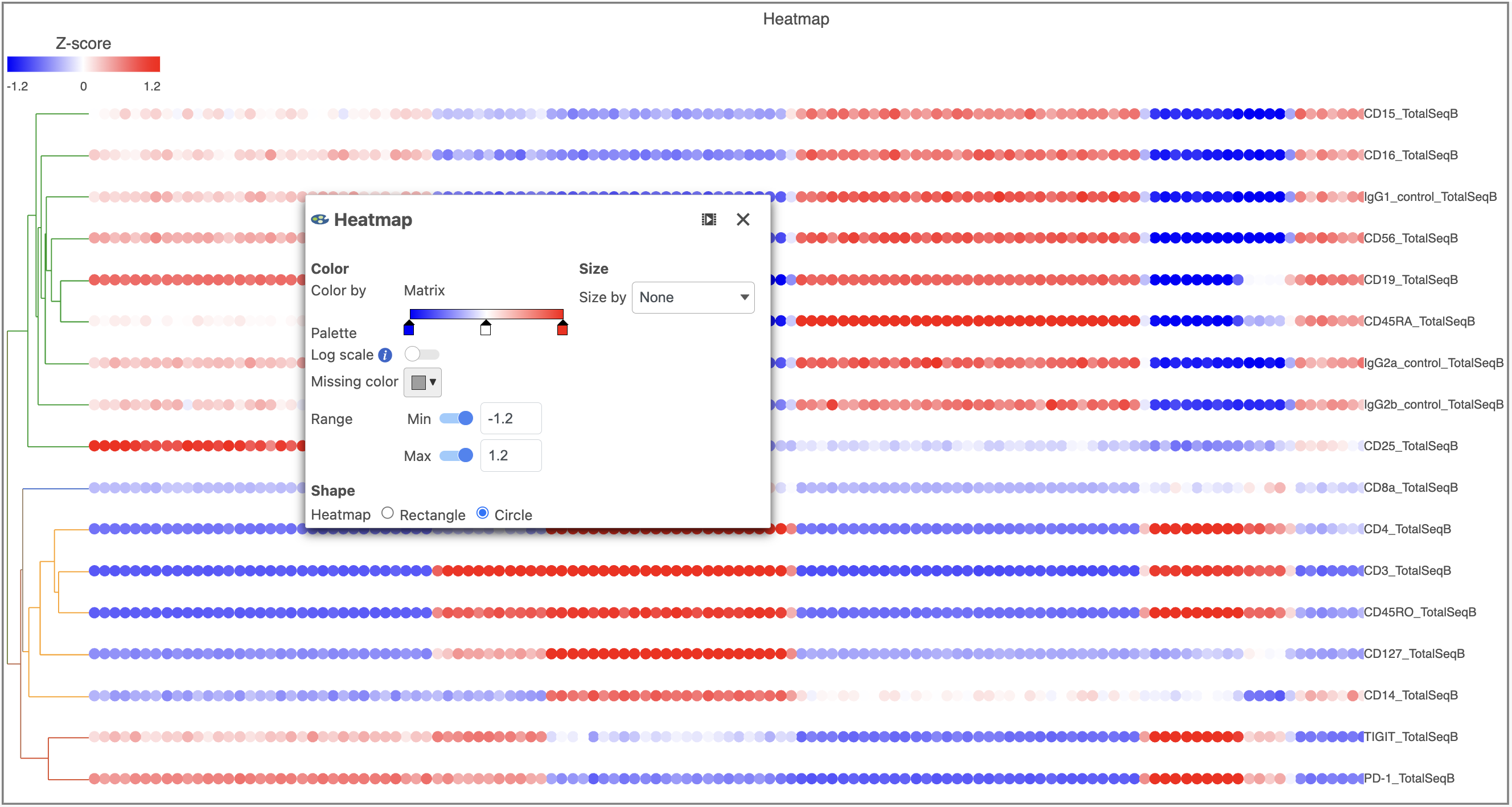 Image Added Image Added
|
- Switch the Shape back to Rectangle
- Change the Color Palette by clicking on the color squares and selecting colors from the rainbow. Click outside of the selection box to exit this selection. The color options can be dragged alone the Palette to highlight value differences (Figure 12).
| Numbered figure captions |
|---|
| SubtitleText | Heatmap showing expression of protein markers after configuration changing the Heatmap settings further |
|---|
| AnchorName | Heatmap of proteins configured |
|---|
|
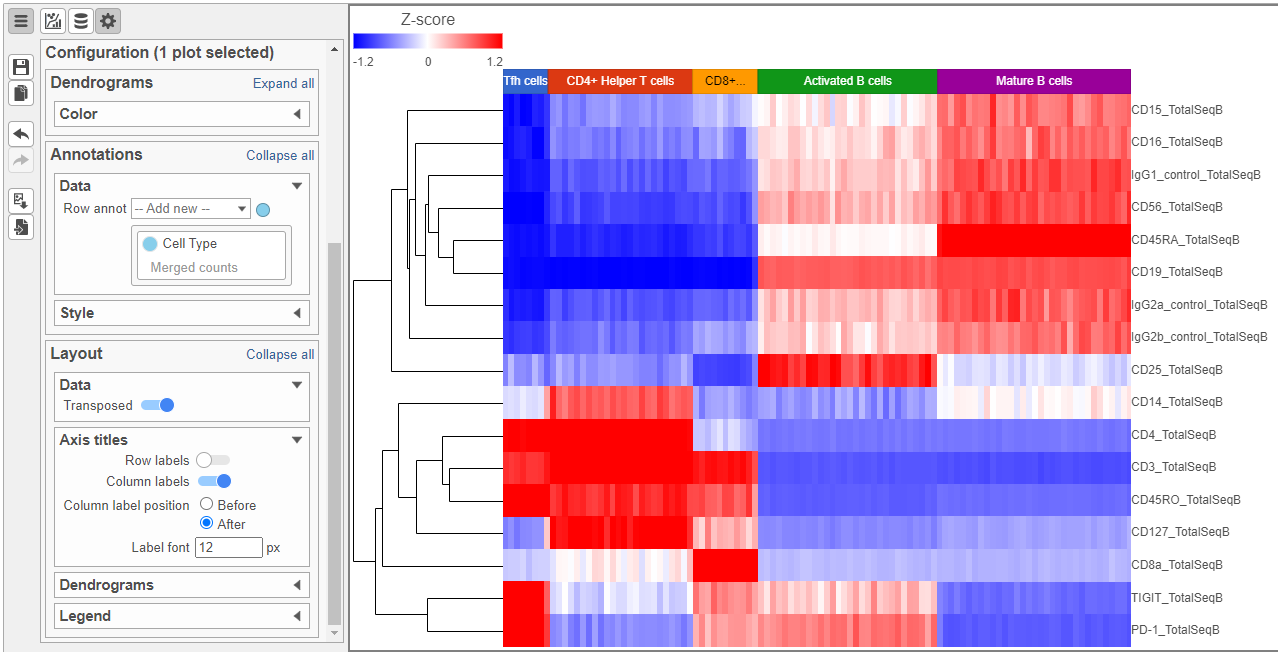 Image Removed Image Removed
|
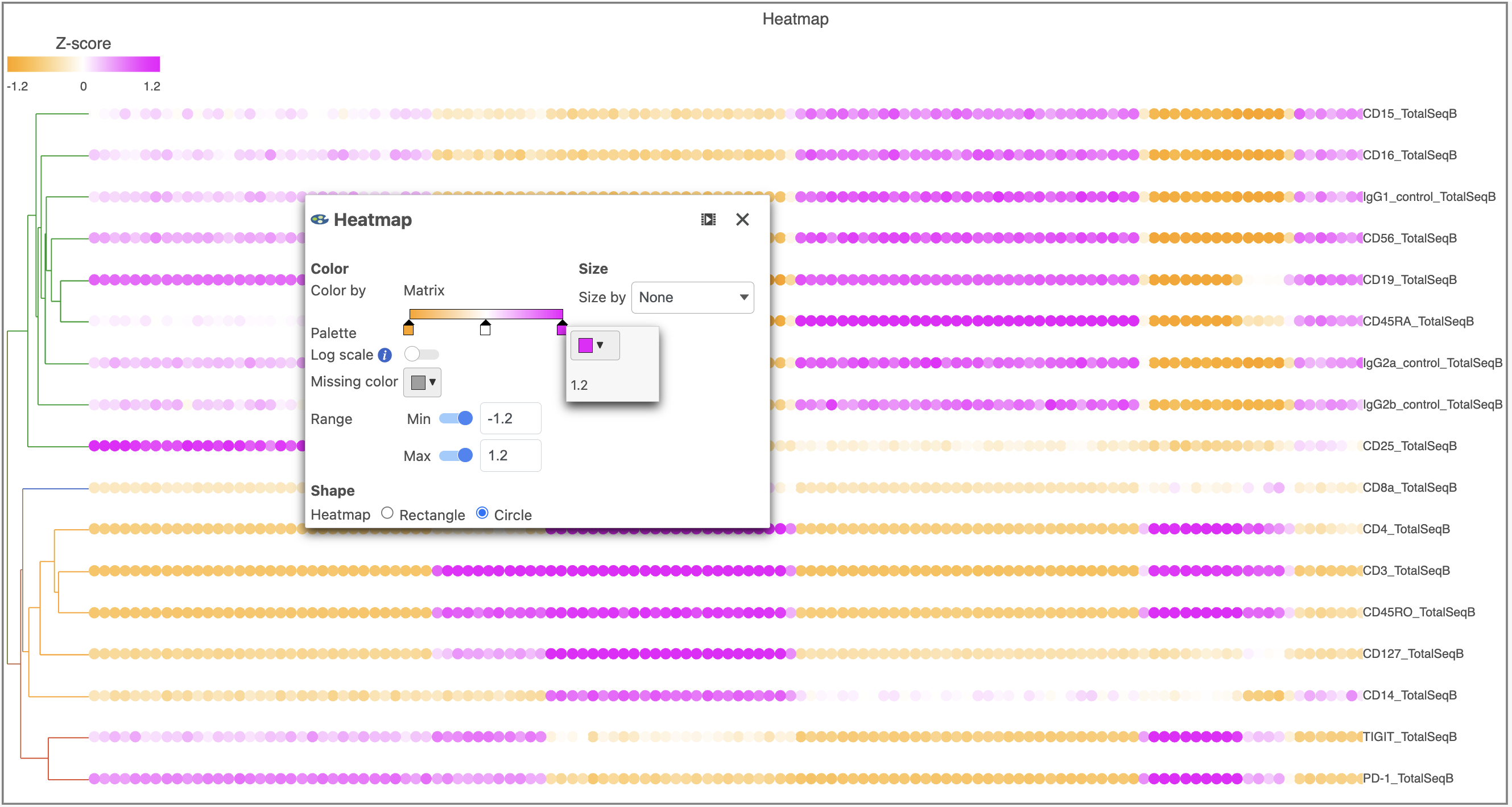 Image Added Image Added
|
Feel free to explore the other tool options on the left to customize the plot further.
Differential Analysis, Visualization, and Pathway analysis - Gene Expression Data
...
- Click the project name to return to the Analyses tab
- Click the Gene Expression data node
- Click the Antibody Capture data node
- Click Statistics
- Click Differential analysis
- Click GSA
- Check Cell type to include it in the statistical test
- ANOVA then click Next
- Click Cell type
- Click Add factor
- Click Next
- Check Drag Activated B cells in the top panel
- Check Drag Mature B cells in the bottom panel
- Click Add comparison Click comparison
The comparison should appear in the table as Activated B cells vs. Mature B cells.
- Click Finish to run the statistical test
As before, this will generate a GSA an ANOVA task node and a GSA n ANOVA data node.
- Double-click the GSA ANOVA task node to open the task report (Figure ?Figure 13)
| Numbered figure captions |
|---|
| SubtitleText | GSA report for the gene expression data |
|---|
| AnchorName | GSA genes result |
|---|
|
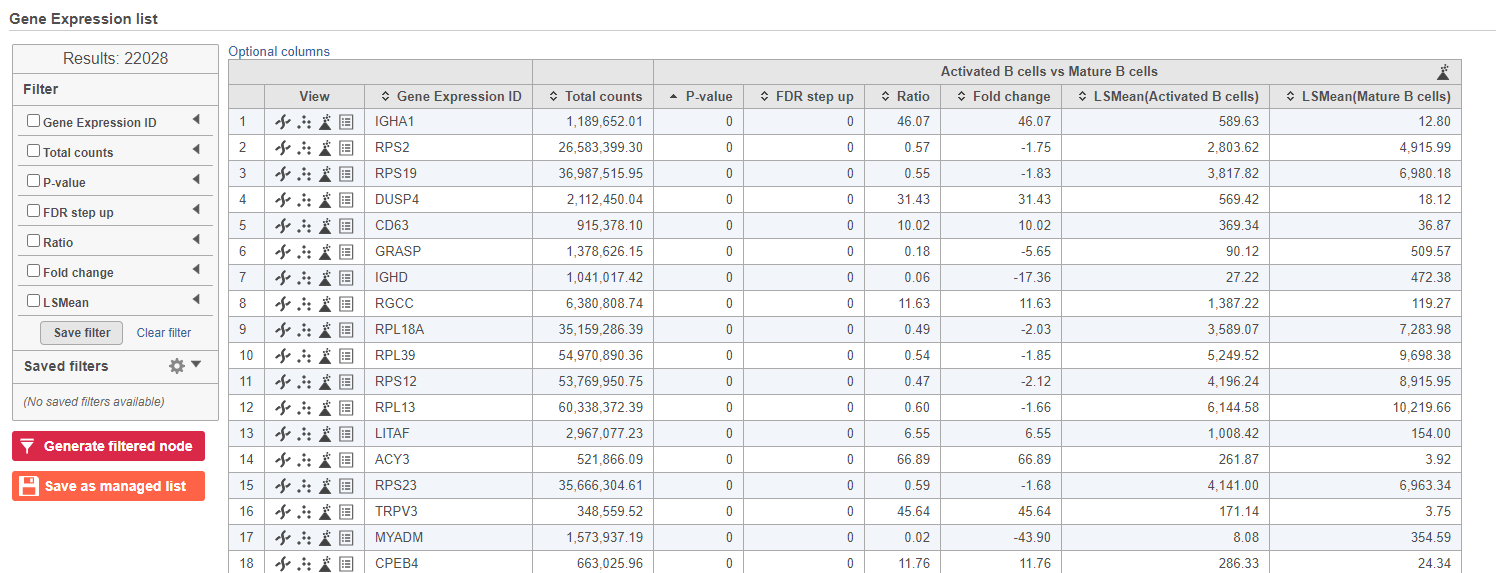 Image Removed Image Removed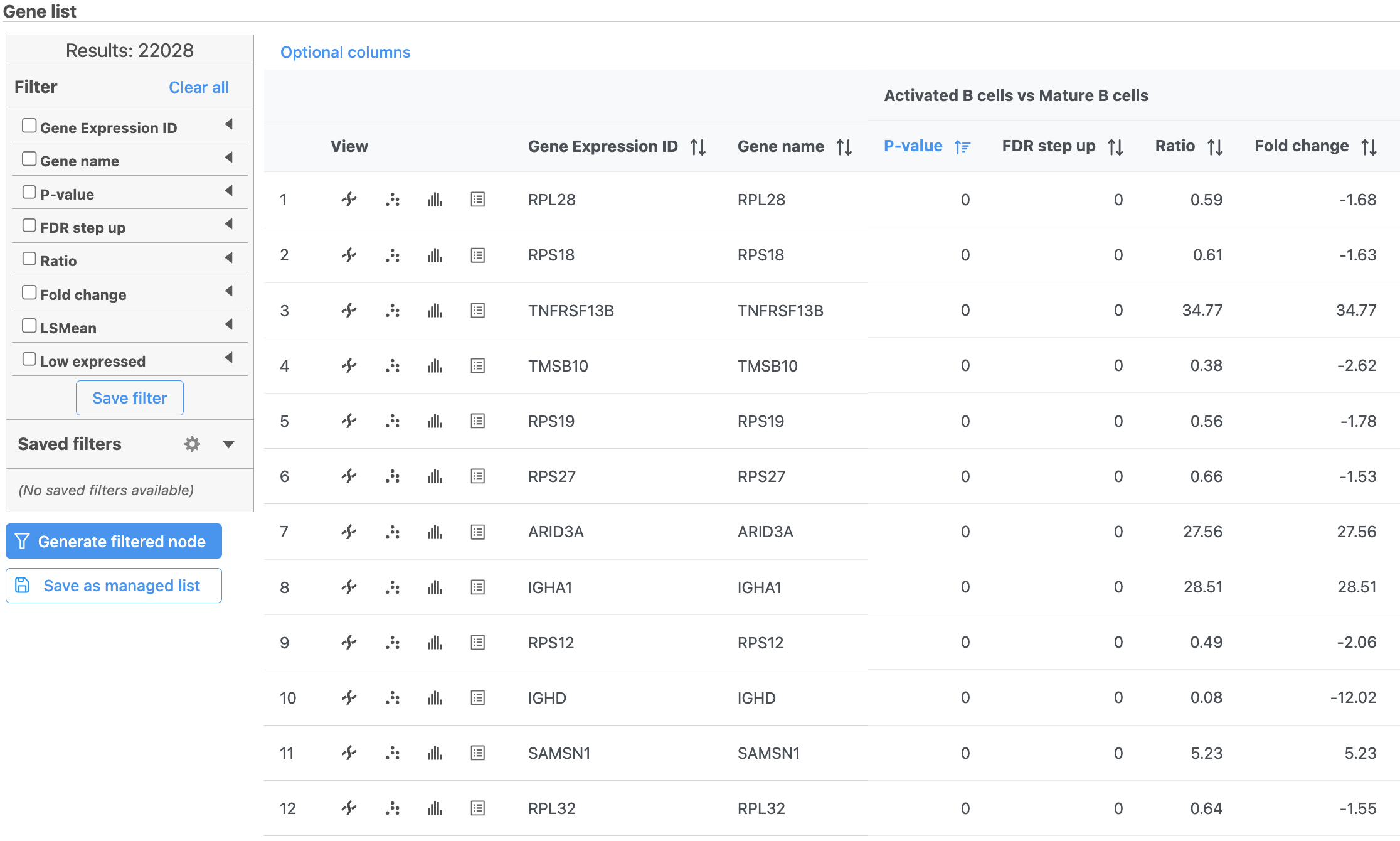 Image Added Image Added
|
Because more than 20,000 genes have been analyzed, it is useful to use a volcano plot to get an idea about the overall changes.
...
The Volcano plot opens in a new data viewer session, in a new tab in the web browser. It shows each gene as a point with cutoff lines set for P-value (y-axis) and fold-change (x-axis). By default, the P-value cutoff is set to 0.05 and the fold-change cutoff is set at |2| (Figure ?Figure 14).
The plot can be configured using various options in the Configuration card on the tools on the left. For example, the Color, Size and Shape cards can the Style icon can be used to change the appearance of the points. The X and Y-axes can be changed in the Data card. The Significance card can the Axes icon. The Statistics icon can be used to set different Fold-change and P-value thresholds for coloring up/down-regulated genes. The in plot controls can be used to transpose  Image Addedthe volcano plot (Figure 14).
Image Addedthe volcano plot (Figure 14).
| Numbered figure captions |
|---|
| SubtitleText | The volcano plot can be configured Configured using various options in the Configuration and Selection cardsthe icons on the left and in plot controls |
|---|
| AnchorName | Volcano plot gene expression |
|---|
|
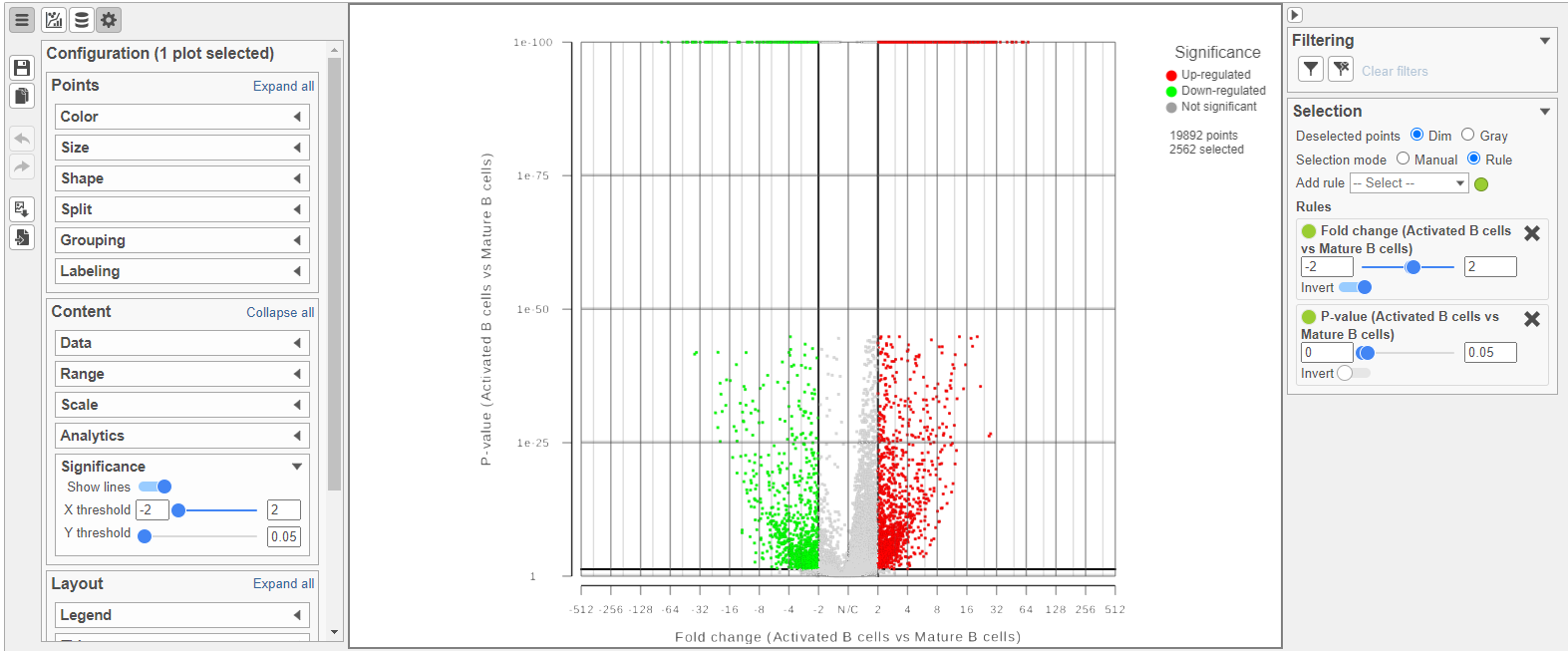 Image Removed Image Removed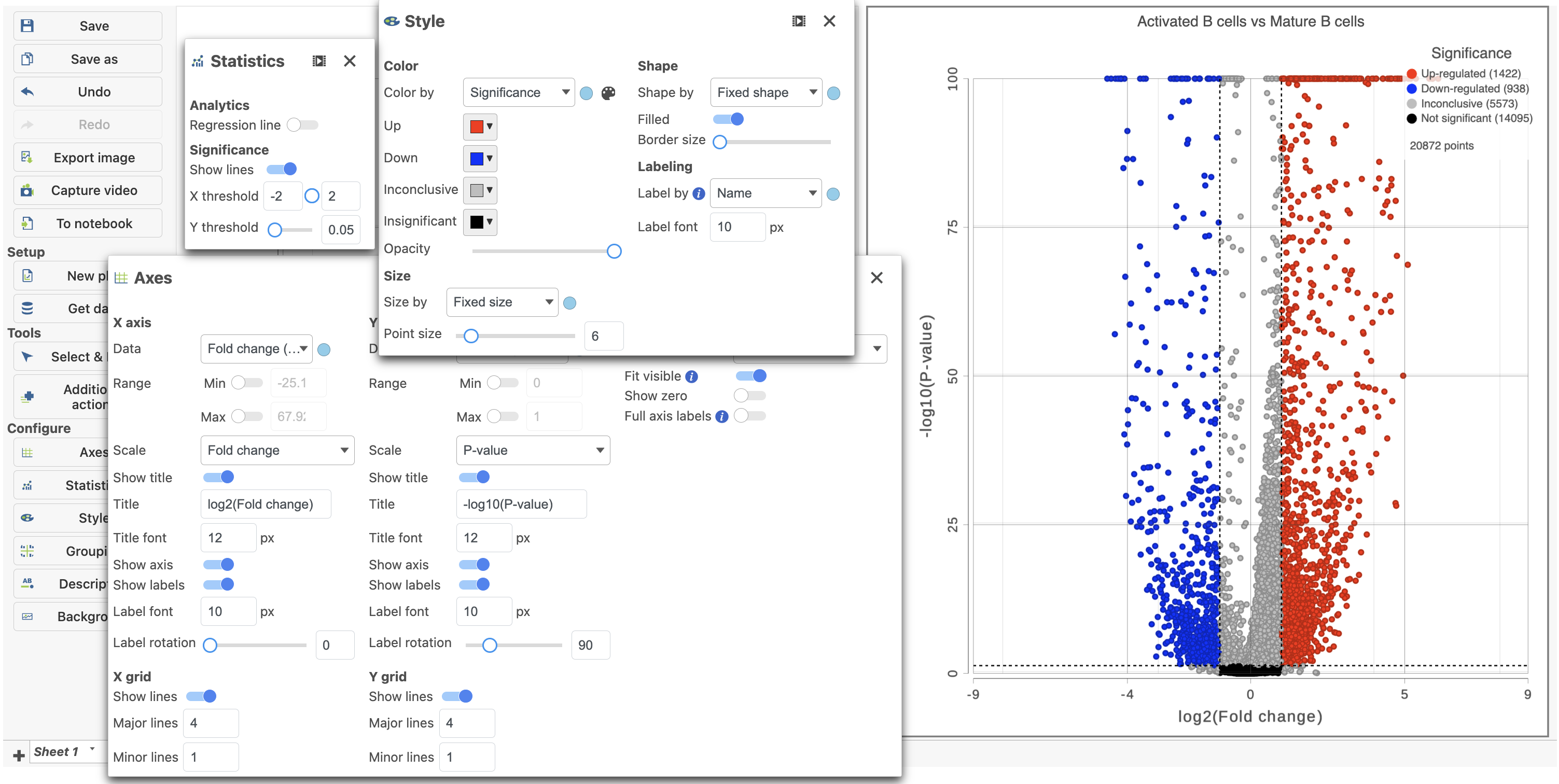 Image Added Image Added
|
- Click the GSA ANOVA report tab in your web browser to return to the full report
...
The number at the top of the filter will update to show the number of included genes (Figure ?Figure 15).
| Numbered figure captions |
|---|
| SubtitleText | Use the panel on the left to filter the list for significant genes |
|---|
| AnchorName | Significant genes |
|---|
|
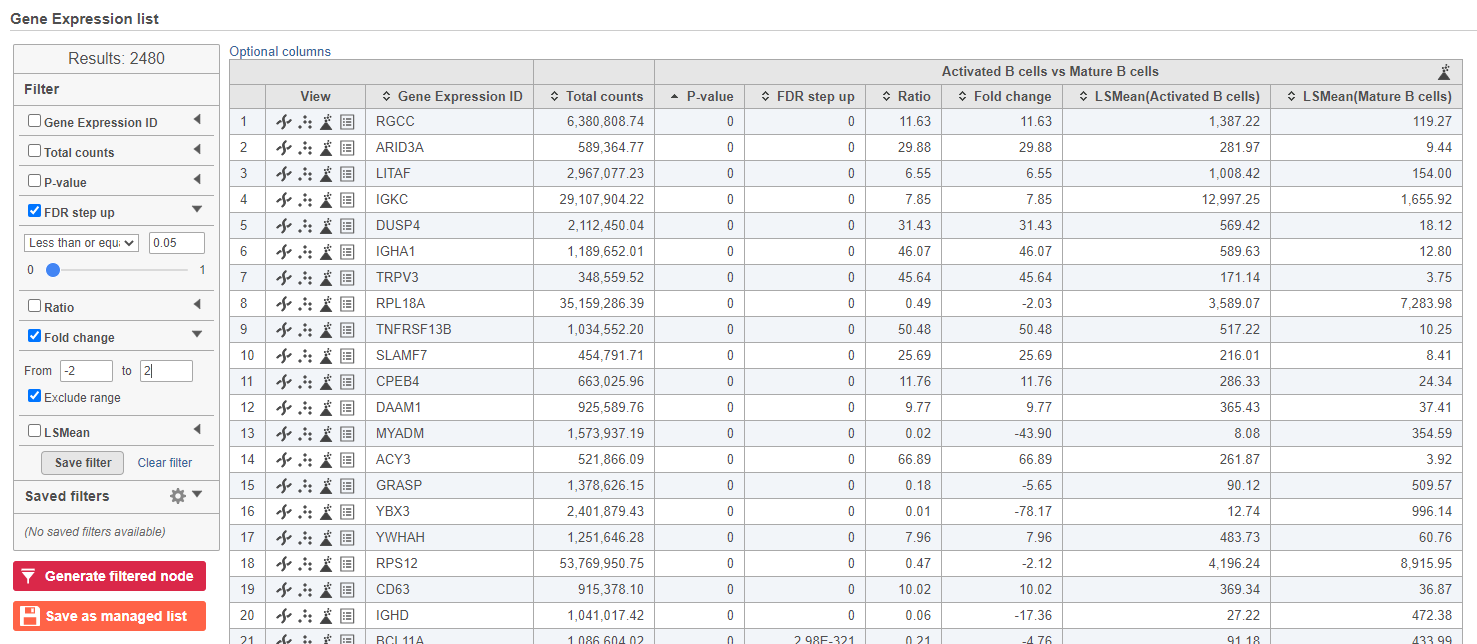 Image Removed Image Removed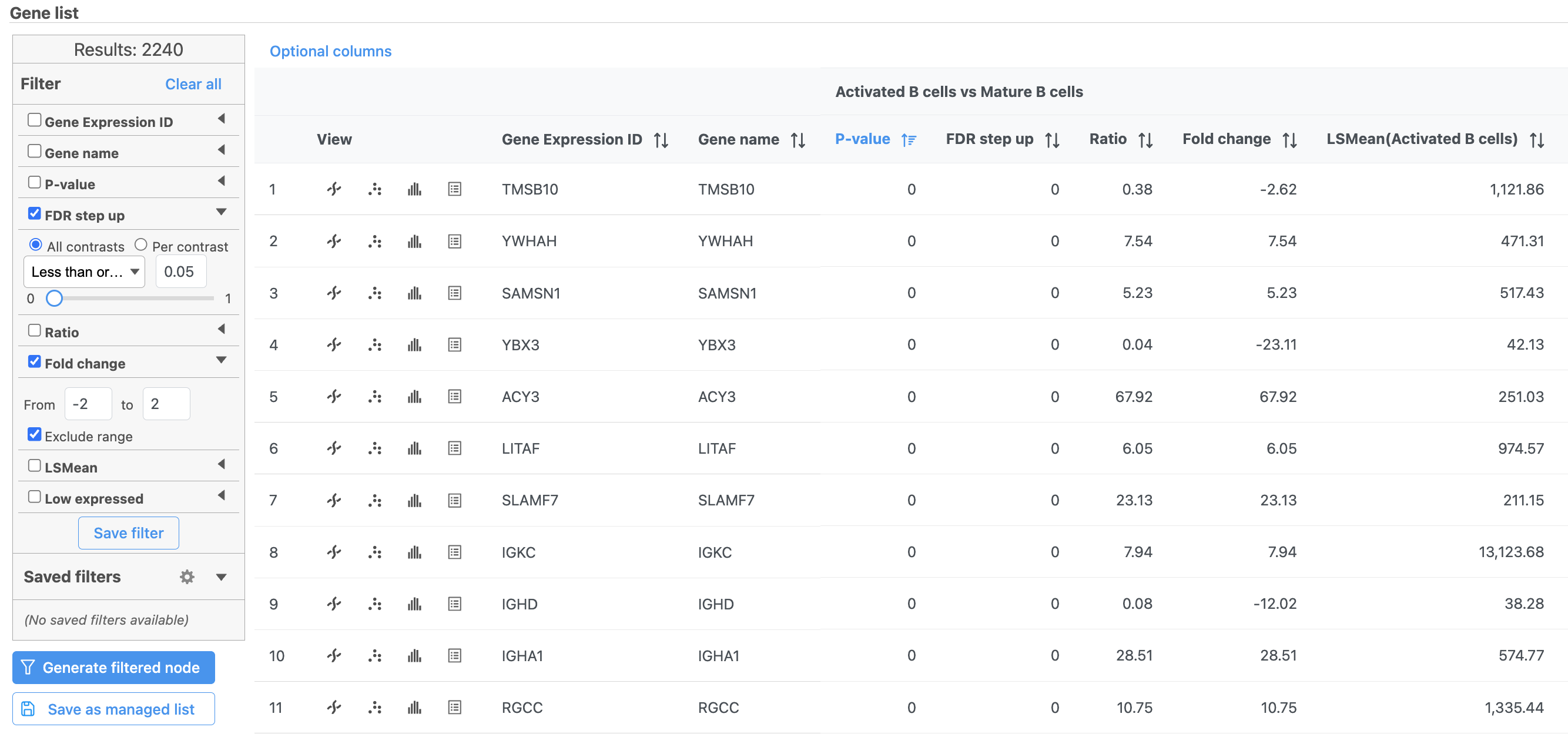 Image Added Image Added
|
- Click
 Image Removed
Image Removed Image Added to create a new data node including only these significantly different genes
Image Added to create a new data node including only these significantly different genes
...
The pathway enrichment results list KEGG pathways, giving an enrichment score and p-value for each (Figure ?Figure 16).
| Numbered figure captions |
|---|
| SubtitleText | Results of pathway enrichment test |
|---|
| AnchorName | Pathway enrichment analysis results |
|---|
|
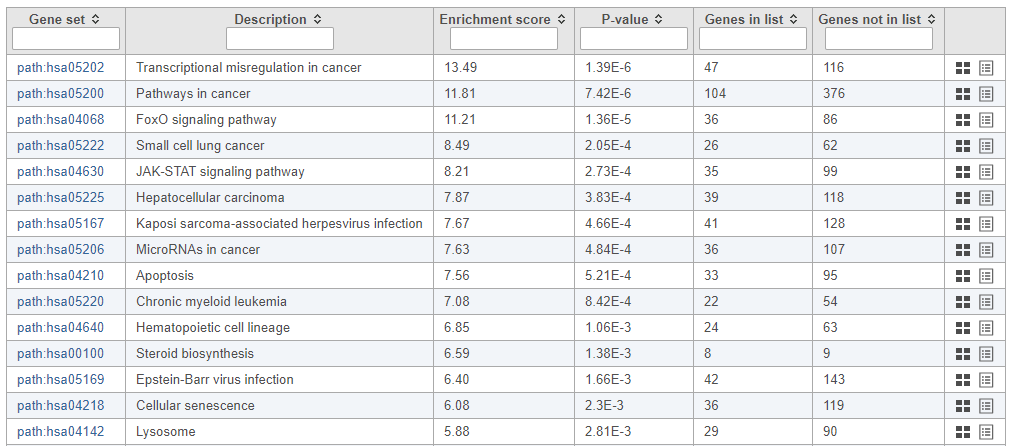 Image Removed Image Removed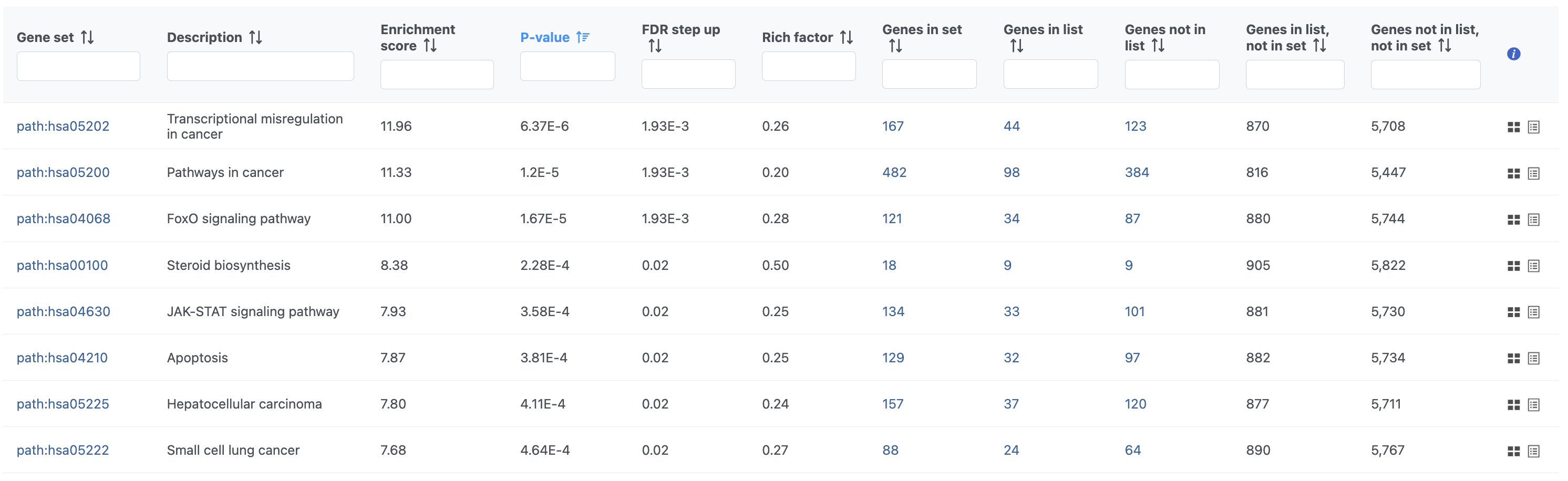 Image Added Image Added
|
To get a better idea about the changes in each enriched pathway, we can view an interactive KEGG pathway map.
...
The KEGG pathway map shows up-regulated genes from the input list in red and down-regulated genes from the input list in green (Figure ?17).
| Numbered figure captions |
|---|
| SubtitleText | Transcriptional misregulation in cancer pathway with significant genes highlighted in green and red |
|---|
| AnchorName | Transcriptional misregulation in cancer |
|---|
|
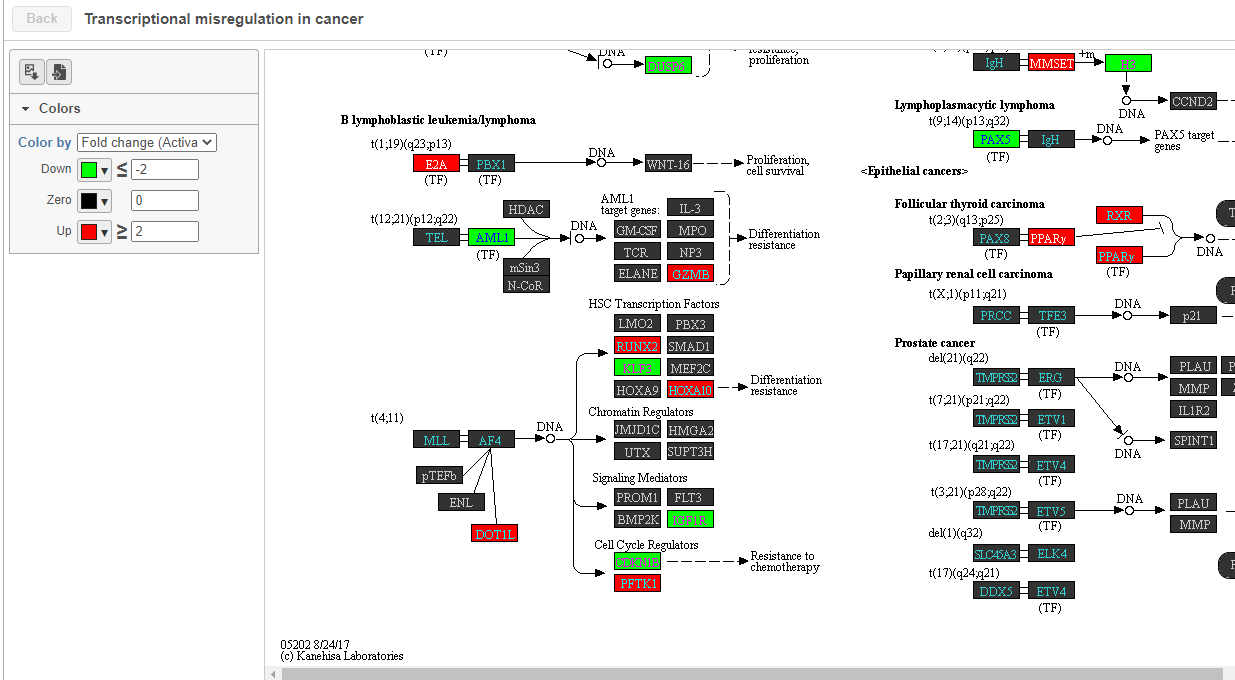 Image Removed Image Removed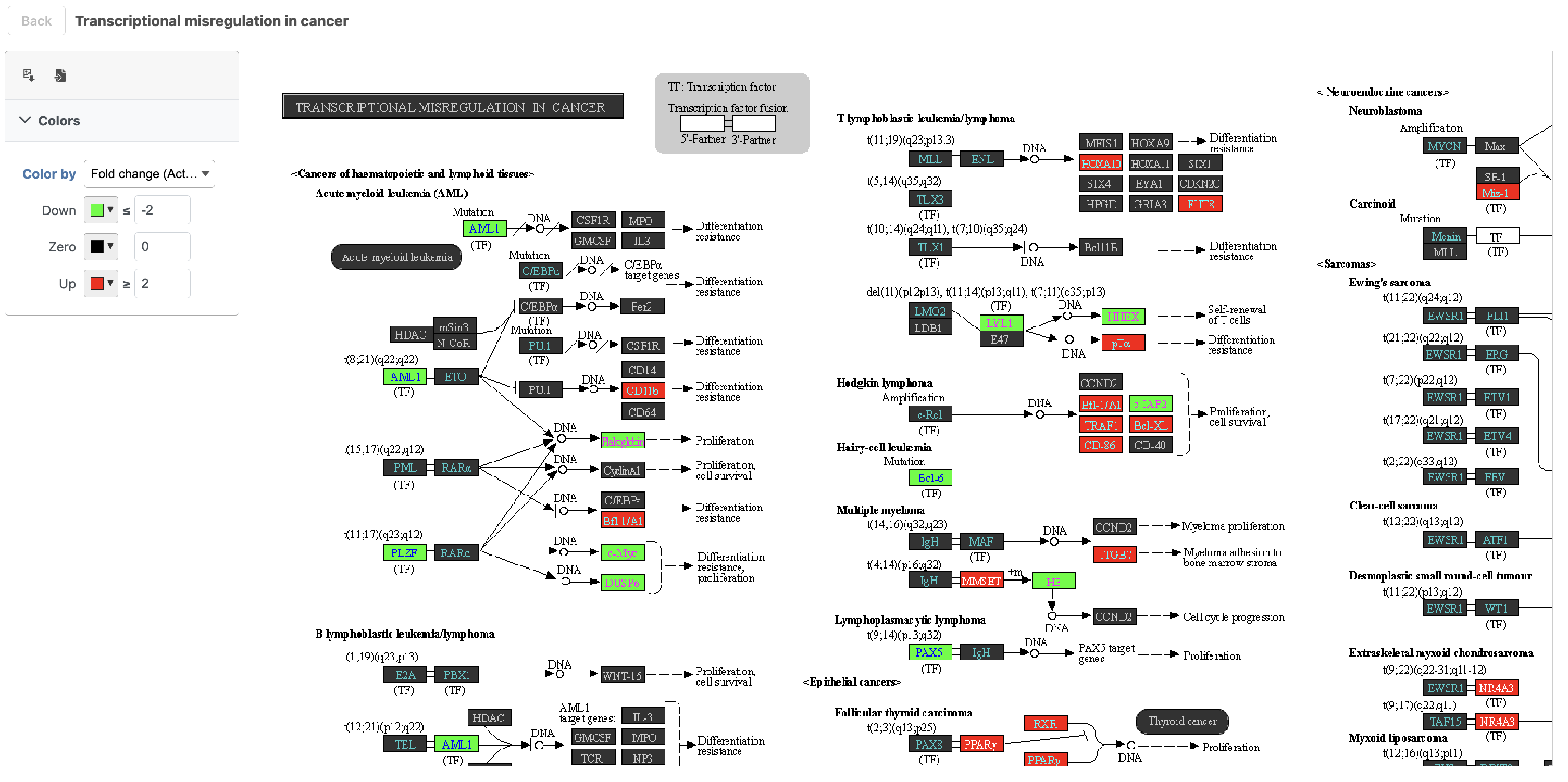 Image Added Image Added
|
| Numbered figure captions |
|---|
| SubtitleText | Final CITE-Seq pipeline |
|---|
| AnchorName | CITE-Seq final pipeline |
|---|
|
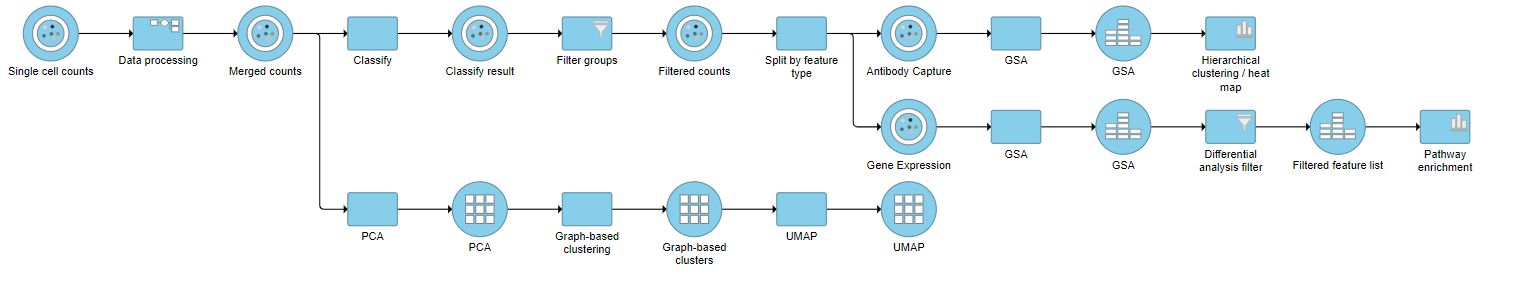 Image Removed Image Removed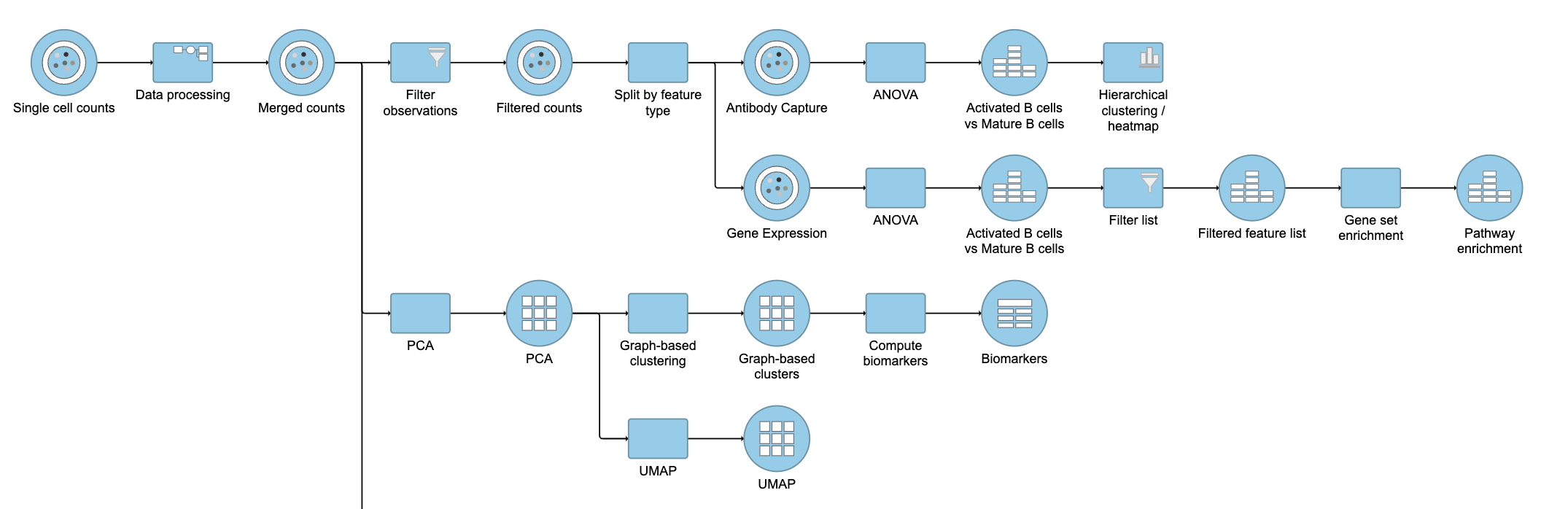 Image Added Image Added
|
...


































