...
| Numbered figure captions |
|---|
| SubtitleText | Invoking the Trim bases task |
|---|
| AnchorName | Trim bases |
|---|
|
 Image Removed Image Removed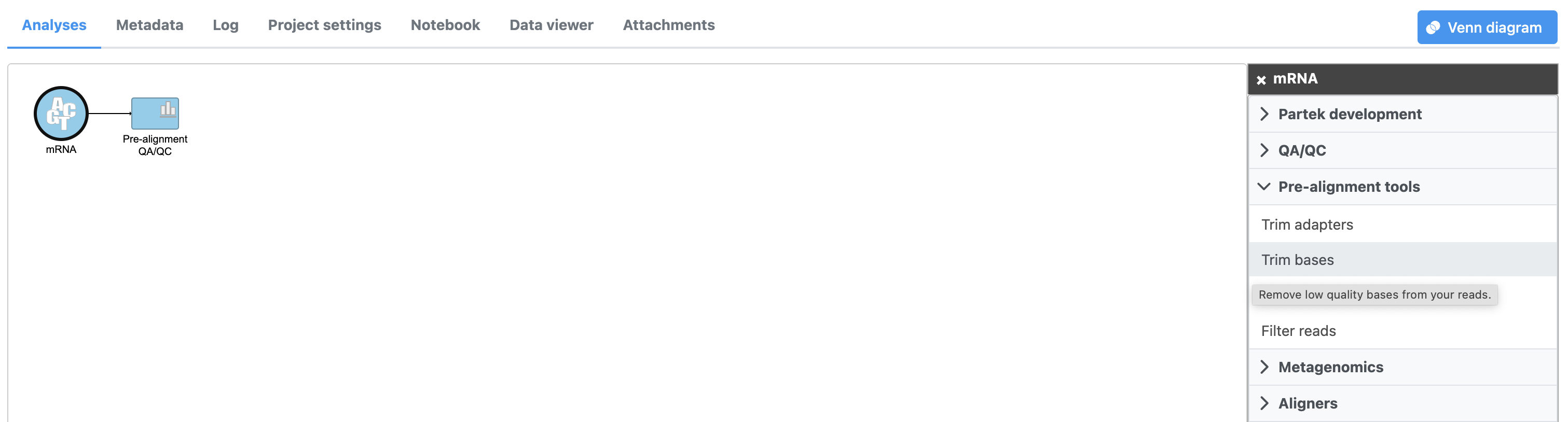 Image Added Image Added
|
By default, Trim bases removes bases removes bases starting at the 3' end and continuing until it finds a base pair call with a Phred score of equal to or greater than 20. Hover over the  Image Removed for additional information on any task option 35 (Figure 2).
Image Removed for additional information on any task option 35 (Figure 2).
- Click Finish to run Trim bases with default settings
| Numbered figure captions |
|---|
| SubtitleText | Viewing additional information about the Quality score trimming methodConfiguring Trim bases |
|---|
| AnchorName | Configuring Trim bases |
|---|
|
 Image Removed Image Removed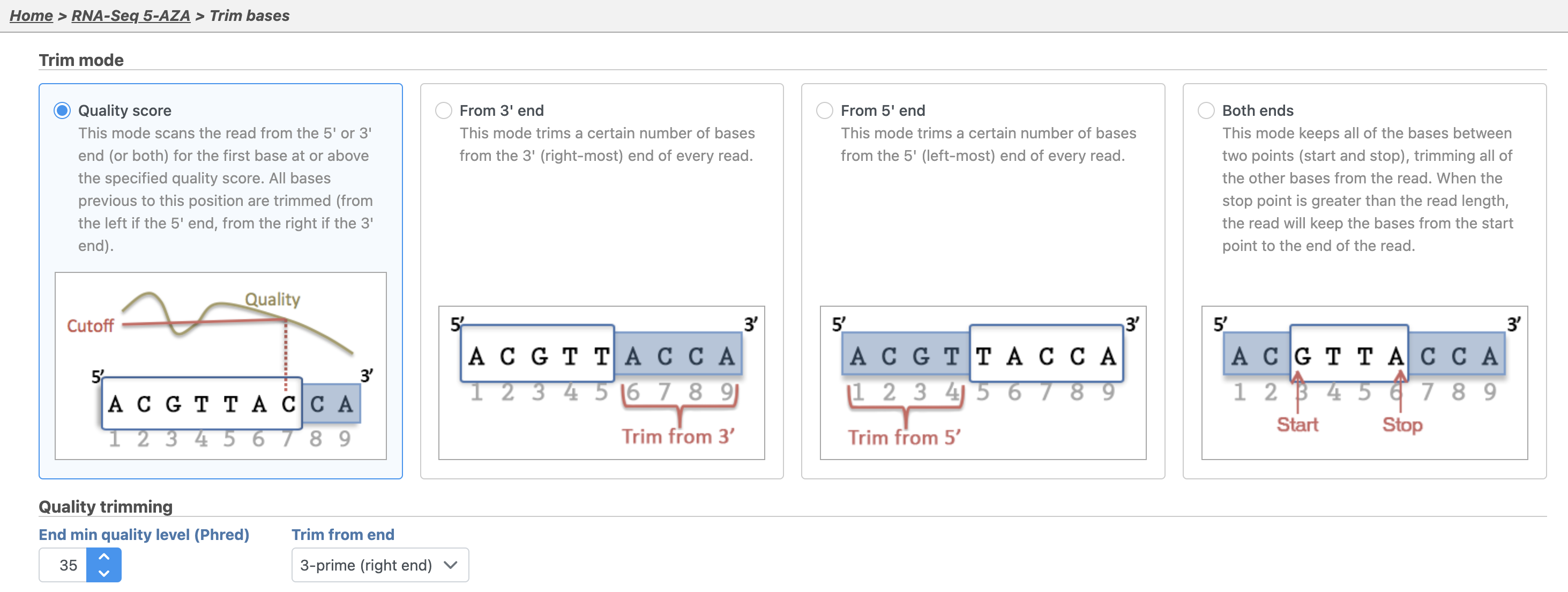 Image Added Image Added
|
The Trim bases task will generate a new data node, Trimmed reads (Figure 3). We can view the task report for Trim bases by double-clicking either the Trim bases task node or the Trimmed reads data node or choosing Task report from the task menu.
| Numbered figure captions |
|---|
| SubtitleText | A Task and a Data node are created from the Trim bases task. Task and Data nodes that have been queued or are in progress are shown in a lighter color than completed tasks. |
|---|
| AnchorName | Queued task node |
|---|
|
|
...
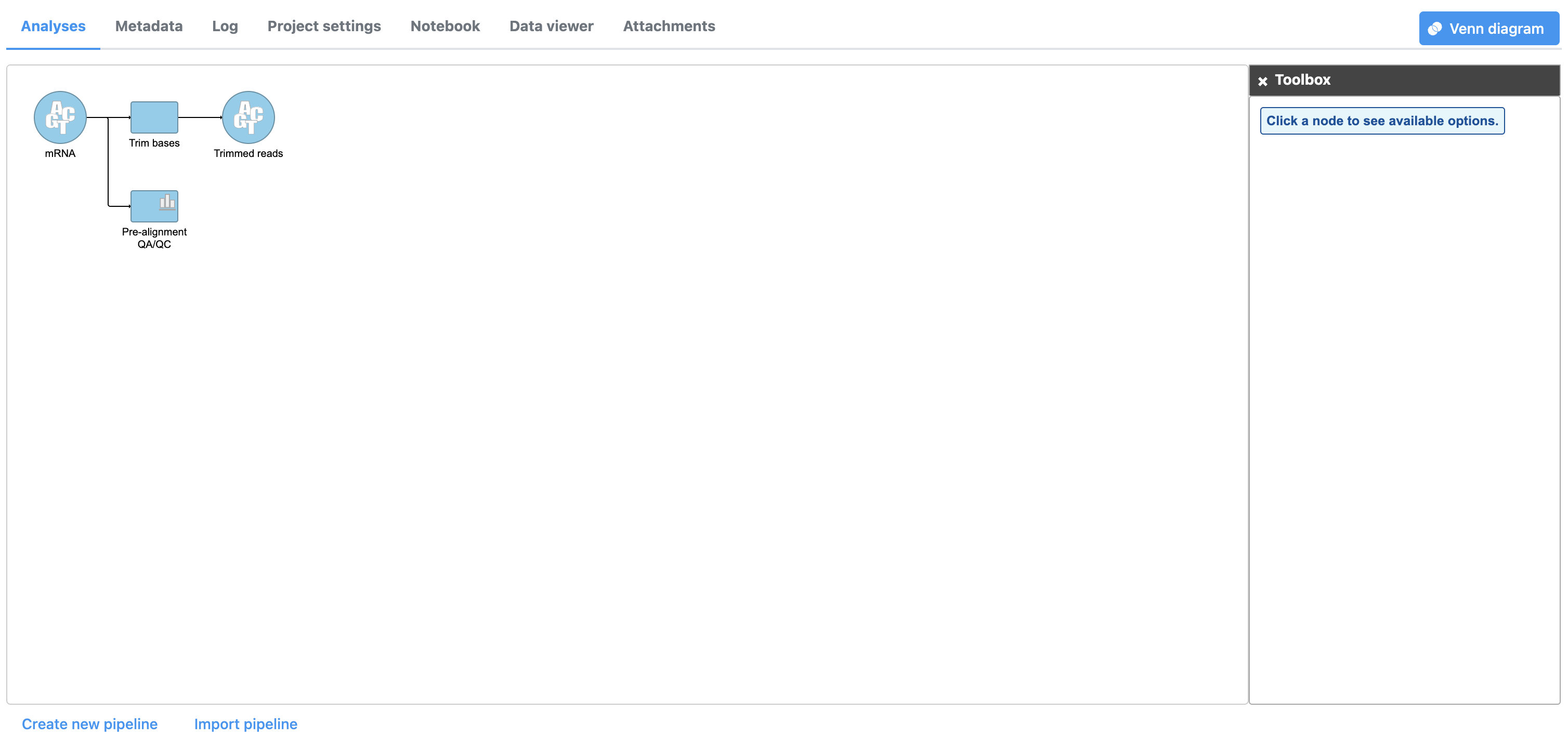 Image Added Image Added
|
- Double-click the Trimmed reads data node to open the task report
The report shows the percentage of trimmed reads and reads removed in a spreadsheet and a two graphs (Figure 4).
| Numbered figure captions |
|---|
| SubtitleText | Results of the Trim bases task |
|---|
| AnchorName | Trim bases report |
|---|
|
 Image Removed Image Removed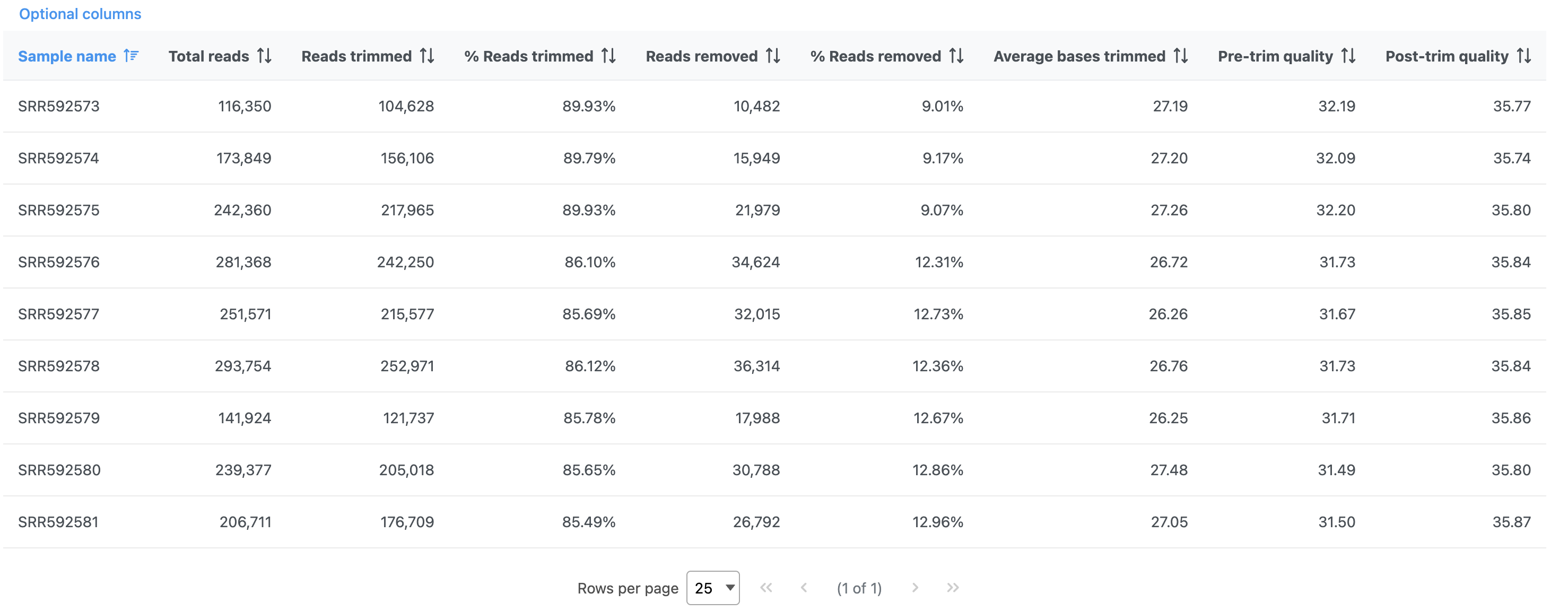 Image Added Image Added
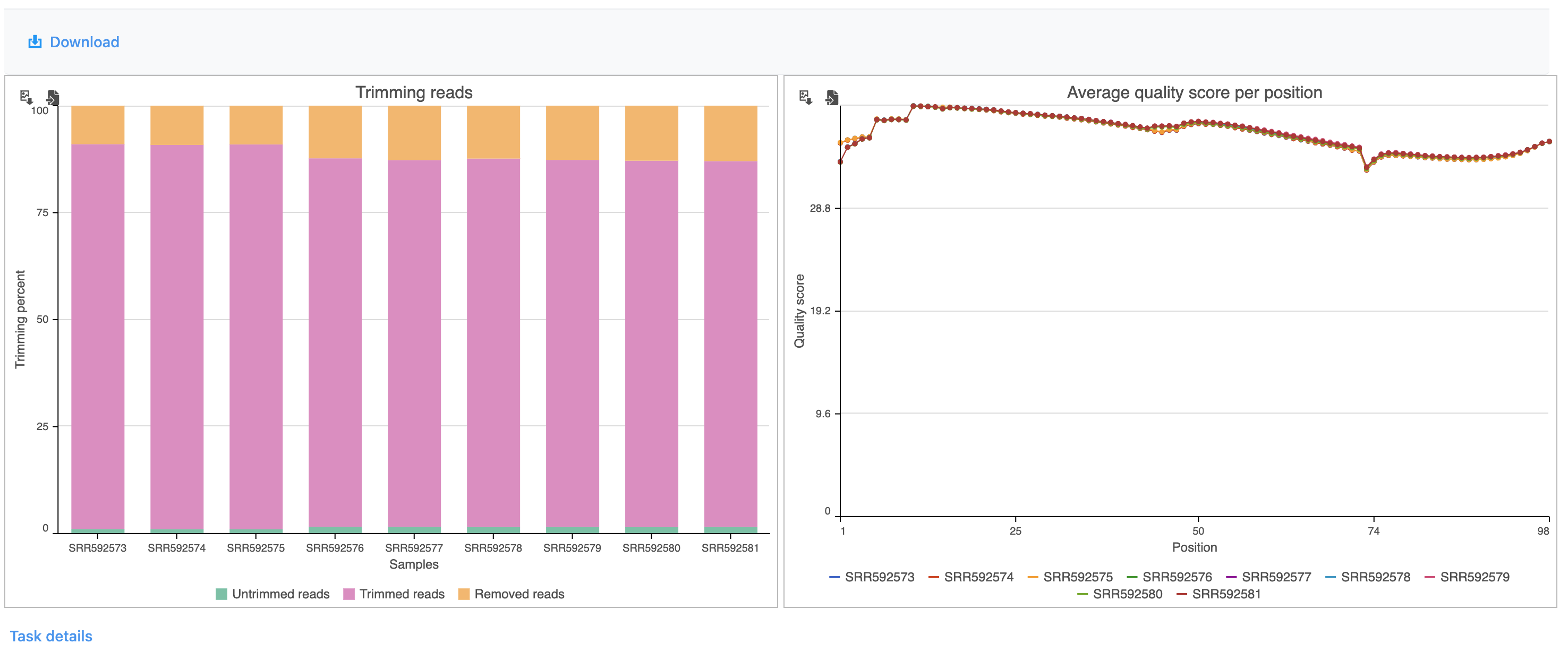 Image Added Image Added
|
The results are fairly consistent across samples with ~65% ~2% of reads untrimmed, ~30% ~86% trimmed, and ~5% ~12% removed for each. The average quality score for each sample is increased with higher average quality scores at the 3' ends.
- Click RNA-Seq 5-AZA to return to the Analyses tab
|
...








