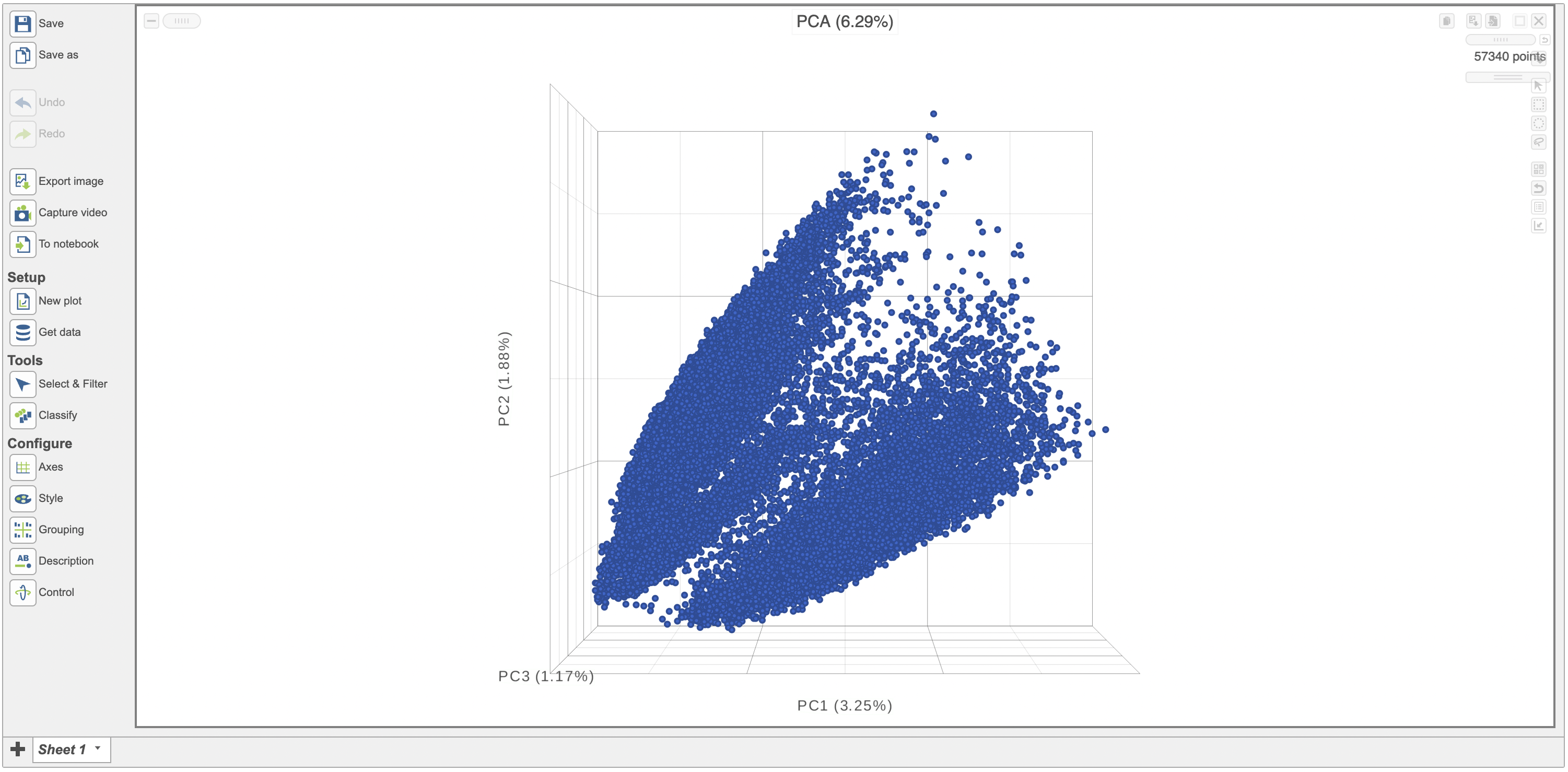
After performing exploratory analyses such as PCA, UMAP and t-SNE is is helpful to visualize the results on a scatterplotscatter plot. This can help visually assess the source of variation affecting the results of an experiment, classify cells and select samples for downstream analysis. Here we have a PCA scatterplot scatter plot generated from the analysis of 12 samples from a scRNA sequencing study. The first three most informative PCs are plotted by default and the percentage of variation explained is stated next to each one of them.
| Numbered figure captions |
|---|
| SubtitleText | Example of a 3D PCA scatterplot . |
|---|
| AnchorName | Example of a 3D PCA scatterplot |
|---|
|
|
...
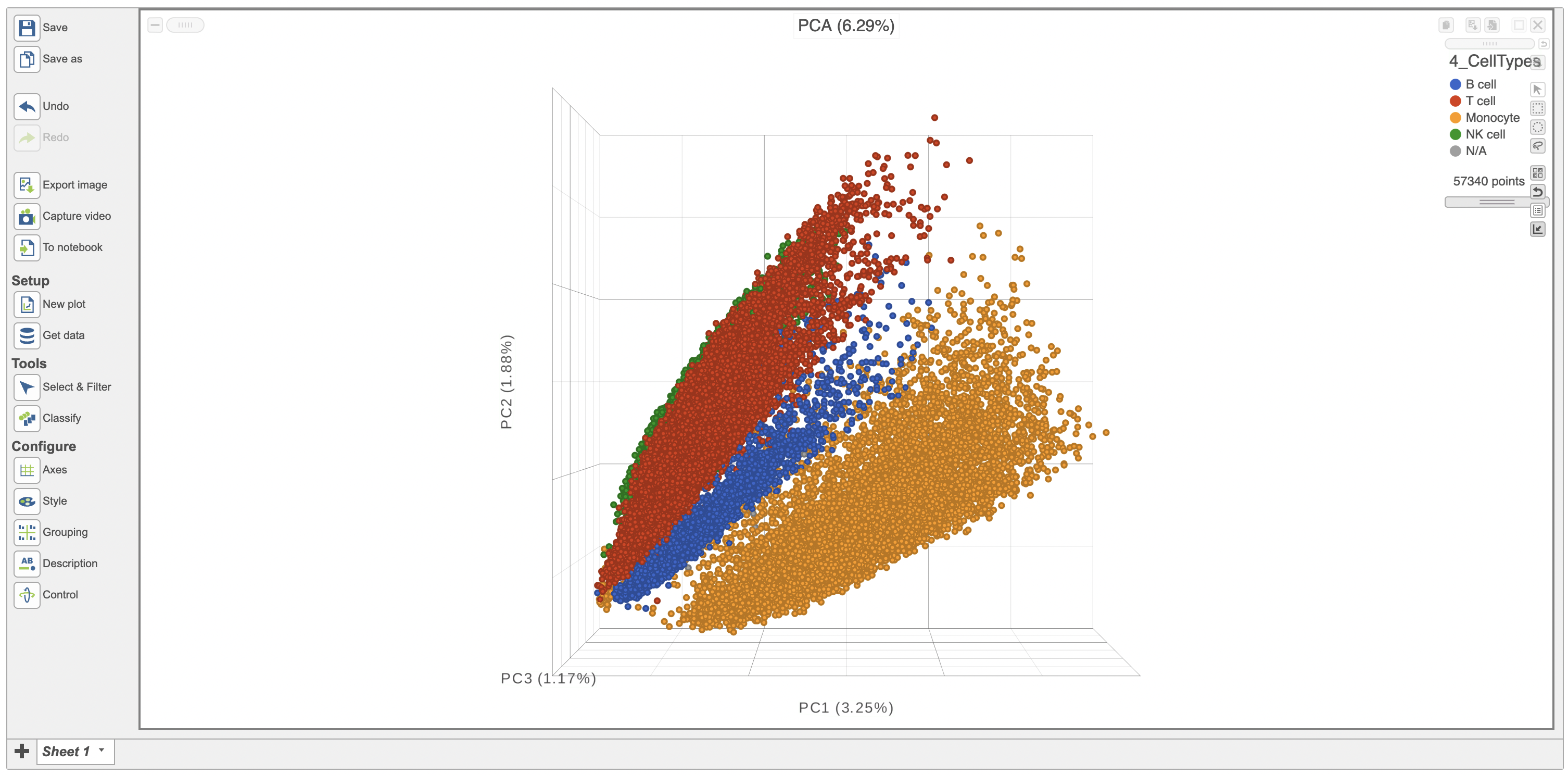
The Configure > Style menu on the left can then be used to color the features in the scatterplot scatter plot based on an attribute (Figure 2). In this case, Figure 3 shows the cells being colored based on their cell-type.
...
| Numbered figure captions |
|---|
| SubtitleText | Customization menu. |
|---|
| AnchorName | Customization menu |
|---|
|
 Image Removed Image Removed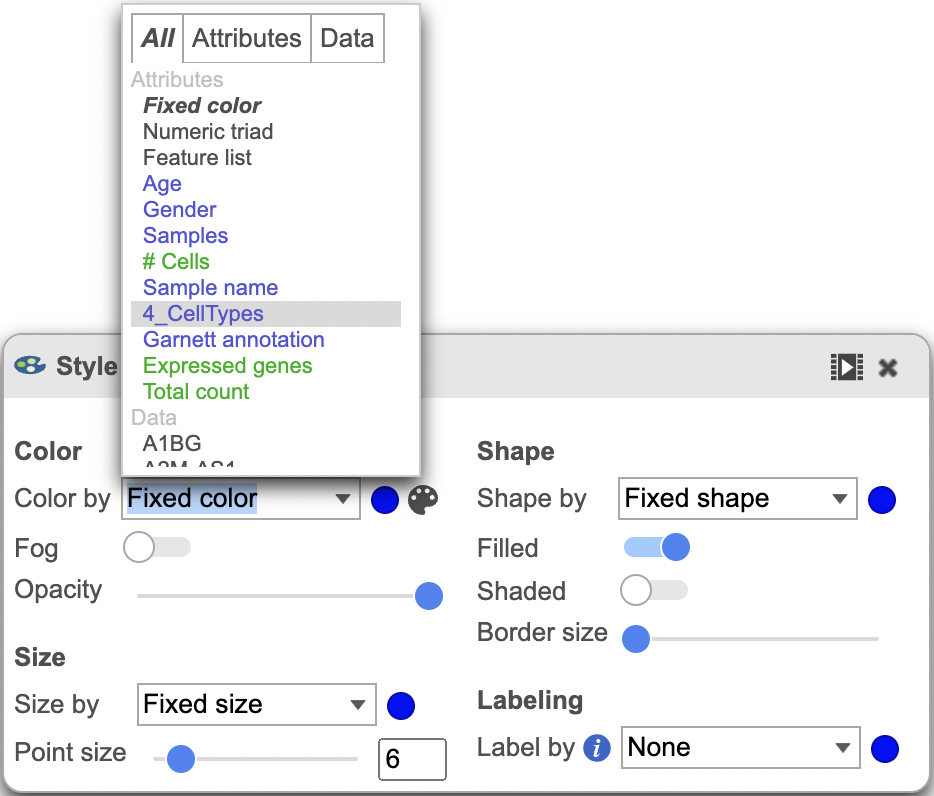 Image Added Image Added
|
| Numbered figure captions |
|---|
| SubtitleText | PCA scatterplot colored by cell-type. |
|---|
| AnchorName | PCA scatterplot colored by cell-type. |
|---|
|
|
...



