| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| minLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
Hierarchical Clustering
The gene list in spreadsheet Down_Syndrome_vs_Normal (A) can be used for hierarchical clustering to visualize patterns in the data.
- Under the Visualization section in the Gene Expression workflow, select Cluster Based on Significant Genes(Figure 1)
| Numbered figure captions |
|---|
| SubtitleText | Select cluster based on significant genes from the Visualization panel of the Gene Expression workflow |
|---|
| AnchorName | Selecting Clustering |
|---|
|
 Image Removed Image Removed
|
The Cluster Significant Genes dialog asks you to specify the type of clustering you want to perform.
...
- Choose Hierarchical Clustering and select OKselect OK
- Choose the Down_Syndrome_vs_Normal (A) spreadsheet under the Spreadsheet with differentially expressed genes
- Choose the Standardize – shift genes to mean of zero and scale to standard deviation of one under the Expression normalization panel . (Figure 1)
This option will adjust all the gene intensities such that the mean is zero and the standard deviation is 1.
| Numbered figure captions |
|---|
| SubtitleText | Configuring Hierarchical Clustering |
|---|
| AnchorName | Hierarchical Clustering options |
|---|
|
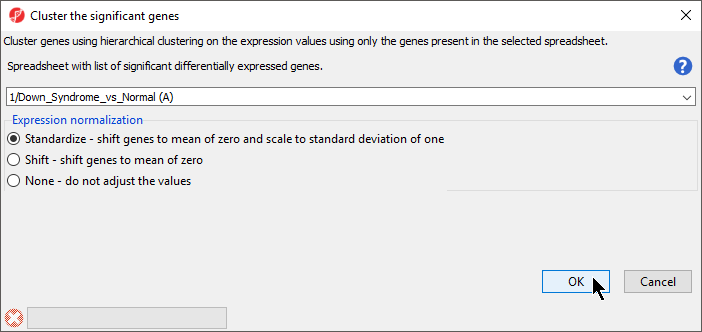 Image Added Image Added
|
- Select OK to generate a Hierarchical Clustering tab (Figure 2)
| Numbered figure captions |
|---|
| SubtitleText | Hierarchical Clustering of Down_Syndrome_vs_Normal (A) |
|---|
| AnchorName | Hierarchical Clustering |
|---|
|
...
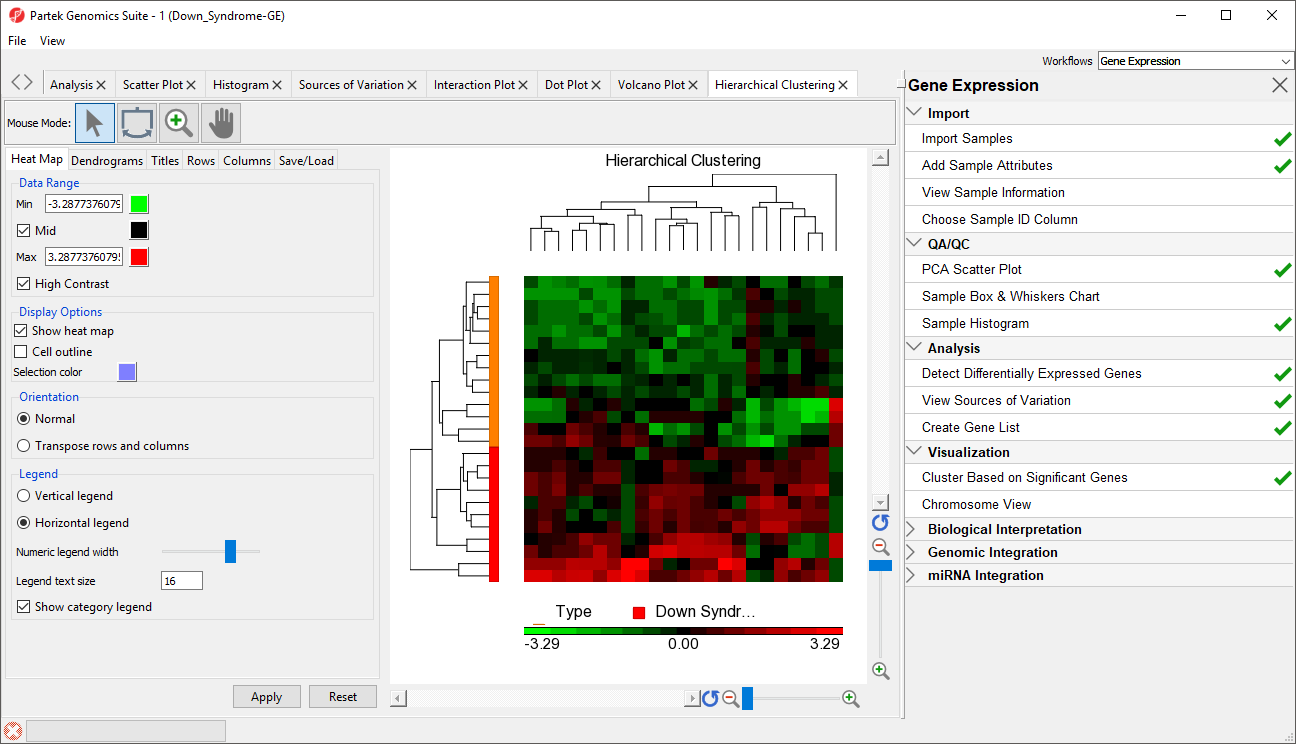 Image Added Image Added
|
The graph (Figure 2) illustrates the standardized gene expression level of each gene in each sample. Each gene is represented in one column, and each sample is represented in one row. Genes which are unchanged are with no difference in expression have a value of zero and are colored black. Genes with increased expression in Down syndrome samples have positive values and are colored red. Genes with reduced expression in Down syndrome samples have negative values and are colored green. Down syndrome samples are colored red and normal samples are colored orange. On the left-hand side of the graph, we can see that the Down syndrome samples cluster together.
For more information on the methods used for clustering, you can refer to Chapter 8: Hierarchical & Partitioning Clustering in in Help > User’s Manual. For a tutorial on configuring the clustering plot, please refer to the user guide that can be downloaded from: here or from Help >On-line Tutorials > User Guides.
Adding Gene Annotation
During data importation, the GeneChip annotation file was linked to the imported data. This linked annotation information can be added as new columns to the ANOVA or gene list spreadsheets. For example, we can add additional annotation to the gene list we created from the ANOVA results as follows:
...
| Numbered figure captions |
|---|
| SubtitleText | Inserting an annotation |
|---|
| AnchorName | Adding Annotation to Gene List |
|---|
|
 Image Removed Image Removed
|
...
| Numbered figure captions |
|---|
| SubtitleText | Adding Chromosomal Location annotation |
|---|
| AnchorName | Add Rows/Columns to Spreadsheet |
|---|
|
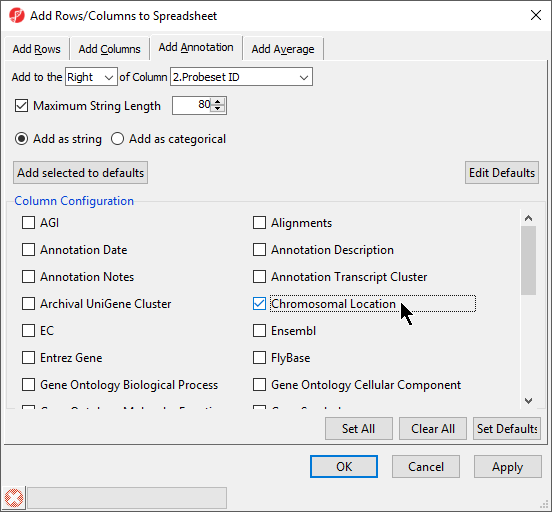 Image Removed Image Removed
|
Interestingly, of the 23 genes of the Down_Syndrome_vs_Normal (A) spreadsheet, 20 genes are located on chromosome 21. To save changes to the spreadsheet, select the Save Active Spreadsheet icon ( Image Removed).
Image Removed)....
| Numbered figure captions |
|---|
| SubtitleText | Dot Plot showing DSCR3 expression levels for each sample |
|---|
| AnchorName | Dot Plot |
|---|
|
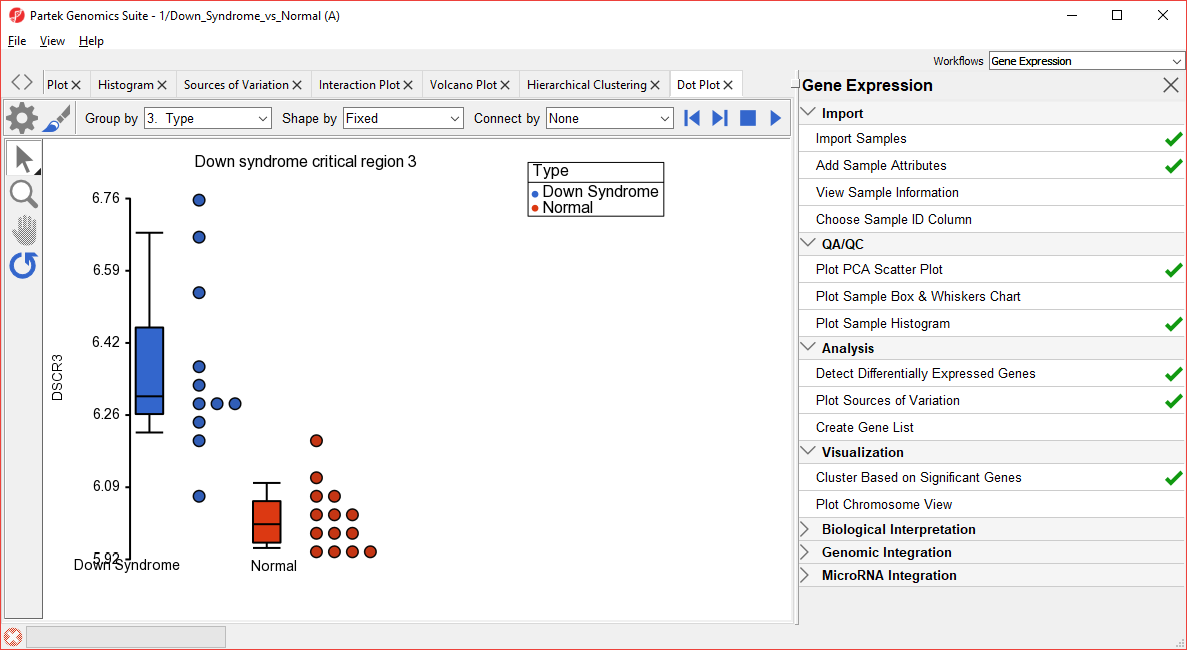 Image Removed Image Removed
|
In the plot, each dot is a sample of the original data. The Y-axis represents the log2 normalized intensity of the gene and the X-axis represents the different types of samples. The median expression of each group is different from each other in this example. The median of the Down syndrome samples is ~6.3, but the median of the normal samples is ~6.0. The line inside the Box & Whiskers represents the median of the samples in a group. Placing the mouse cursor over a Box & Whiskers plot will show its median and range. Hierarchical Clustering Analysis.





