Page History
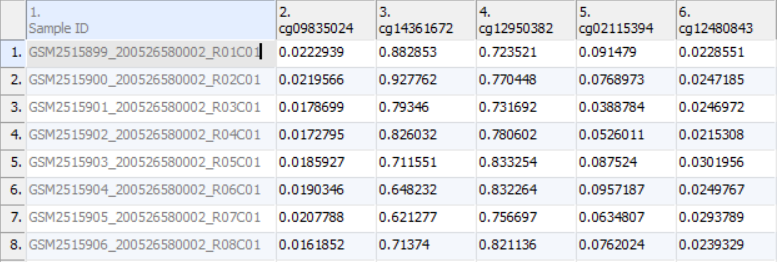
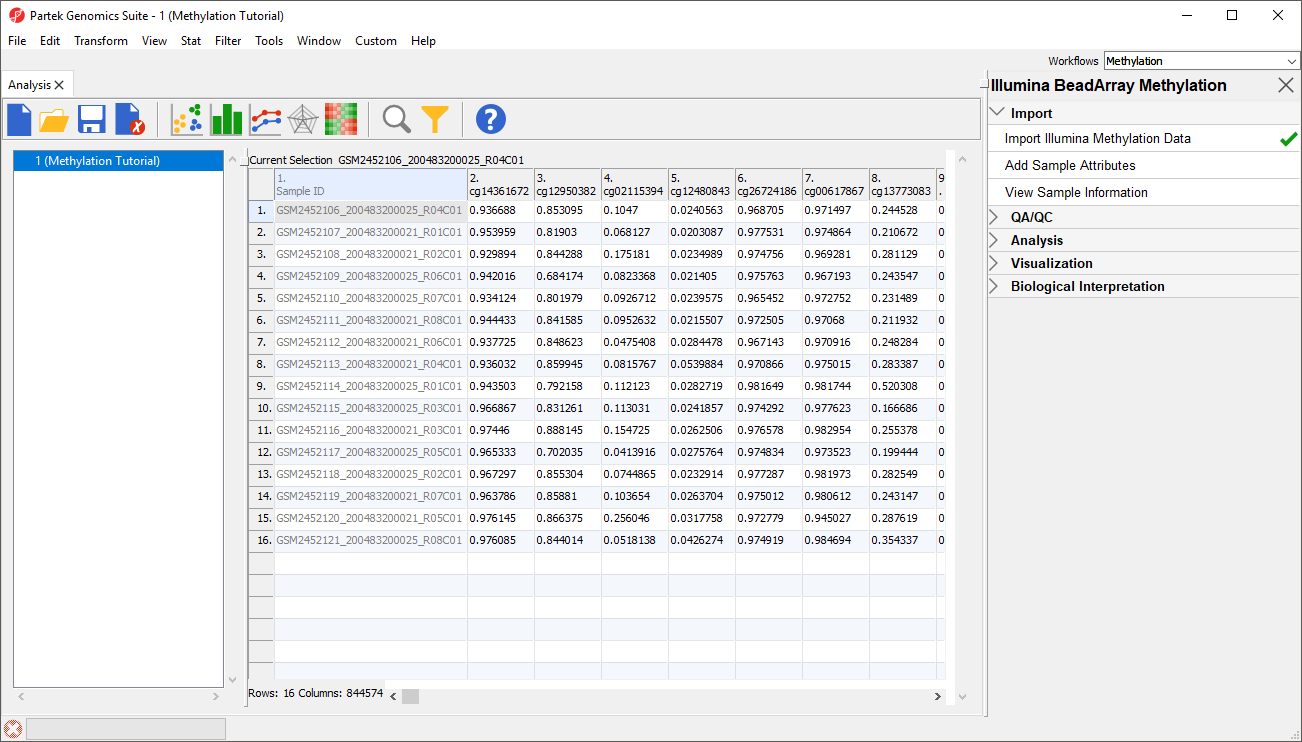
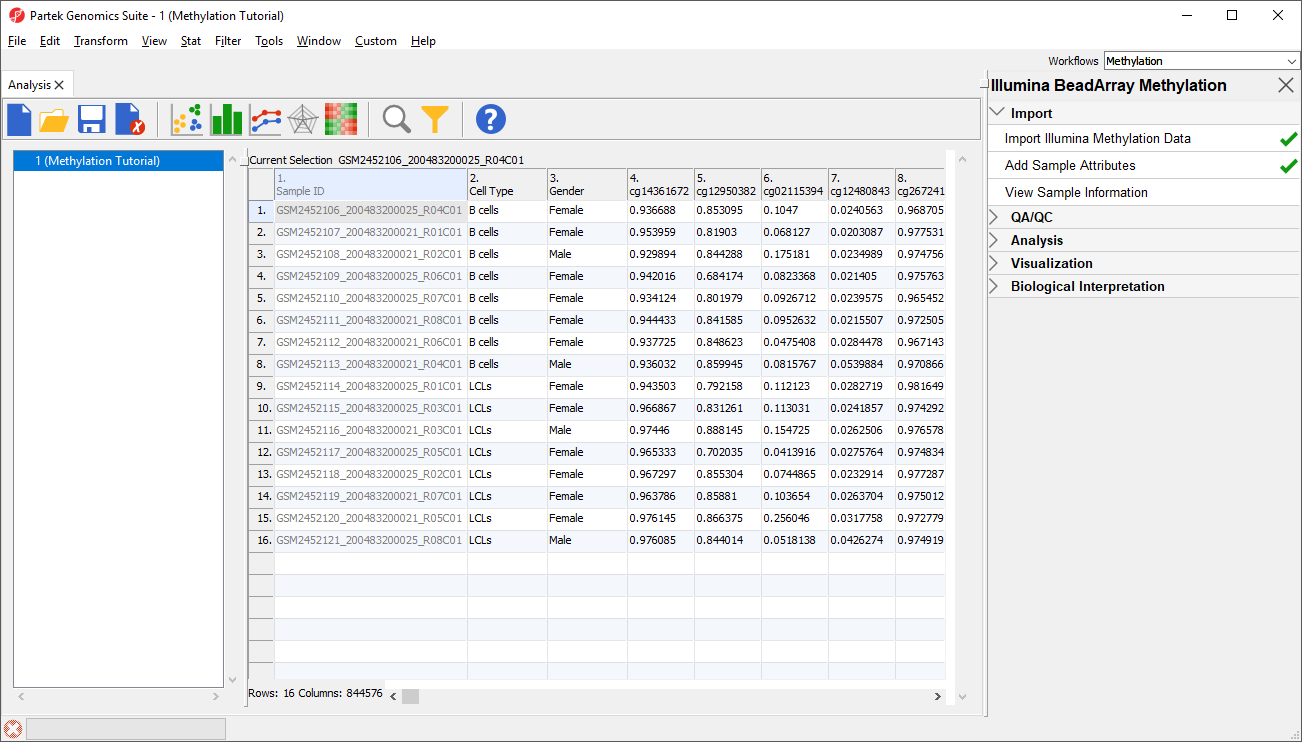
Once the .idat files are imported, you will be facing the Scatter Plot. To move forward with the tutorial, switch to the Analysis tab and take a look at the top level (i.e. initial) Each row of the spreadsheet (Figure 1) . Samples identifiers are corresponds to a single sample. The first column is the names of the .idat files as shown by the Sample ID column while all the and the remaining columns are the array probes.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
As discussed in the previous chapter, the values in the cells are normalized The table values are β-values, which which correspond to the percentage of methylation at each site and are calculated as . A β-value is calculated as the ratio of methylated probe intensity over the overall intensity at each site (the overall intensity is the sum of methylated and unmethylated probe intensities). An alternative metric for measurement of methlyation levels are M-values. β-values can be easily converted to M-values using the following equation:
M-value = log2( β / (1 - β))
Here's the interpretation: a M-value close to 0 indicates a similar intensity between the methylated and unmethylated probes, which means the CpG site is about half-methylated. Positive M-values mean that more molecules are methylated than unmethylated, while negative M-values mean that more molecules are unmethylated than methylated. As discussed by Du and colleagues, the β-value has a more intuitive biological interpretation, but the M-value is more statistically valid for the differential analysis of methylation levels.
Therefore, the differential methylation analysis of the tutorial data will be performed on the M-values; select Convert Beta Value to M Value from the workflow. Note that the original data (β-values) will be overwritten.
Before proceeding to exploratory data
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Before we can perform any analysis, the study samples need to be organized in four groups, two biological replicates each. In the Import section of the workflow, select Add Sample Attributes and, in the next into their experimental groups.
- Select Add Sample Attributes from the Import section of the Illumina BeadArray Methylation workflow
- Select Add a Categorical Attribute from the Add Sample Attributes dialog (Figure 2)
...
Section Heading
Section headings should use level 2 heading, while the content of the section should use paragraph (which is the default). You can choose the style in the first dropdown in toolbar.
...
| Numbered figure captions | ||||
|---|---|---|---|---|
|
...
...
- Select OK
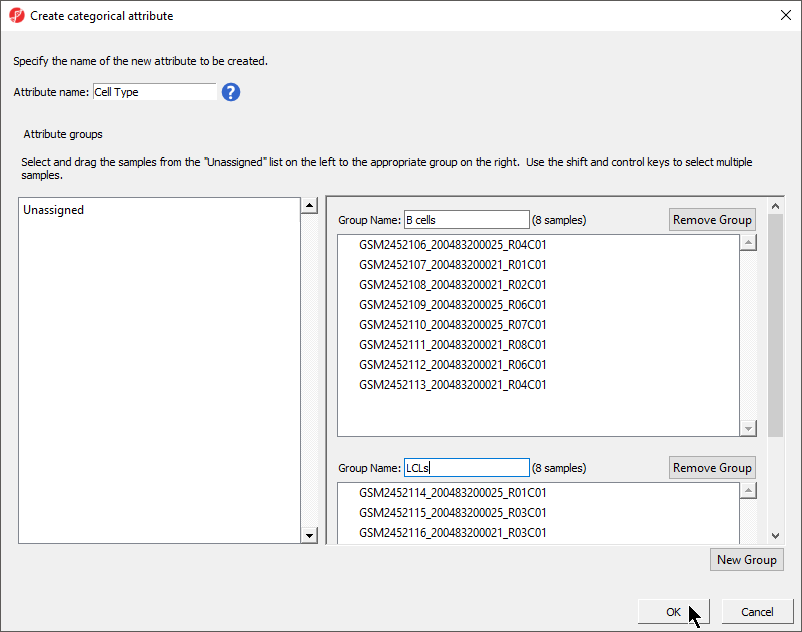
The Create categorical attribute dialog allows us to create groups for a categorical attribute. By default, two groups are created, but additional groups can be added.
- Set Attribute name: to Cell Type
- Rename the groups B cells and LCLs
- Drag and drop the samples from the Unassigned list to their groups as listed in the table below
| Sample ID | Cell Type |
|---|---|
| GSM2452106_200483200025_R04C01 | B cells |
| GSM2452107_200483200021_R01C01 | B cells |
| GSM2452108_200483200021_R02C01 | B cells |
| GSM2452109_200483200025_R06C01 | B cells |
| GSM2452110_200483200025_R07C01 | B cells |
| GSM2452111_200483200021_R08C01 | B cells |
| GSM2452112_200483200021_R06C01 | B cells |
| GSM2452113_200483200021_R04C01 | B cells |
| GSM2452114_200483200025_R01C01 | LCLs |
| GSM2452115_200483200025_R03C01 | LCLs |
| GSM2452116_200483200021_R03C01 | LCLs |
| GSM2452117_200483200025_R05C01 | LCLs |
| GSM2452118_200483200025_R02C01 | LCLs |
| GSM2452119_200483200021_R07C01 | LCLs |
| GSM2452120_200483200021_R05C01 | LCLs |
| GSM2452121_200483200025_R08C01 | LCLs |
There should now be two groups with eight samples in each group (Figure 3).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select OK
- Select Yes from the Add another categorical attribute dialog
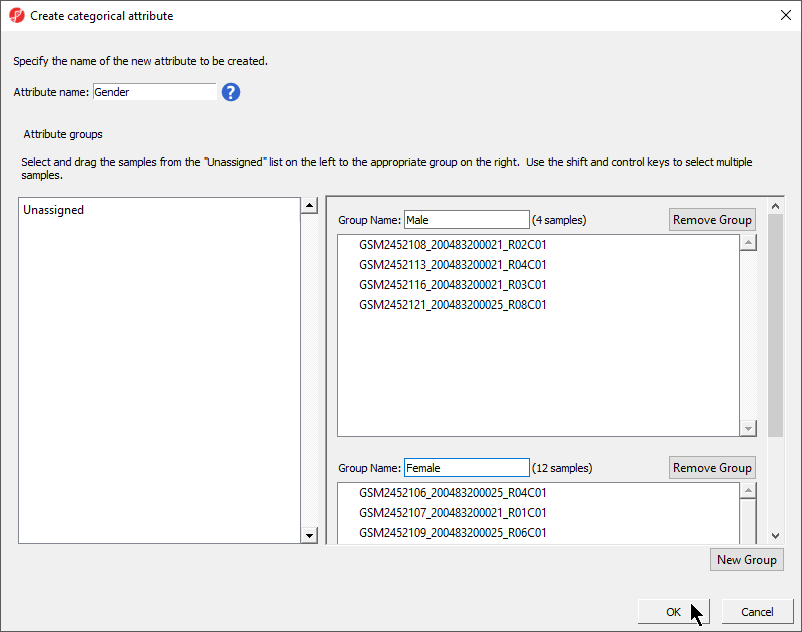
- Set Attribute name: to Gender
- Rename the groups Male and Female
- Drag and drop the samples from the Unassigned list to their groups as listed in the table below
| Sample ID | Gender |
|---|---|
| GSM2452106_200483200025_R04C01 | Female |
| GSM2452107_200483200021_R01C01 | Female |
| GSM2452108_200483200021_R02C01 | Male |
| GSM2452109_200483200025_R06C01 | Female |
| GSM2452110_200483200025_R07C01 | Female |
| GSM2452111_200483200021_R08C01 | Female |
| GSM2452112_200483200021_R06C01 | Female |
| GSM2452113_200483200021_R04C01 | Male |
| GSM2452114_200483200025_R01C01 | Female |
| GSM2452115_200483200025_R03C01 | Female |
| GSM2452116_200483200021_R03C01 | Male |
| GSM2452117_200483200025_R05C01 | Female |
| GSM2452118_200483200025_R02C01 | Female |
| GSM2452119_200483200021_R07C01 | Female |
| GSM2452120_200483200021_R05C01 | Female |
| GSM2452121_200483200025_R08C01 | Male |
There should now be two groups with four samples in Male and twelve samples in Female (Figure 4).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select OK
- Select No from the Add another categorical attribute dialog
- Select Yes to save the spreadsheet
Two new columns have been added to spreadsheet 1 (Methylation) with the cell type and gender of each sample (Figure 5).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
| Page Turner | ||
|---|---|---|
|
| Additional assistance |
|---|
|
| Rate Macro | ||
|---|---|---|
|