Page History
| Table of Contents | ||||||
|---|---|---|---|---|---|---|
|
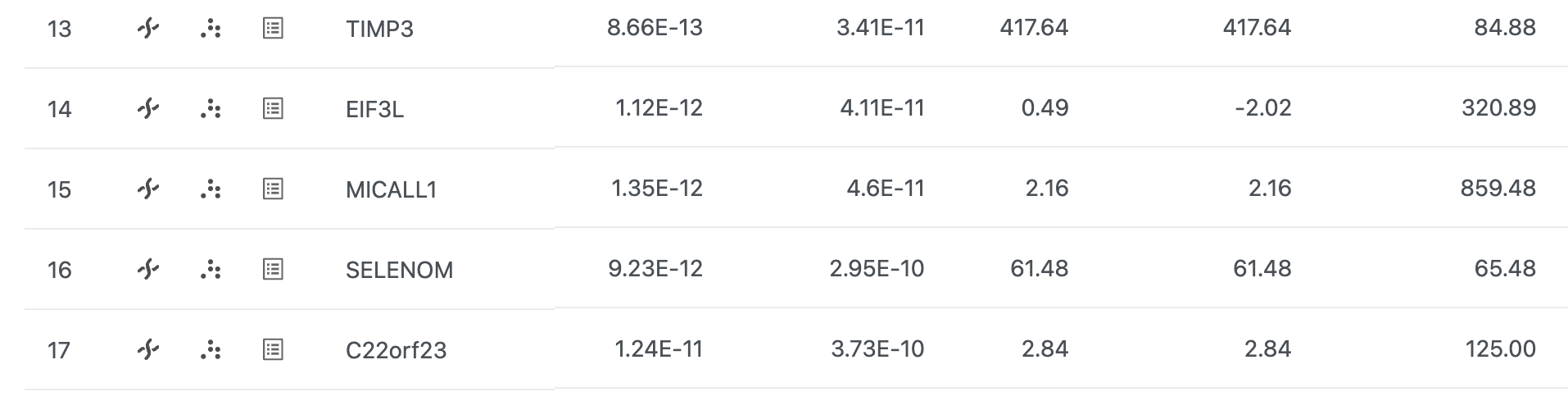
From the GSA the DESeq2 task report, we can browse to any gene in the Chromosome view.
- Select Click in the SELM SELENOM row to open Chromosome view (Figure 1)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
|
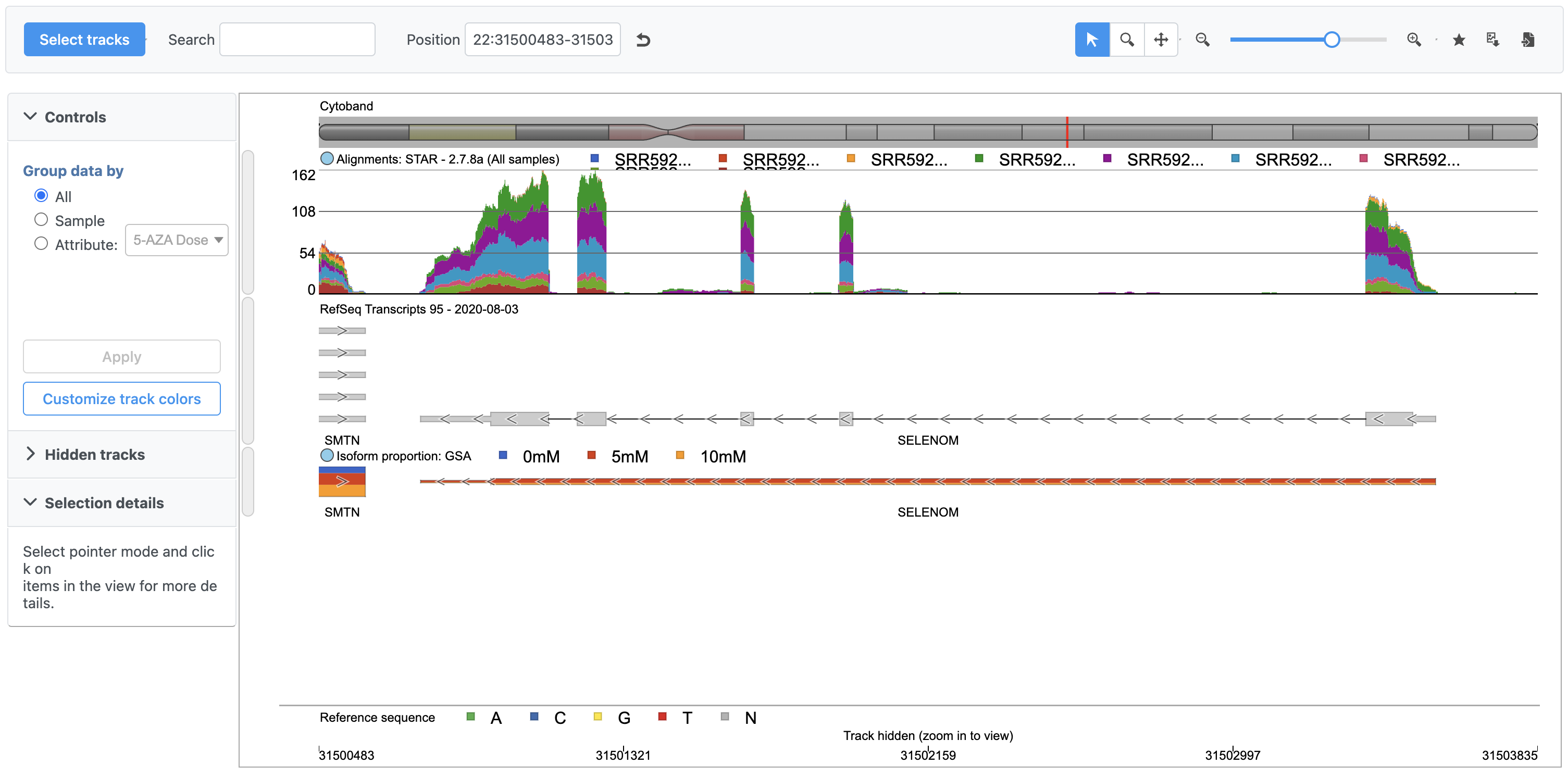
A new tab will open showing SELM SELENOM in the Chromosome view (Figure 2).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Chromosome View shows reference genome, annotation, and data set information together aligned at genomic coordinates.
...
We can add tracks from any data node using Select Tracks.
- Click Select Select tracks
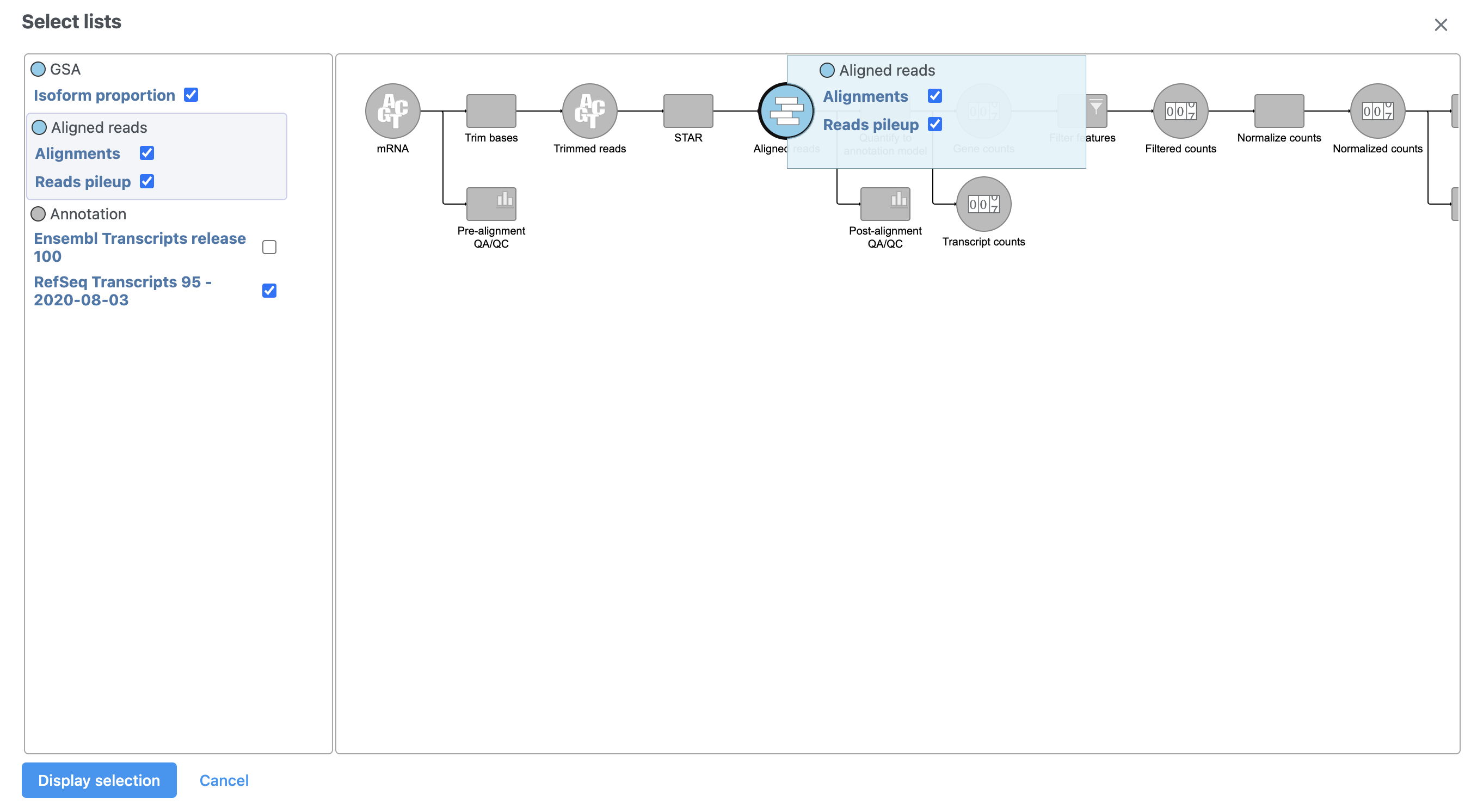
A pop-up dialog showing the pipeline allows us to choose which data to display as tracks in Chromosome view (Figure 3).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select Click Reads pileup under Align Aligned reads on the left-hand side of the dialog
- Select Click Display tracks to make the change
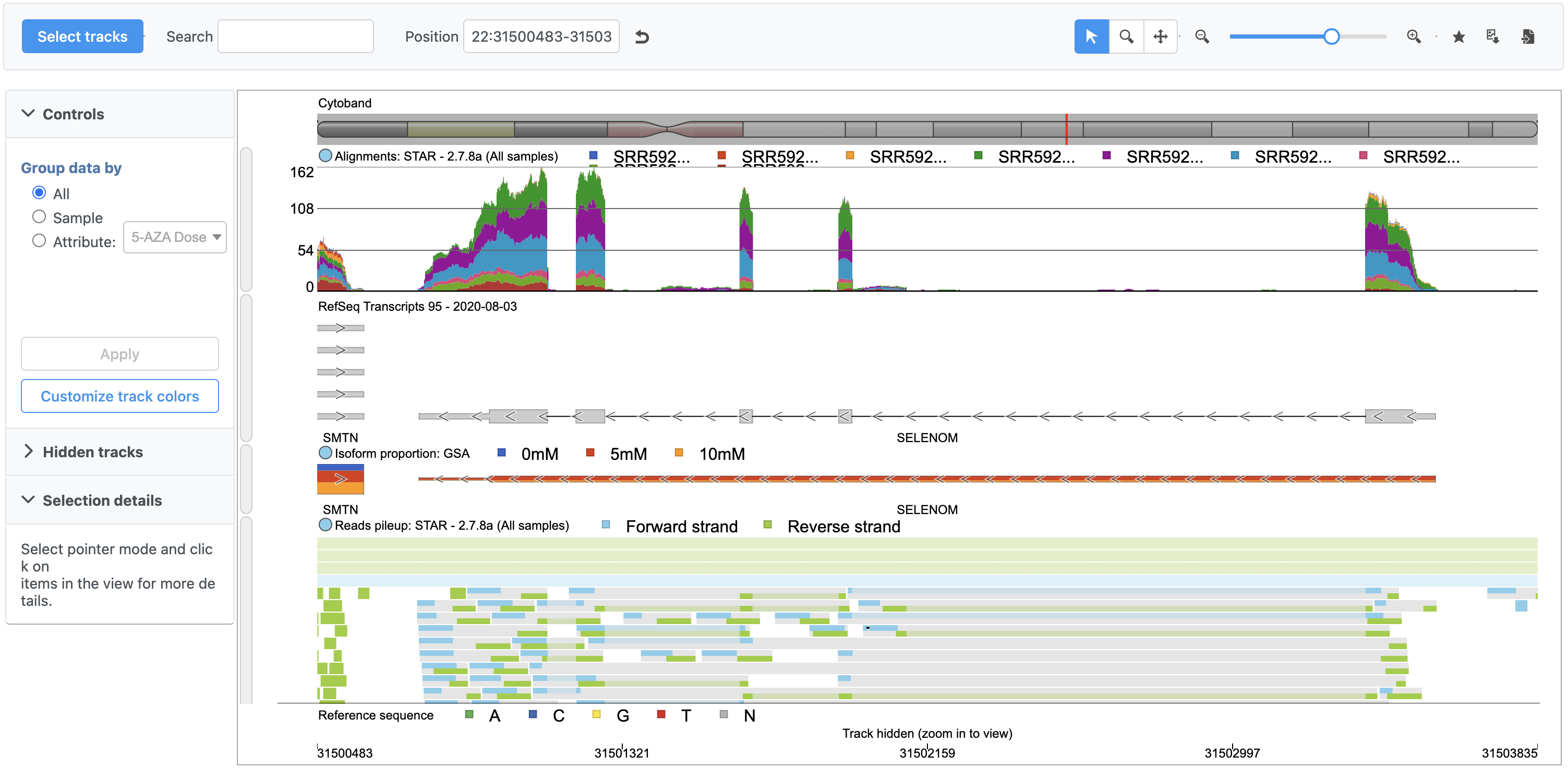
The reads pileup track is now included (Figure 4).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
With multiple tracks, it may be useful to pin a track to the top so we can scroll down the reads pileup track without losing sight of the Alignments or RefSeq tracks.
- Select next to the Alignments track
- Select to pin the track to the top
- Repeat for RefSeq
- Scroll down to view individual reads
Selecting a read brings up detailed information about the read in the selection details panel (Figure 5).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
To learn more about Chromosome view, please consult the Chromosome View user guide.
| Page Turner | ||
|---|---|---|
|
| Additional assistance |
|---|
| Rate Macro | ||
|---|---|---|
|
...