Page History
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
|
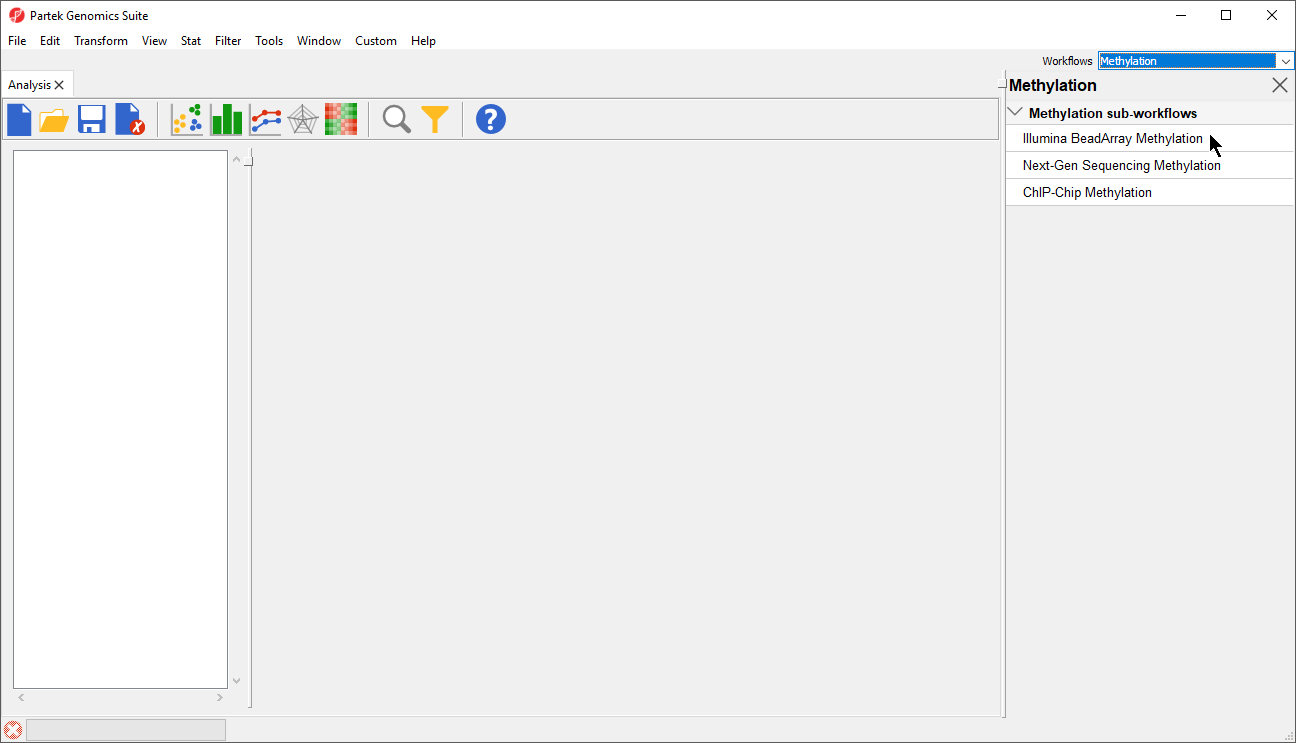
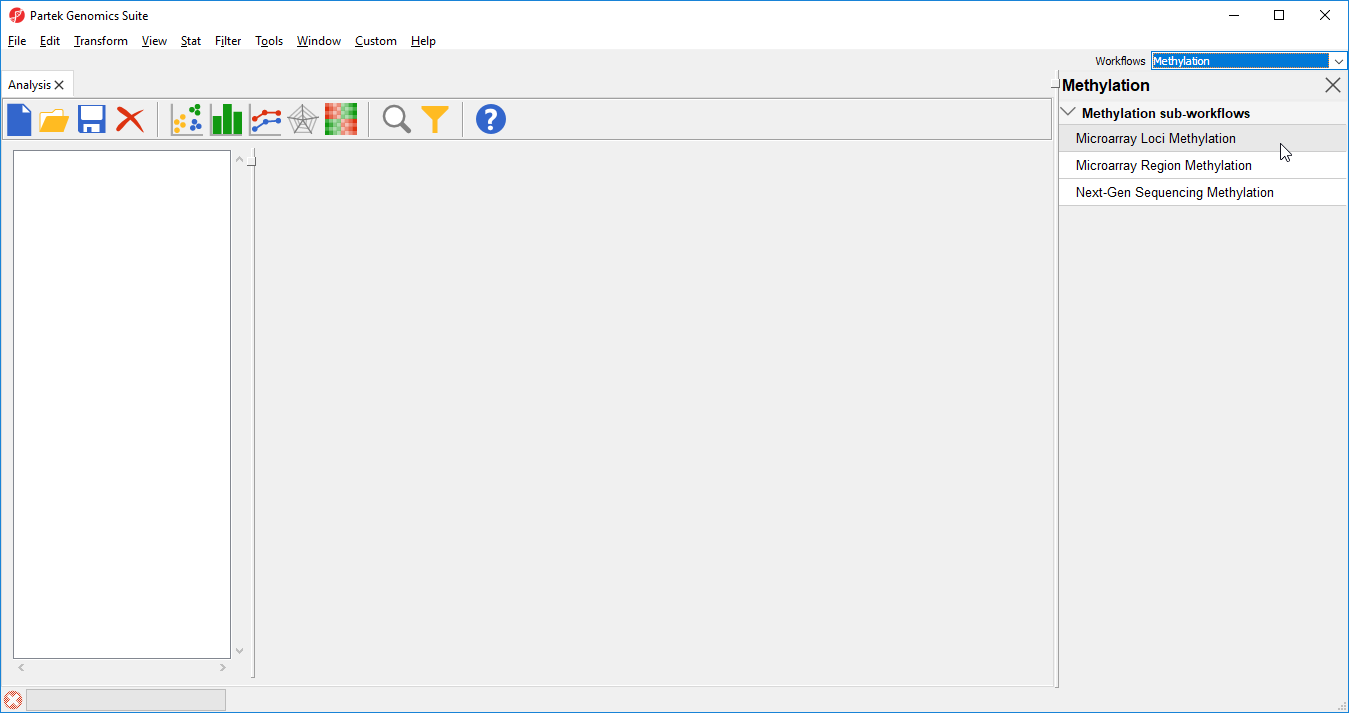
- Select Illumina BeadArray Microarray Loci Methylation from the Methylation sub-workflows panel (Figure 2)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- That will open Illumina BeadArray Methylation workflow (Figure 3)
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
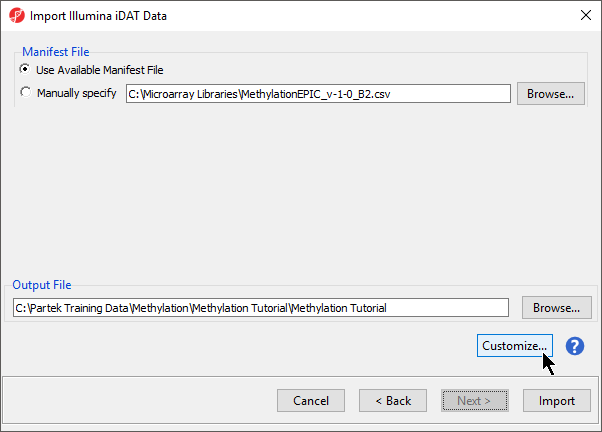
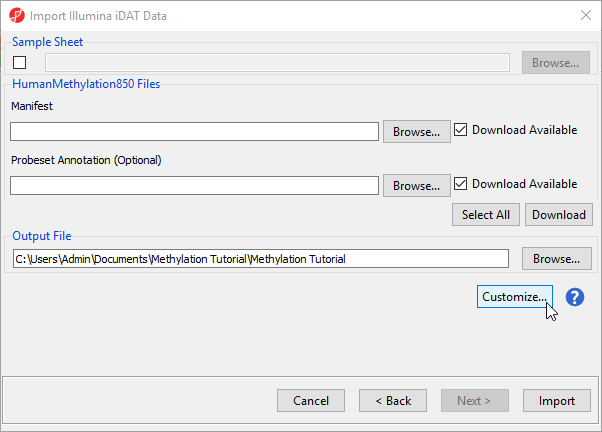
By default the output file destination is set to the file containing your .idat files and the name matches the file folder name. The name and location of the output file can be changed using the Output File panel.
...
- Select Exclude X and Y Chromosomes
Analysis of differentially methylated loci in humans and mice often excludes probes on the X and Y chromosomes because of the difficulties caused by the inactivation of one X chromosome in female samples.
- Select Exclude probes using detection p-value and leave the default settings of 0.05 and 1 out of 16 samples.
...
Overview
Content Tools