Page History
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
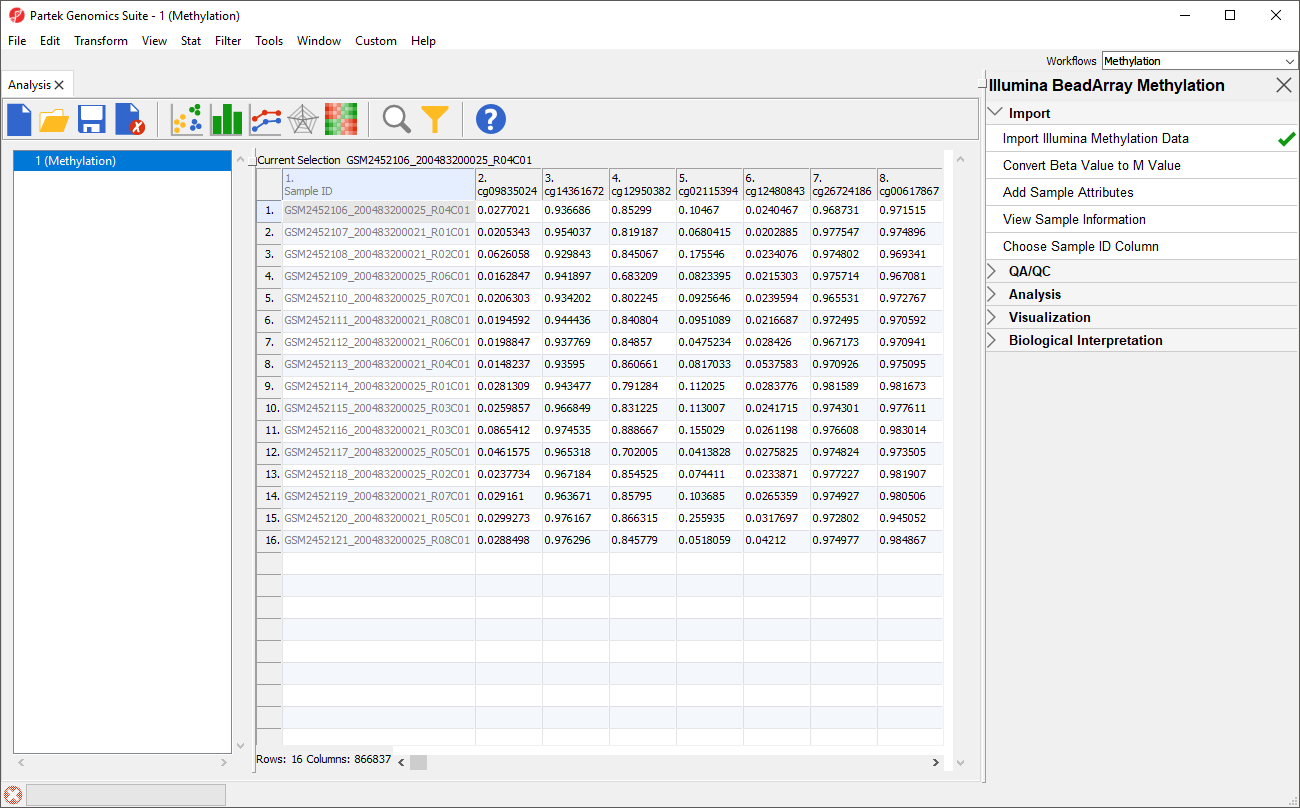
An alternative metric for measurement of methlyation levels are M-values. β-values can be easily converted to M-values using the following equation:
M-value = log2( β / (1 - β))
An M-value close to 0 for a CpG site indicates a similar intensity between the methylated and unmethylated probes, which means the CpG site is about half-methylated. Positive M-values mean that more molecules are methylated than unmethylated, while negative M-values mean that more molecules are unmethylated than methylated. As discussed by Du and colleagues, the β-value has a more intuitive biological interpretation, but the M-value is more statistically valid for the differential analysis of methylation levels.
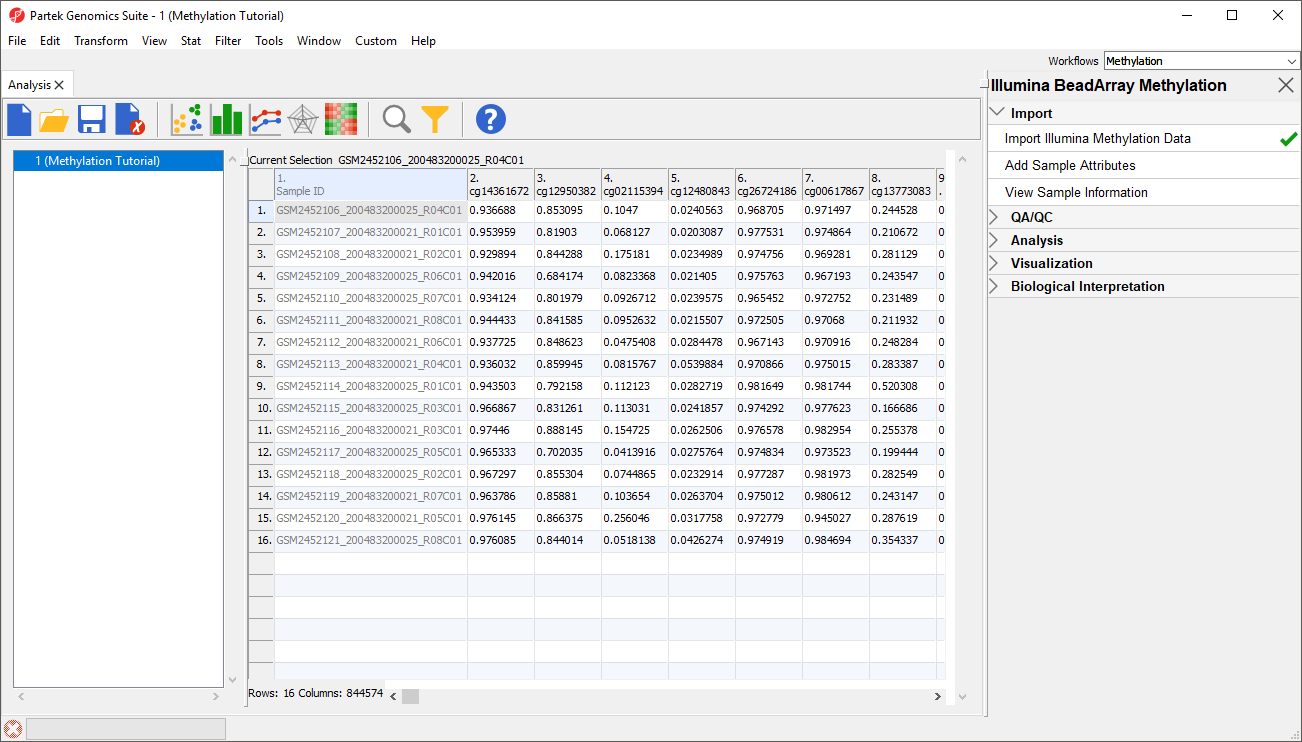
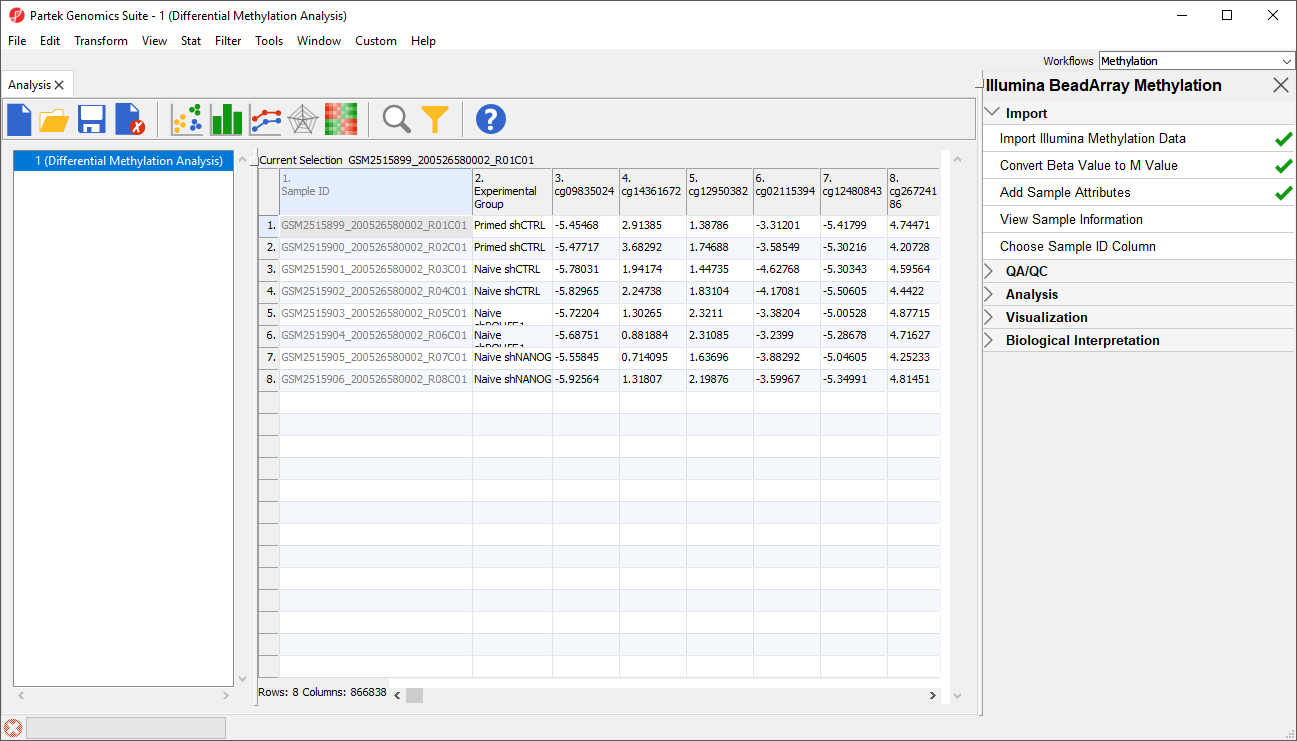
Because we are performing differential methylation analysis, we need to convert our data to from β-values to M-values.
- Select Convert Beta Value to M Value from the Import section of the Illumina BeadArray Methylation workflow
The original data (β-values) will be overwritten.
- Select () from the icon bar to save the current spreadsheet
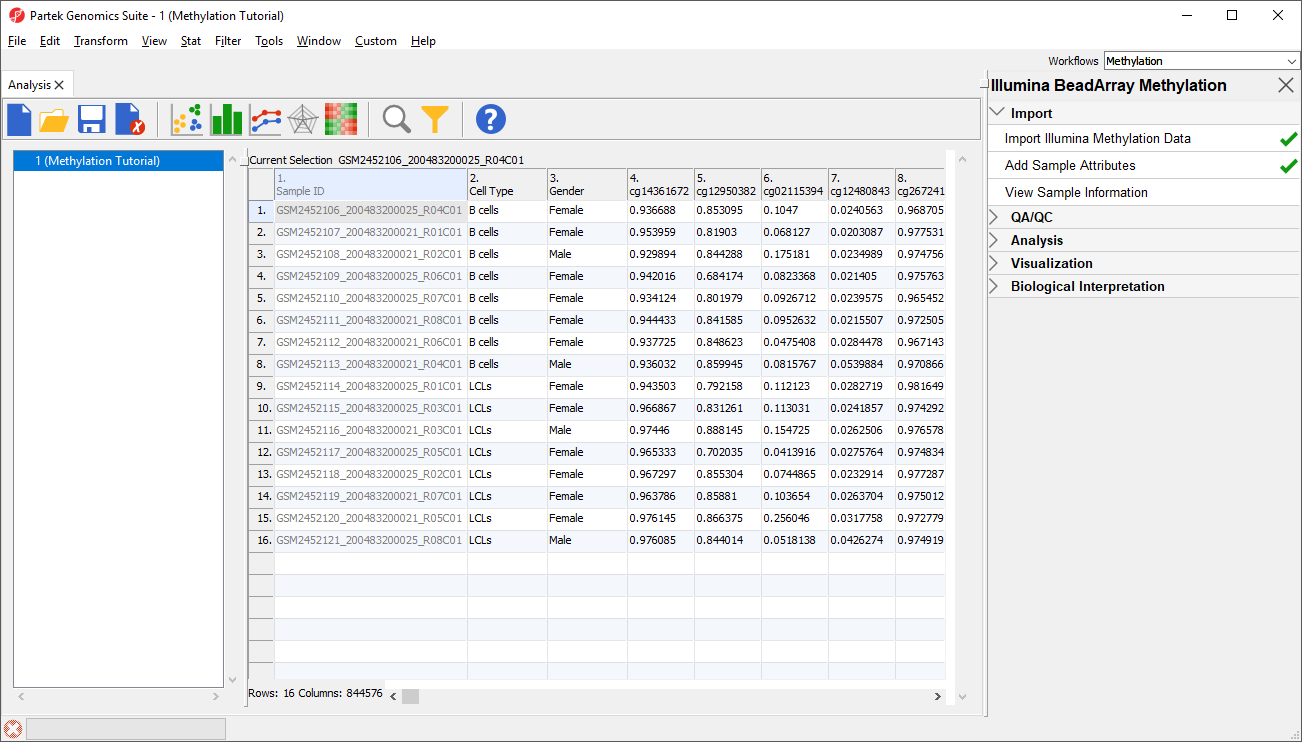
Before we can perform any analysis, the study samples need to be organized into their experimental groups.
...
Two new columns have been added to spreadsheet 1 (Methylation) with the cell type and gender of each sample .
For this tutorial, we want to exclude all male samples to simplify our analysis. To do this, we can use the interactive filter.
- Select () from the quick action bar to invoke the interactive filter
- Select 4. CHR for Column
For categorical columns, the interactive filter displays each category of the selected column as a colored bar. For 4. CHR, each bar represents a chromosome with the height of bar representing the number of probes from that chromosome in the selected spreadsheet. To filter out a category, left-click on its bar. Right clicking on a bar will include only the selected category. A pop up balloon will show the category label as you mouse over each bar.
...
(Figure 5).
The yellow and black bar on the right-hand side of the spreadsheet panel shows the fraction of excluded cells in black and included cells in yellow. Right-clicking this bar brings up an option to clear the filter.
Now that we have filtered out probes on the X and Y chromosomes, we will create a spreadsheet containing only probes on the autosomes.
- Right click on the ANOVA-1way (ANOVAResults) spreadsheet in the spreadsheet tree panel (Figure 4)
| Numbered figure captions | |
|---|---|
|
...
| |||
| Page Turner | ||
|---|---|---|
|
...