Page History
...
Analysis of variance (ANOVA) is a very powerful technique for identifying differentially expressed genes in a multi-factor experiment. In this data set, ANOVA will be used to generate a list of genes that are significantly differentially regulated by each treatment by two-fold.
Adding factors and interactions
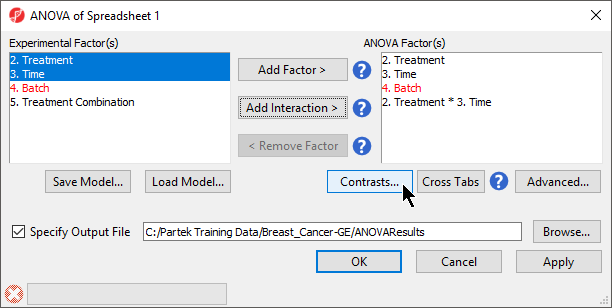
When setting up the ANOVA, the primary factors of interest, Treatment and Time, should be included. We will also include the interaction between Treatment and Time, Treatment * Time, because we are interested in whether different treatments behave differently over time. From our exploratory analysis using the PCA, we also know that Batch is a major source of variation and needs to be included. Including Batch as a random factor will allow us to account for the batch effect.
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Adding linear contrasts
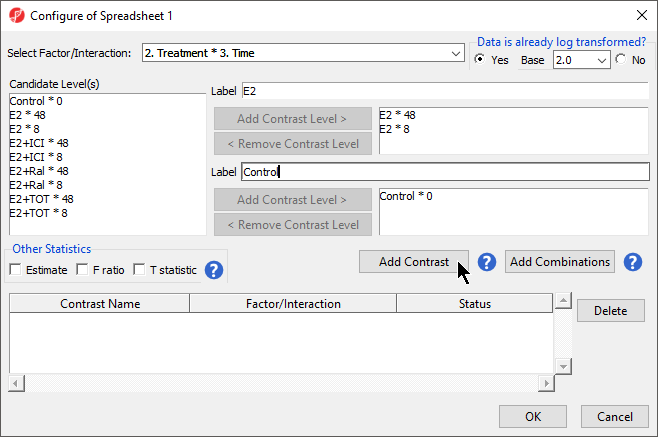
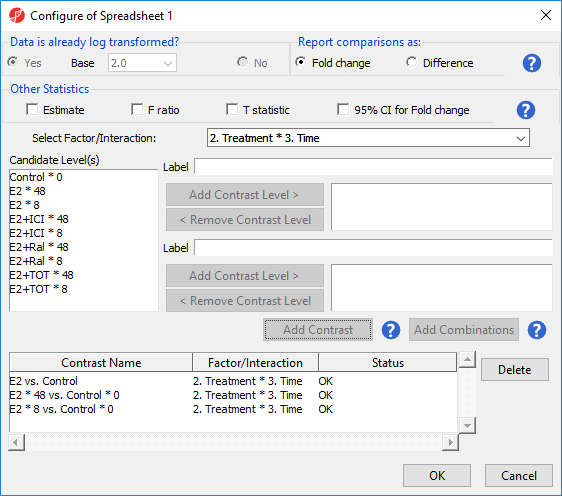
ANOVA will output a p-value and F ratio for each factor or interaction; to get the fold-change and ratio between the different levels of a factor or interaction, a linear contrasts, or comparisons, must be added.
...
The lower panel (Group 2) is considered the reference level. Because the data is log2 transformedlog2 transformed, the geometric mean will be used to calculate the fold change and mean ratio to place both on a linear scale instead of a log scale.
- Select Add Contrast (Figure 2)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
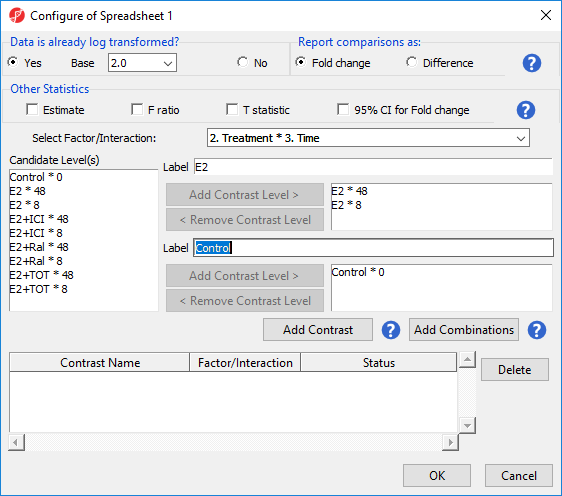
To examine the time points of each treatment condition separately, we would can select Add Combinations instead of Add Contrast. This adds every possible contrast for the levels in the Group 1 and Group 2 panels.
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
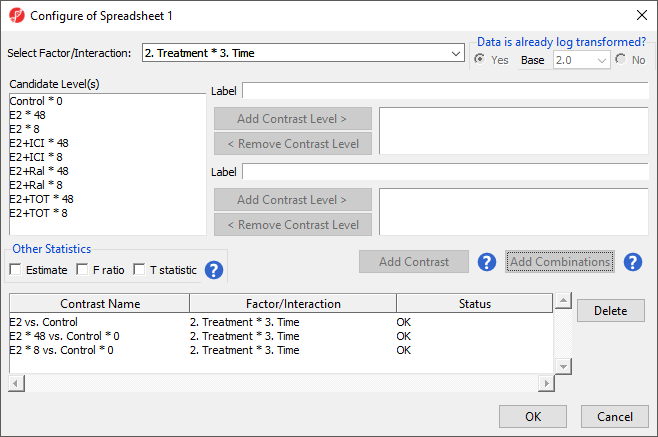
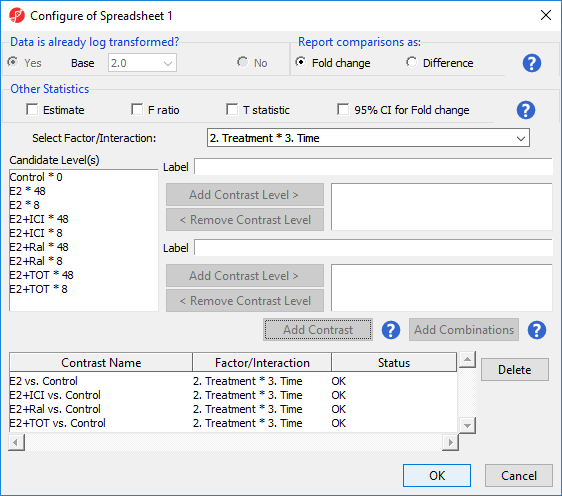
For this tutorial, we will not be considering the time points of each treatment condition individually. We can remove the E2 * 8 vs. Control * 0 and E2 * 48 vs. Control * 0 contrasts.
...
There should now be four contrasts added to the contrasts panel (Figure 4).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
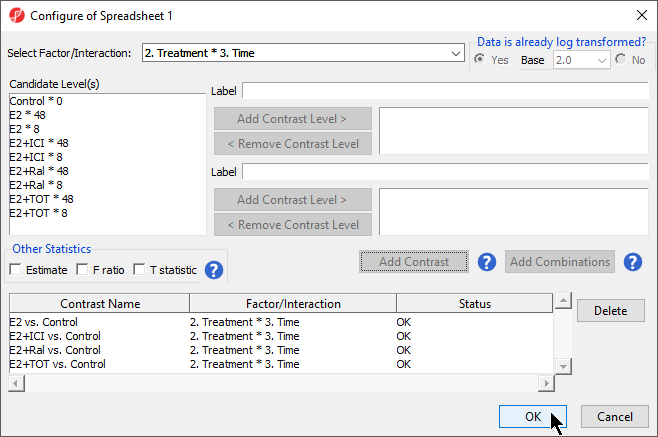
- Select OK to add the contrasts to the ANVOA model
...
- Select OK to perform the ANOVA
ANOVA results spreadsheet
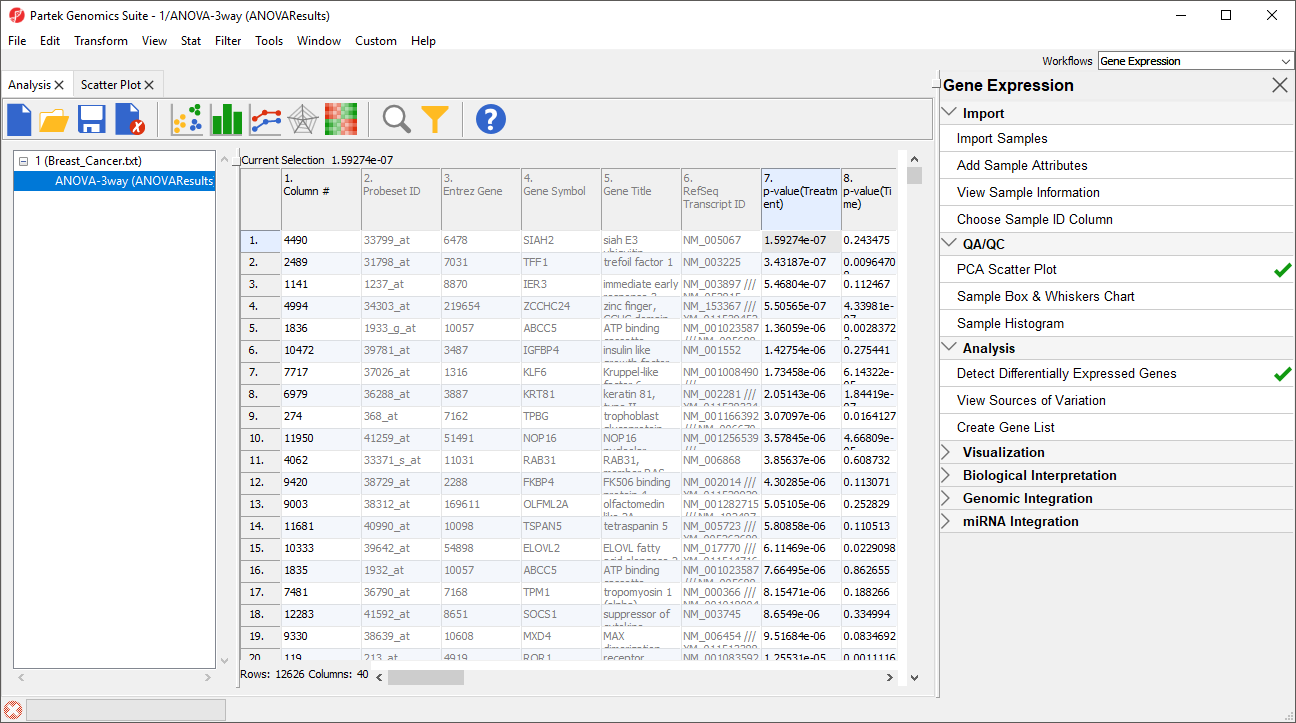
The result of the 3-way mixed model ANOVA is displayed in a new spreadsheet, ANOVA-3way (ANOVAResults) that is a child of the Breast_Cancer.txt spreadsheet. In ANOVAResults, each row represents a probe(set)/gene with the columns containing the results of the ANOVA (Figure 5).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
Each factor in the ANOVA adds p-value, F value, and SS value columns. F value is a ratio of signal to noise; high values indicate that the probe(set)/gene explains variation in the data set due to the factor. SS value is the sum of squares.
Each contrast in the ANOVA adds p-value, ratio, and fold-change columns. The p-value is calculated using log space. The ratio and fold change are calculated using linear space.
Viewing the sources of variation
Sources of variation captured in the ANOVA can be viewed for the entire data set or for individual probe(sets)/genes.
...
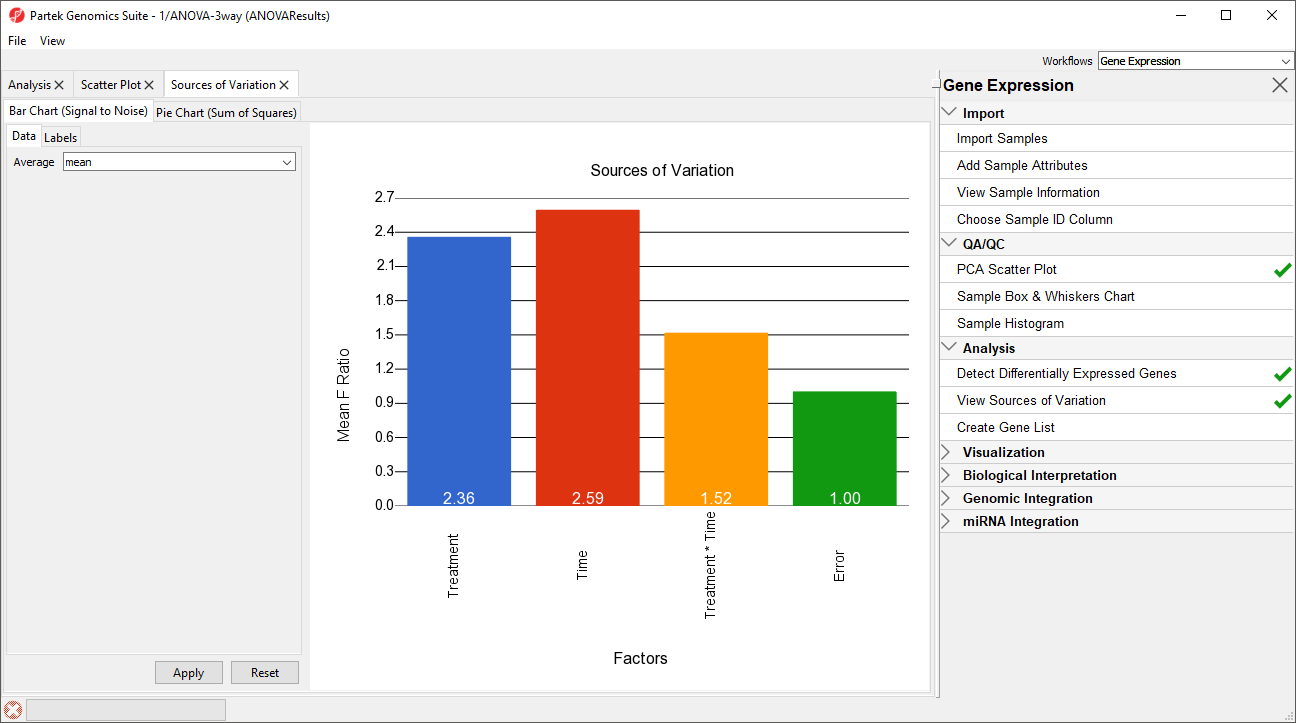
The Sources of Variation plot will open in a new tab (Figure 6).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
- Right-click on a row header in the ANOVAResults spreadsheet
- Select Sources of Variation from the pop-up menu
The plot will open in a new tab. For additional plots that can be invoked from the ANOVA results spreadsheet, see the Visualizations user guide.
| Page Turner | ||
|---|---|---|
|
| Additional assistance |
|---|
|
| Rate Macro | ||
|---|---|---|
|