| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| minLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
t-SNE (t-distributed stochastic neighbor embedding) is a visualization method commonly used to analyze single-cell RNA-Seq data. Each cell is shown as a point on the plot and each cell is positioned so that it is close to cells with similar overall gene expression. When working with multiple samples, a t-SNE plot can be drawn for each sample or all samples can be combined into a single plot. Viewing samples individually is the default in Partek Flow Partek® Flow® because sample to sample variation and outlier samples can obscure cell type differences if all samples are plotted together. However, as you will see in this tutorial, in some data sets, cell type differences can be visualized even when samples are combined.
Using the t-SNE plot, cells can be classified based on clustering results or and differences in gene and pathway expressionexpression of key marker genes.
Multiple single-sample t-SNE plots
By default, each sample in a multi-sample data set is plotted on its own t-SNE.
...
Prior to performing t-SNE, it is a good idea to reduce the dimensionality of the data using principal components analysis (PCA).
- Click the Filtered counts data node after the Filter features task
- Select PCA from the Exploratory analysis section of the task menu (Figure 1)
| Numbered figure captions |
|---|
| SubtitleText | Select the PCA task from the Exploratory analysis menu |
|---|
| AnchorName | Select PCA task |
|---|
|
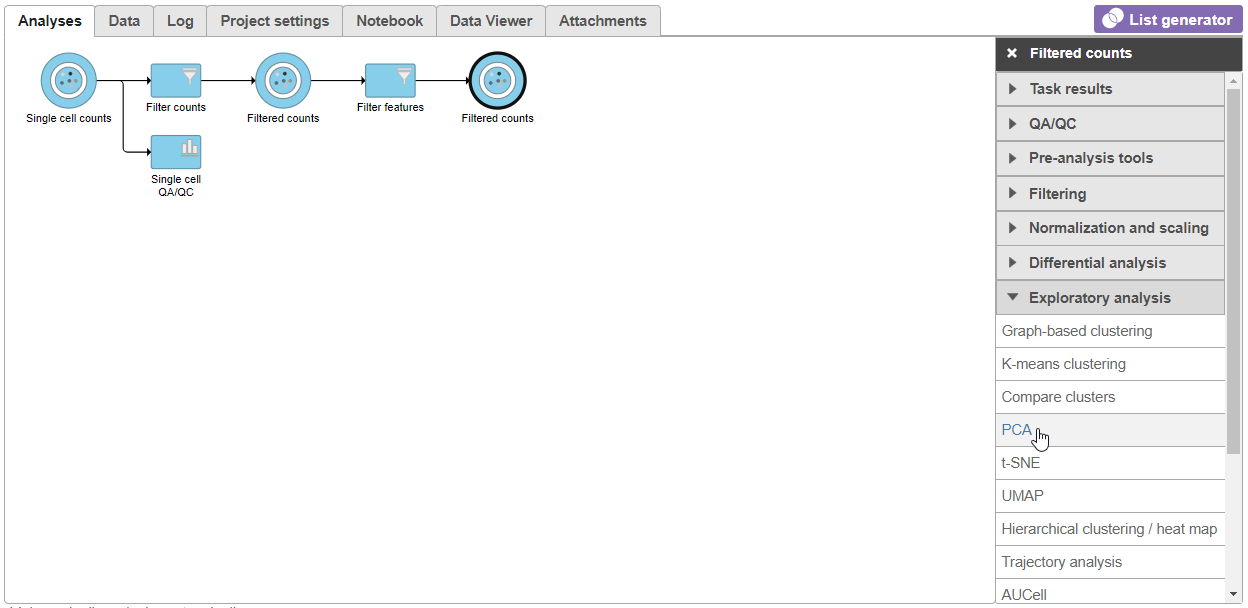 Image Added Image Added
|
- Click Finish to run PCA with default settings (Figure 2)
Note, the default settings include the Split by sample checkbox being selected. This means that the dimensionality reduction will be performed on each sample separately.
| Numbered figure captions |
|---|
| SubtitleText | PCA task set up page with default settings |
|---|
| AnchorName | PCA task set up |
|---|
|
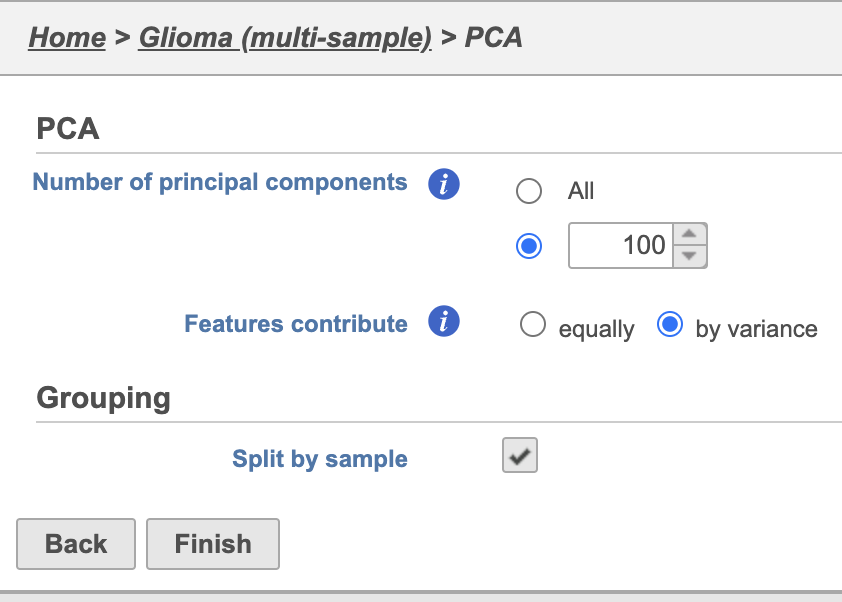 Image Added Image Added
|
PCA task and data nodes will be generated.
- Click the PCA data node
- Select t-SNE from the Exploratory analysis section of the task menu (Figure 1Figure 3)
| Numbered figure captions |
|---|
| SubtitleText | Invoking t-SNE from the task menu |
|---|
| AnchorName | Invoking t-SNE |
|---|
|
 Image Removed Image Removed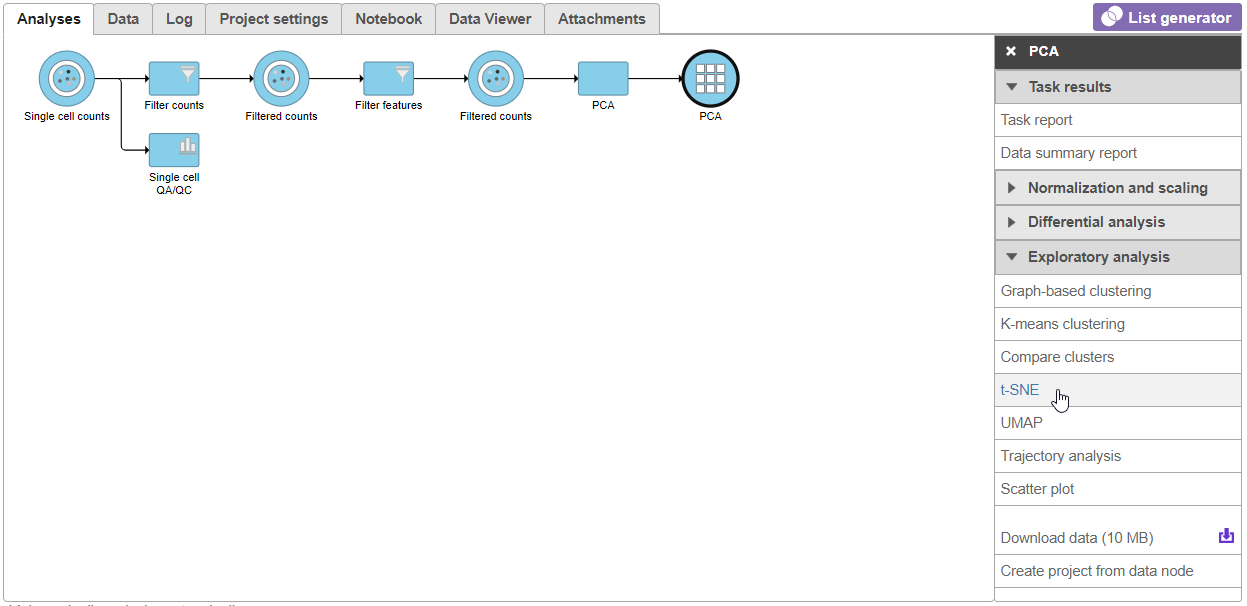 Image Added Image Added
|
- Click Finish from the t-SNE dialog to run t-SNE with the default settings
...
...
| set up with default settings | | AnchorName | t-SNE task set up |
|---|
|
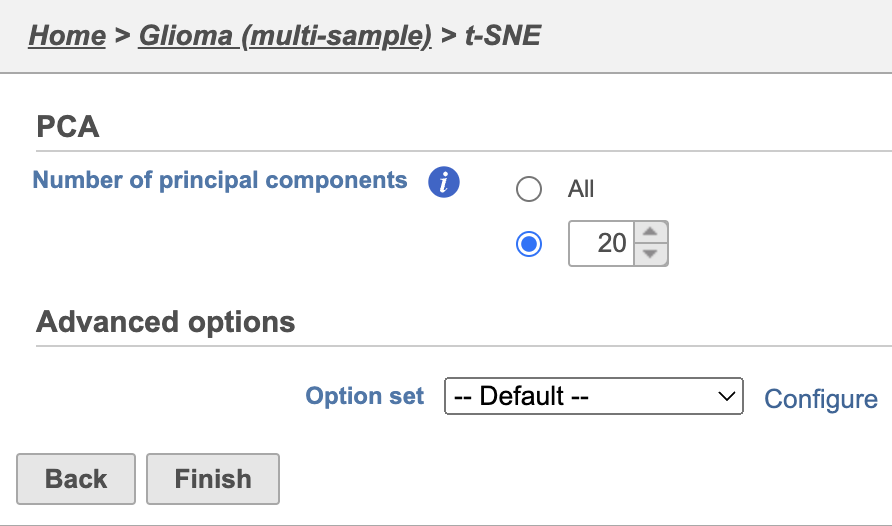 Image Added Image Added
|
Because the upstream PCA task was performed separately for each sample, the t-SNE task will also be performed separately for each sample. t-SNE task and data nodes will be generated (Figure 5).
| Numbered figure captions |
|---|
| SubtitleText | t-SNE task node |
|---|
| AnchorName | t-SNE task node |
|---|
|
 Image Removed Image Removed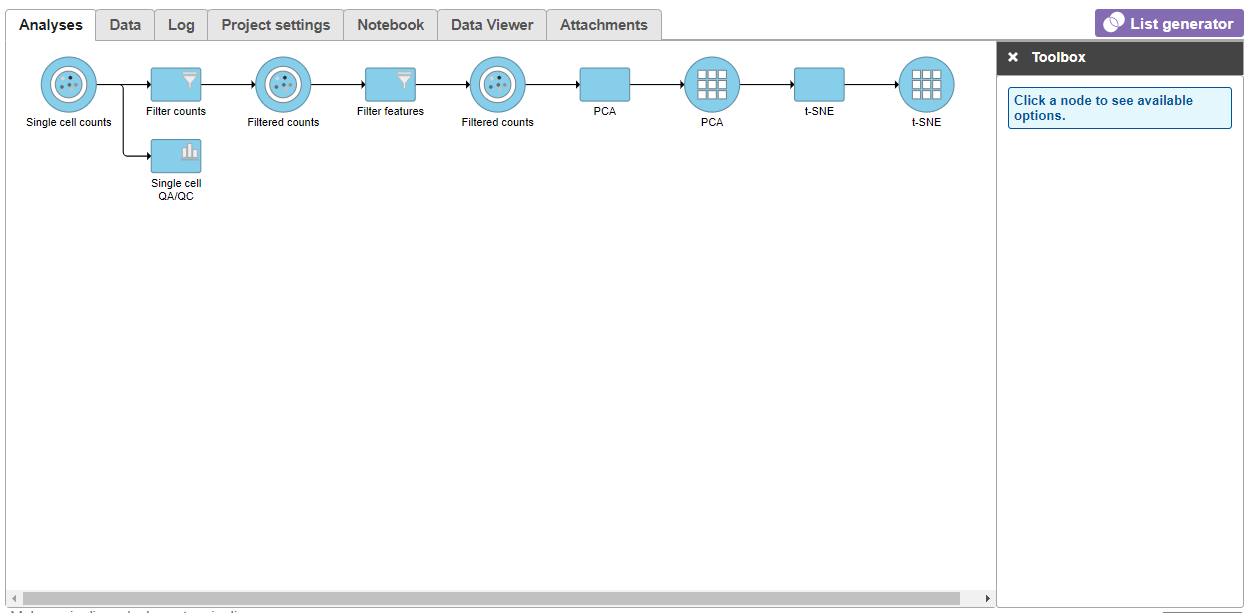 Image Added Image Added
|
Once the t-SNE task has completed, we can view the t-SNE plot.plots
- Click the t-SNE node
- Click Task report from the task menu or double click the t-SNE SNE node
The t-SNE plot will open to in a new data viewer session. The t-SNE plot for the first sample in the data set, MGH36 (Figure 3)Figure 6), will open on the canvas. Please note that the appearance of the t-SNE plot will may differ each time it is drawn so your t-SNE plots will may look different than those shown in this tutorial; however. However, the cell-to-cell relationships indicated will be the same.
| Numbered figure captions |
|---|
| SubtitleText | Viewing t-SNE plot of a single sample |
|---|
| AnchorName | Viewing single-sample t-SNE |
|---|
|
 Image Removed Image Removed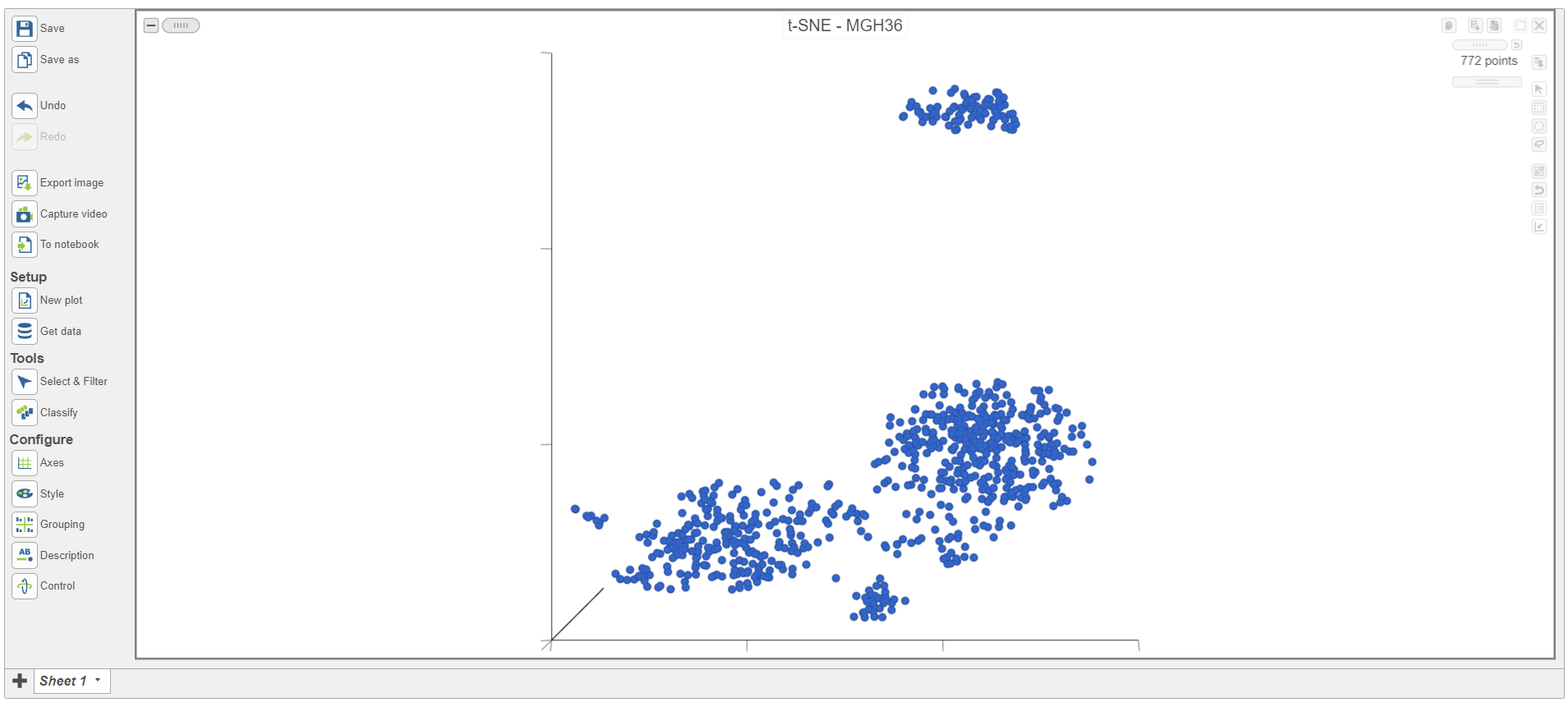 Image Added Image Added
|
The t-SNE plot is in 3D by default. To change the default, click your avatar in the top right > Settings > My Preferences and edit your graphics preferences and change the default scatter plot format from 3D to 2D.
You can rotate the 3D plot by lefleft-clicking and dragging your mouse. You can zoom in and out using your mouse wheel. The 2D t-SNE is also calculated and you can switch between the 2D and 3D plots using the Plot style radio buttons. on the canvas. We will do this later on in the tutorial.
Each sample has its own plot. We can switch between samples using the Back and Next buttons .
- Open the Axes icon on the
...
- left under Configure (Figure 7)
- Navigate to Misc
- Select the
 Image Addedicon below the Sample name to go to the next sample
Image Addedicon below the Sample name to go to the next sample
The t-SNE plot has switched to show the next sample, MGH42 (Figure 47).
| Numbered figure captions |
|---|
| SubtitleText | Viewing t-SNE plot of MGH42 |
|---|
| AnchorName | Viewing single-sample t-SNE (2) |
|---|
|
 Image Removed Image Removed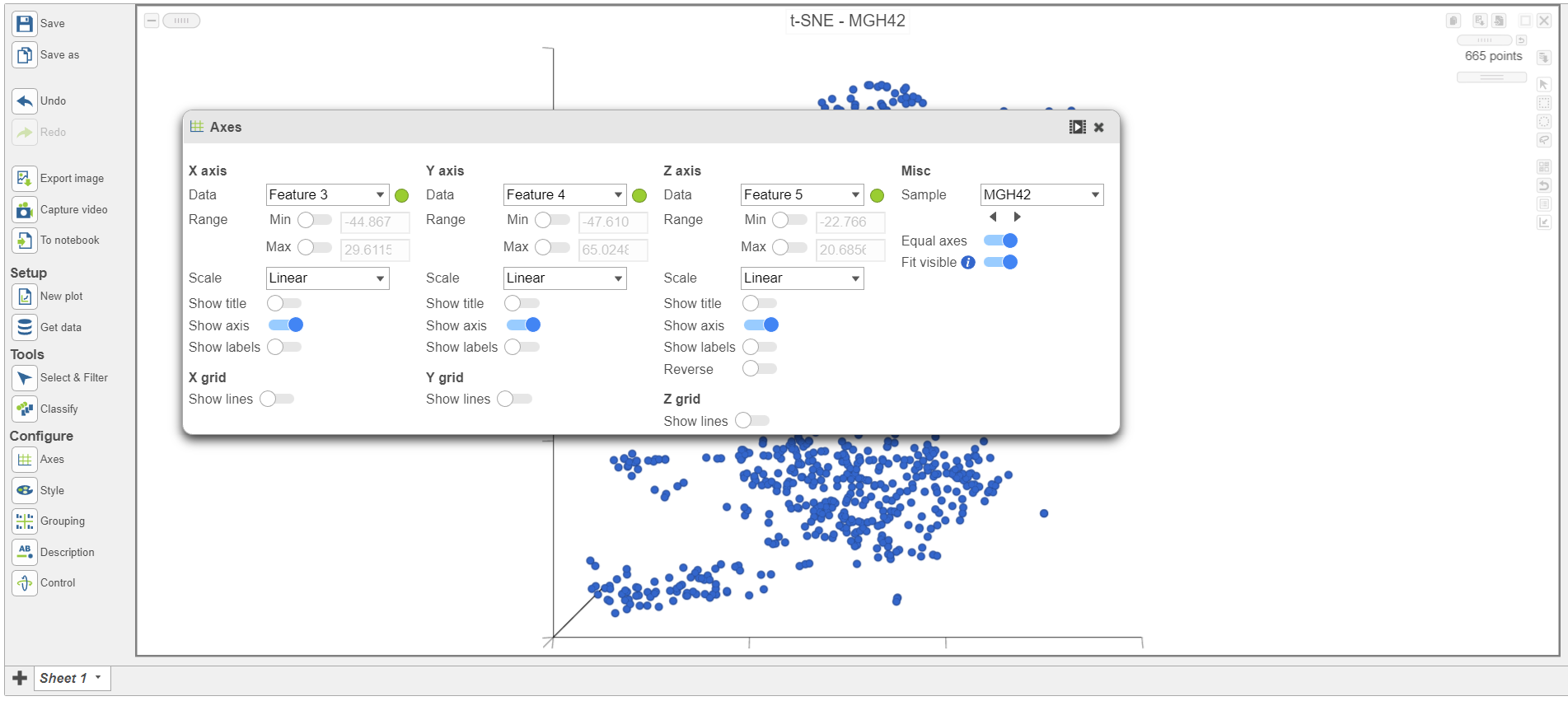 Image Added Image Added
|
The goal of this analysis is to compare malignant cells from two different glioma subtypes, astrocytoma and oligodendroglioma. To do this, we need to identify which cells are the malignant cells we want to include and which cell cells are the normal cells we want to exclude.
The t-SNE plot in Partek Flow offers several options for identifying, selecting, and classifying cells. In this tutorial, we will use the expression of known marker genes to identify normal cellscell types.
To visualize the expression of a marker gene, we can color cells on the t-SNE plot by their expression level.
...
| Numbered figure captions |
|---|
| SubtitleText | Selecting color by gene expression |
|---|
| AnchorName | Selecting color by gene expression |
|---|
|
 Image Removed Image Removed
|
The cells will turn black and a text box Gene ID will open below the drop-down box.
- Type BCAN in the Gene ID text box
- Select BCAN from the list of genes in the data set (Figure 6)Select any of the count data nodes from Get data on the left (Single cell counts, or any of the Filtered counts, Figure 8)
- Search for the BCAN gene
- Click and drag the BCAN gene onto the plot and drop it over the Green (feature) option
| Numbered figure captions |
|---|
| SubtitleText | Coloring cells by BCAN expression |
|---|
| AnchorName | Selecting gene |
|---|
|
 Image Removed Image Removed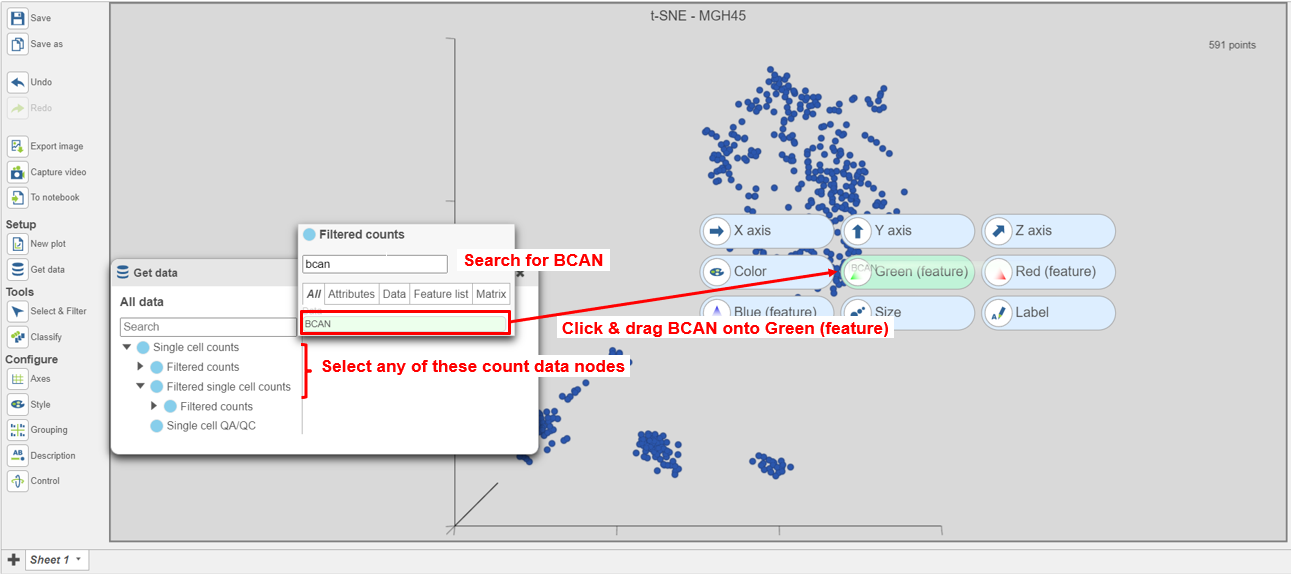 Image Added Image Added
|
The cells will be colored from black to green based on their expression level of BCAN, with cells expressing higher levels more green (Figure 7Figure 9). BCAN is highly expressed in glioma cells.
| Numbered figure captions |
|---|
| SubtitleText | Cells colored by CD14 BCAN expression |
|---|
| AnchorName | Colored by CD14 |
|---|
|
 Image Removed Image Removed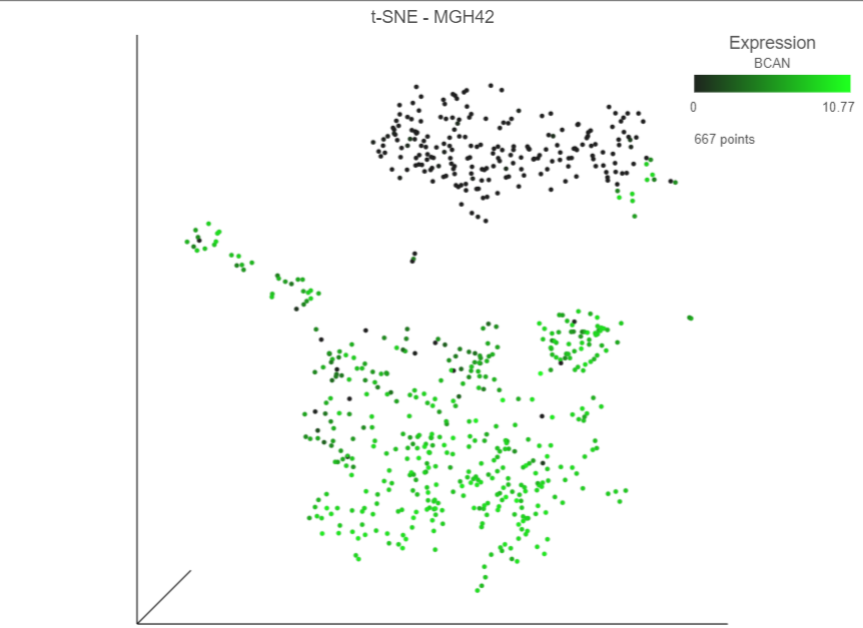 Image Added Image Added
|
In Partek Flow, we can color cells by more than one gene. We will now add a second glioma marker gene, GPM6A.
- Select the
 Image Removed icon next to BCAN
Image Removed icon next to BCAN - Type GPM6A in the new Gene ID box
- Select GPM6A from the list of genes in the data setany of the count data nodes from the Data card on the left (Single cell counts, or any of the Filtered counts)
- Search for the GPM6A gene
- Click and drag the GPM6A gene onto the plot and drop it over the Red (feature) option
Cells expressing GPM6A are now colored red and cells expressing BCAN are colored green. Cells expressing both genes are colored yellow, while cells expressing neither are colored black (Figure 8Figure 10).
| Numbered figure captions |
|---|
| SubtitleText | Coloring cells by BCAN and GPM6A |
|---|
| AnchorName | Coloring by two genes |
|---|
|
 Image Removed Image Removed
|
Relative expression of the two genes for selected cells can be visualized on the legend.
...
| Numbered figure captions |
|---|
| SubtitleText | Selecting a group of cells using the 3D lasso tool |
|---|
| AnchorName | Selecting a group of cells |
|---|
|
 Image Removed Image Removed
|
Selected cells are shown in bold and unselected cells are dimmed.
The relative expression of the two genes for the selected cells will be shown on the legend as dots (Figure 10).
| Numbered figure captions |
|---|
| SubtitleText | Viewing expression levels for a group of cells |
|---|
| AnchorName | Viewing expression levels for a group of cells |
|---|
|
 Image Removed Image Removed
|
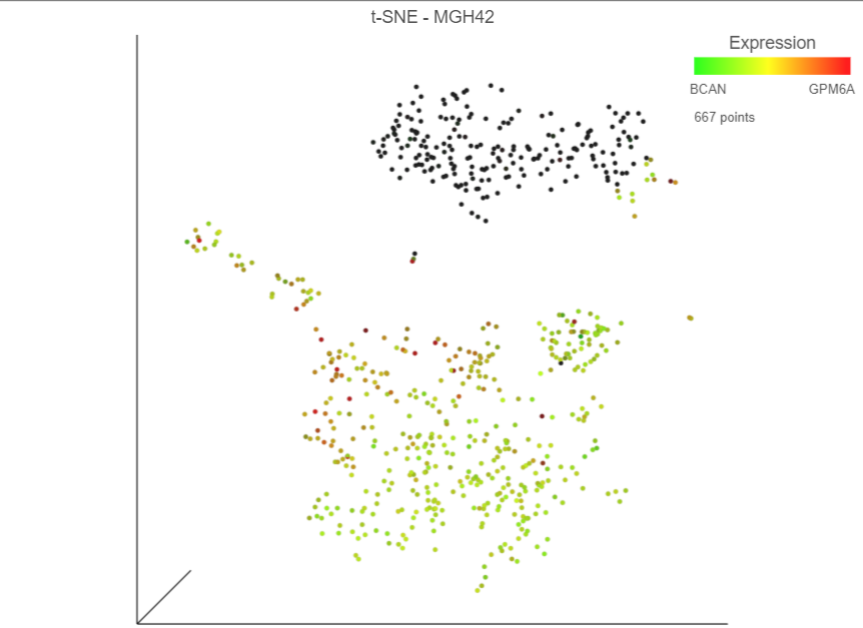 Image Added Image Added
|
Numerical expression levels for each gene can be viewed for individual cells.
- Switch modes to pointer mode by selecting
 Image Removedclicking
Image Removedclicking  Image Added in the top right corner of the plot
Image Added in the top right corner of the plot - Select a cell by pointing and clicking
The expression level for that cell is displayed on the legend for each gene (Figure 11. Expression values can also be viewed by mousing over a cell (Figure 11).
- Deselect the cell by clicking on any black blank space on the plot
Expression values can also be viewing by selecting Gene Expression from the Label by drop-down menu and mousing over a cell.
| Numbered figure captions |
|---|
| SubtitleText | Viewing expression levels for an individual cell. The dots on the legend indicate the expression level of the selected cell. The expression levels also appear in the label when you mouse over a cell |
|---|
| AnchorName | Viewing expression levels for a cell |
|---|
|
 Image Removed Image Removed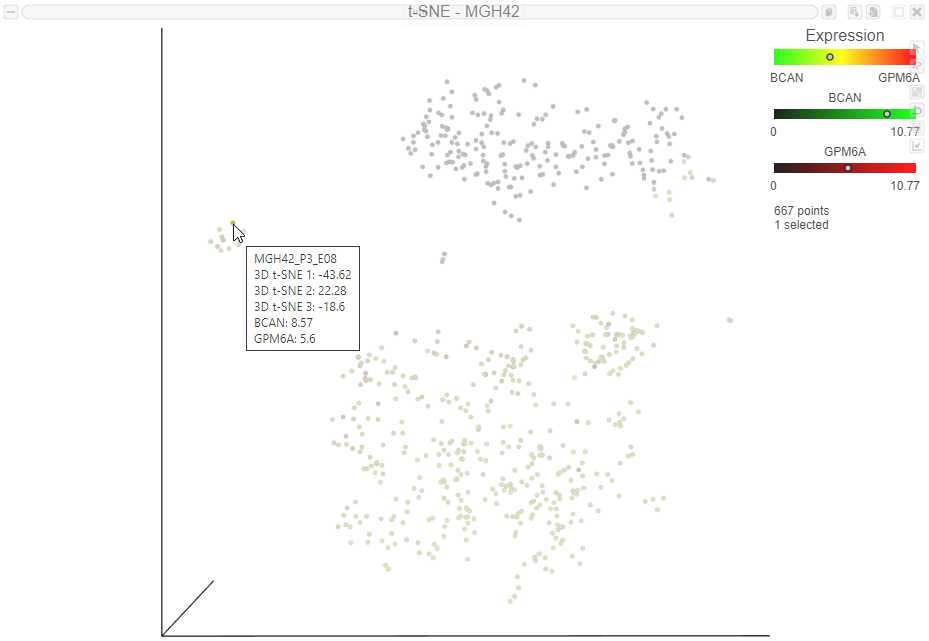 Image Added Image Added
|
Now that cells are colored by the expression of two glioma cell markers, we can classify any cell that expresses these genes as glioma cells. Because t-SNE groups cells that are similar across the high-dimensional gene expression data, we will consider cells that form a group with BCAN where the majority of cells express BCAN and/or GPM6A -expressing cells as the same cell type, even if they do not express the either marker gene.
- Click anywhere on the t-SNE plot without a cell to clear the selection
- Activate the 3D lasso tool by selecting
 Image RemovedSwitch to lasso mode by clicking
Image RemovedSwitch to lasso mode by clicking  Image Added in the top right of the plot
Image Added in the top right of the plot - Draw the lasso around the cluster of green, red, and yellow cells and click the circle to close the lasso . You may need to switch to selection mode and rotate the 3D plot to select only cells from the yellow cluster(Figure 12)
| Numbered figure captions |
|---|
| SubtitleText | Selecting a group of cells using the 3D lasso tool |
|---|
| AnchorName | Selecting a group of cells |
|---|
|
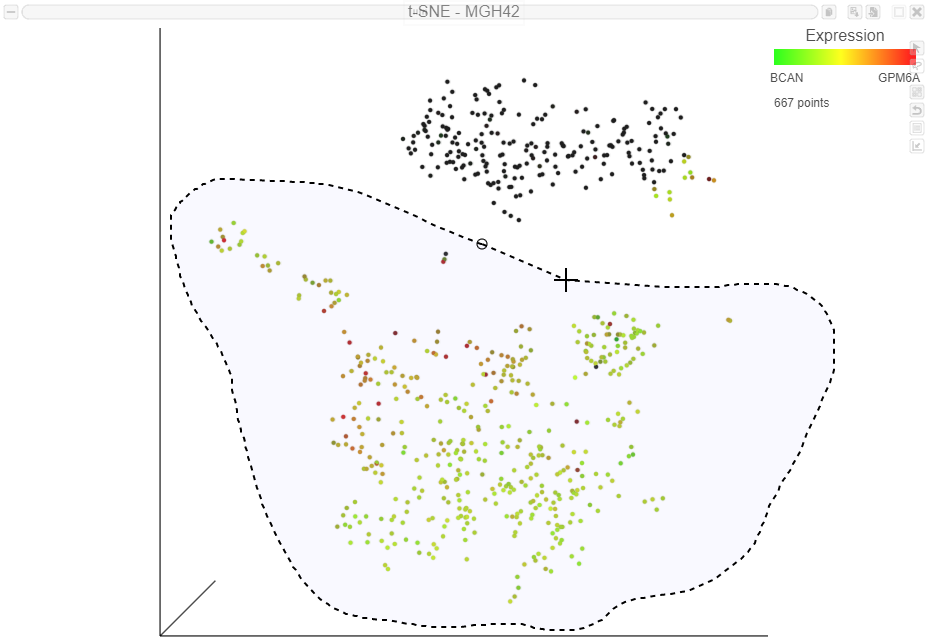 Image Added Image Added
|
Selected cells are shown in bold and unselected cells are dimmed. The number of selected cells is indicated in the Selection section of the menu.
...
figure legend. The cells are plotted on the color scale depending on their relative expression levels of the two marker genes (Figure 13)
...
| Viewing expression levels for a group of cells | | AnchorName |
|---|
|
...
| Viewing expression levels for a group of cells |
|
...
 Image Removed
Image Removed
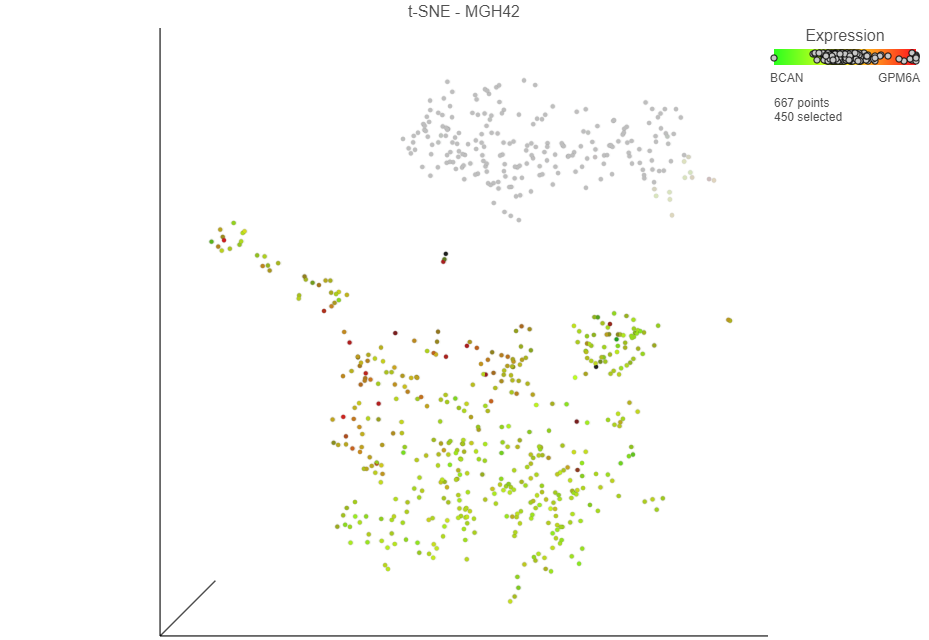 Image Added Image Added
|
- Click Classify selection in the Classify icon under Tools
A dialog to give the classification a name will appear.
- Name the classification Glioma
- Select Click Save (Figure 13Figure 14)
| Numbered figure captions |
|---|
| SubtitleText | Classifying selection |
|---|
| AnchorName | Classifying cells |
|---|
|
 Image Removed Image Removed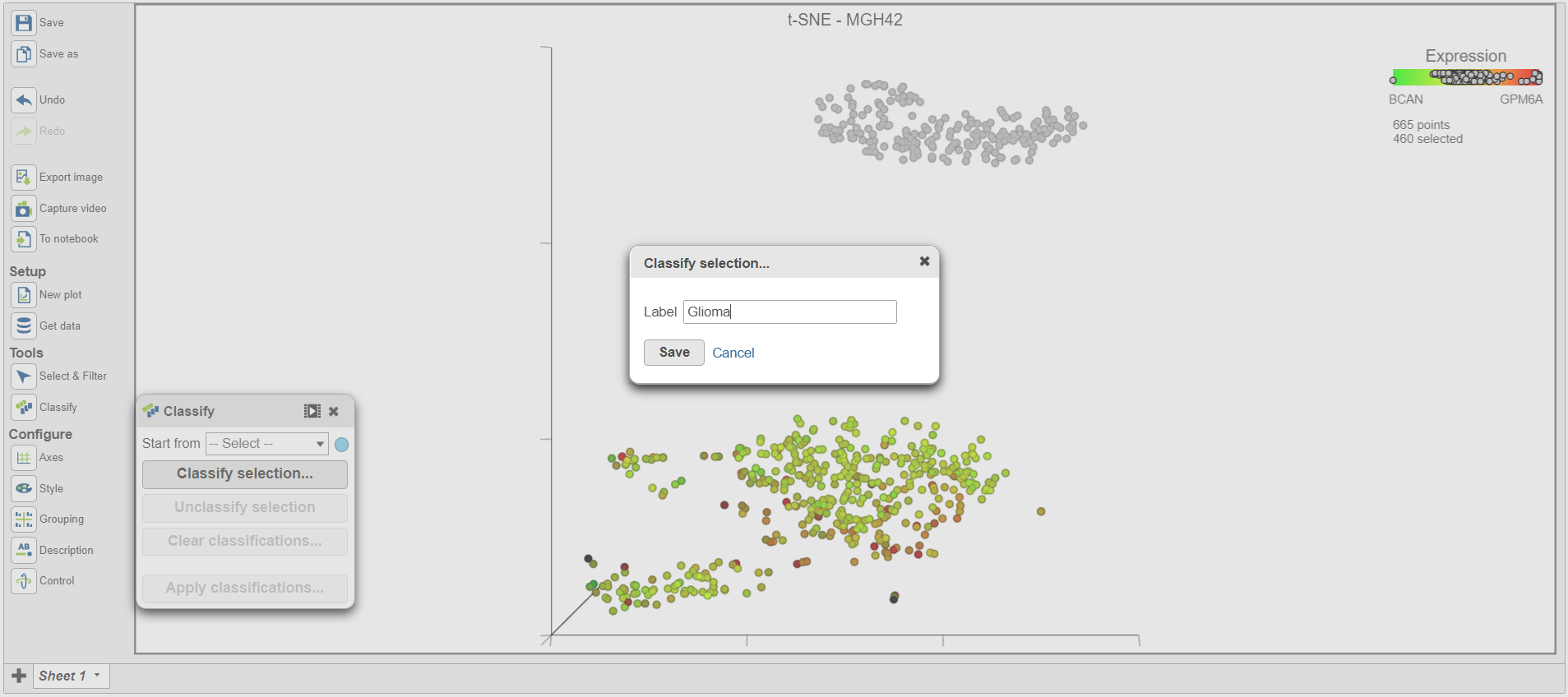 Image Added Image Added
|
Once cells have been classified, the classification is added to the Classifications section of the panel Classify. The number of cells belonging to the classiciation classification is listed; in . In MGH42, there are 413 460 glioma cells (Figure 14Figure 15).
| Numbered figure captions |
|---|
| SubtitleText | The number of cells in each classification is displayed |
|---|
|
...
| | AnchorName | Viewing classifications |
|---|
|
...
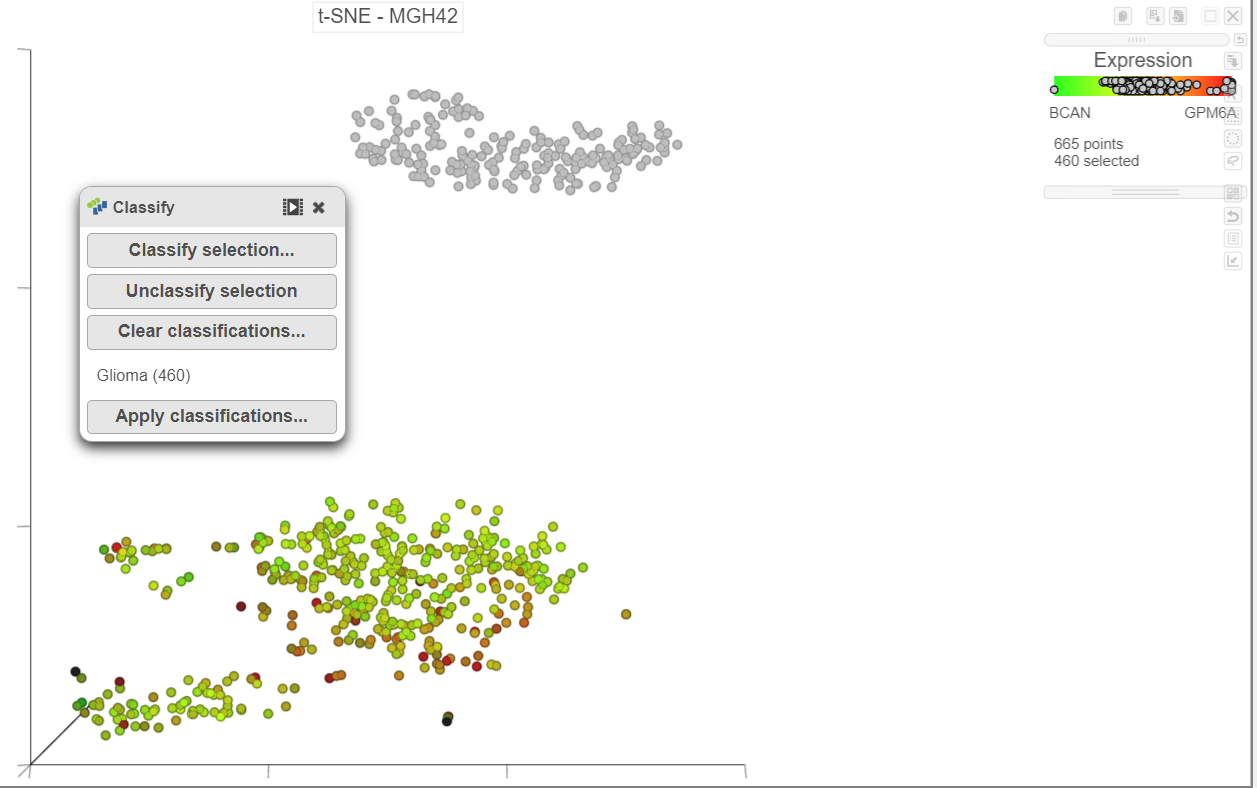 Image Added Image Added
|
Classifications made on the t-SNE plot are retained as a draft after you exit the t-SNE task report. The Save classifications button runs a task, Classify cells, which generates a new Classified cells data nodeas part of the data viewer session. In this tutorial, we will classify malignant cells for each sample before we save and apply the classifications, but if necissarynecessary, you can exit the t-SNE task report and continue classifying the next sample later.
...
save the data viewer session by clicking the  Image Added Save icon on the left to retain all of the formatting and draft classifications. The data viewer session will be stored under the Data viewer tab and can be re-opened to continue making classifications at a later time.
Image Added Save icon on the left to retain all of the formatting and draft classifications. The data viewer session will be stored under the Data viewer tab and can be re-opened to continue making classifications at a later time.
- Switch to pointer mode by clicking
 Image Added in the top right corner of the plot
Image Added in the top right corner of the plot - Deselect the cells by clicking on any blank space on the plot
- Open Axes and navigate to Sample under Misc
- Select the
 Image Addedicon below the sample name to go to the next sample, MGH45
Image Addedicon below the sample name to go to the next sample, MGH45 - Rotate the 3D t-SNE plot to allow you to select only get a better view of cells from the green, red, and yellow cluster
- Activate the 3D lasso tool Switch to lasso mode by selecting
 in the top right corner of the plot
in the top right corner of the plot - Draw the lasso around the cluster of black colored cells and click the circle to close the lasso (Figure 14Figure 16).
| Numbered figure captions |
|---|
| SubtitleText | Classifying malignant cells in sample MGH45 |
|---|
| AnchorName | Classifying cells |
|---|
|
 Image Removed Image Removed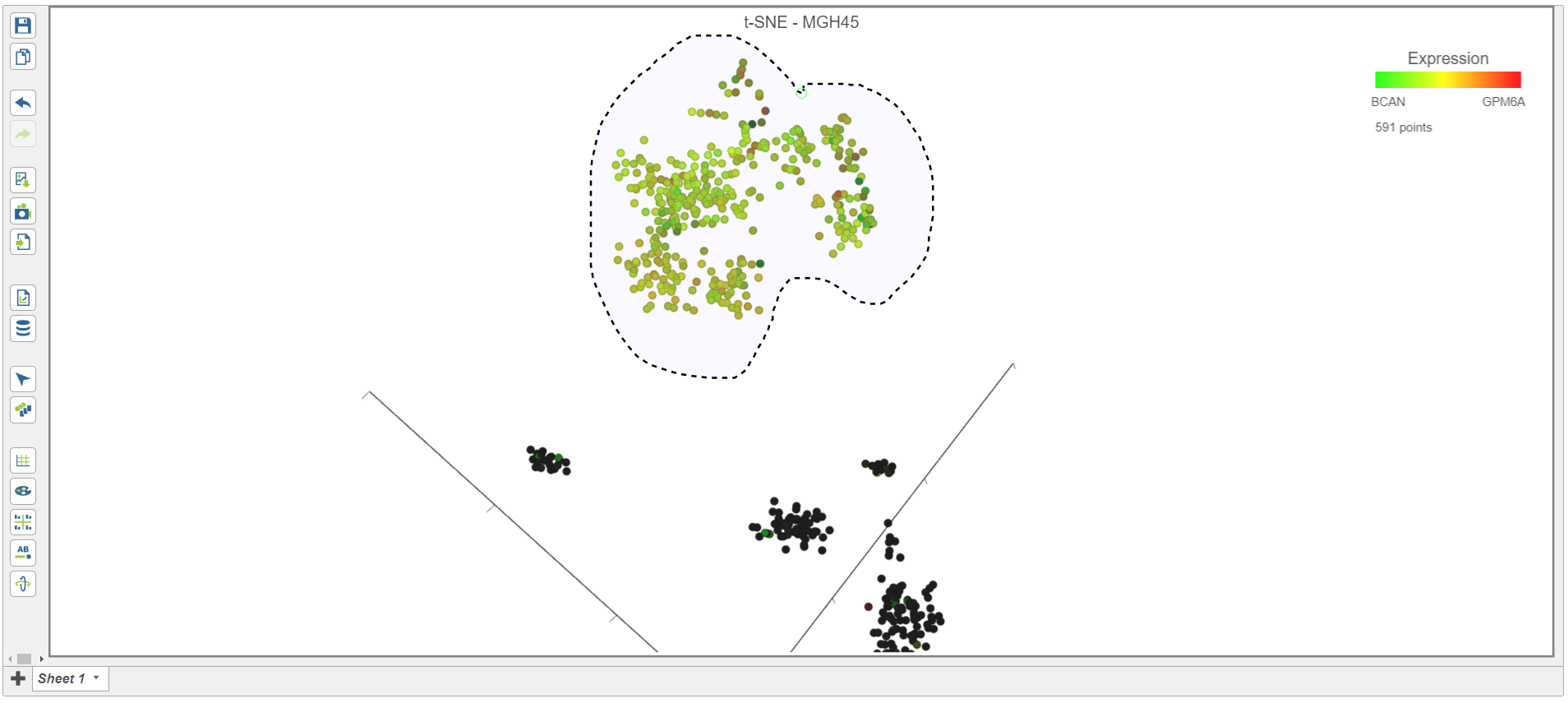 Image Added Image Added
|
- Select Classify selection selection in the Classify icon
- Type Glioma or select Glioma from the prompt (Figure 15drop-down list (Figure 17)
- Select Click Save
| Numbered figure captions |
|---|
| SubtitleText | Adding cells in a second sample to an existing classification |
|---|
| AnchorName | Classifying cells as an existing classification |
|---|
|
 Image Removed Image Removed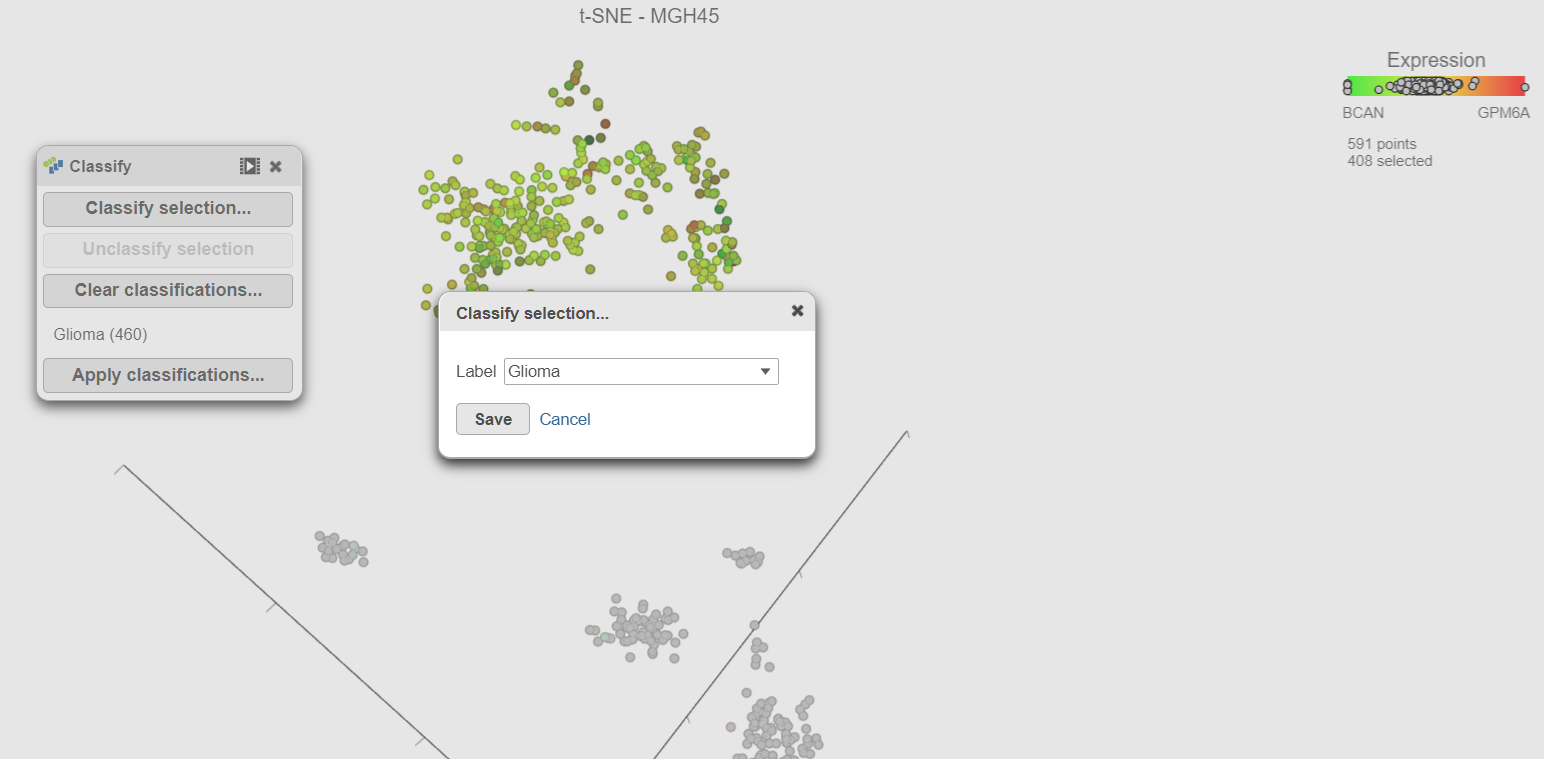 Image Added Image Added
|
- Repeat these steps for each of the 6 remaining samples
Once all samples have been classified, it is useful to check the number of cells in each sample assigned to each classification.
...
| Numbered figure captions |
|---|
| SubtitleText | Navigating to the classification summary |
|---|
| AnchorName | Navigating to classification summary |
|---|
|
 Image Removed Image Removed
|
The classifications summary lists every sample, the number of cells in the sample, the number of cells in each classification, and the percentage of cells in each sample that belong to each classification (Figure 17).
- . Remember to go back to the first sample (MGH36) to classify the glioma cells in that samples too.
There should be 5,322 glioma cells in total across all 8 samples.
- The classification name can be edited or deleted (Figure 18).
| Numbered figure captions |
|---|
| SubtitleText | Viewing Edit or delete the classification summary |
|---|
| AnchorName | Classification summary |
|---|
|  Image Removed
Image Removed| Edit or delete the classification |
|
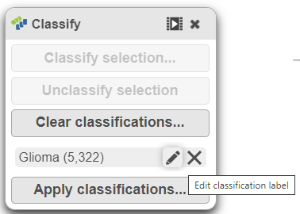 Image Added Image Added
|
With the malignant cells in every sample classified, it is time to save the classifications.
...
- Click Apply classifications
- Select Apply when asked to confirm
The pipeline view will open and the Classify cells tasks will run, generating a Classified groups data node (Figure 18).
- classifications in the Classify icon
- Name the classification attribute Cell type (sample level)
- Click Run (Figure 19)
| Numbered figure captions |
|---|
| SubtitleText | The Classify cells tasks generates a Classified groups data node |
|---|
| AnchorName | Classify cells task |
|---|
|
 Image Removed Image Removed
|
| Name the cell-level attribute | | AnchorName | Classifiy attribute name |
|---|
|
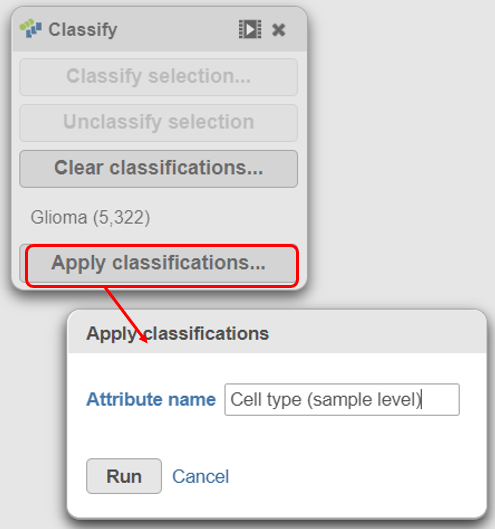 Image Added Image Added |
The new attribute is stored in the Data tab and is available to any node in the project.
- Click on the Glioma (multi-sample) project name at the top to go back to the Analyses tab
- Your browser may warn you that any unsaved changes to the data viewer session will be lost. Ignore this message and proceed to the Analyses tab
One multi-sample t-SNE plot
For some data sets, cell types can be distinguished when all samples can be visualized together on one t-SNE plot. We will use a t-SNE plot of all samples to classify glioma, microglia, and oligodendrocyte cell types.
- Select the Click on the Glioma (multi-sample) project name at the top to go back to the Analyses tab
- Click the Filtered counts data node Select t-SNE from the Exploratory anlaysis after the Filter features task
- Click PCA in the Exploratory analysis section of the task menu
- Select Configure on the t-SNE dialog Uncheck the Split by sample checkbox (Figure 1922)
- Click Finish
...
| Numbered figure captions |
|---|
| SubtitleText | Setting t-SNE to plot all samples together |
|---|
| AnchorName | Setting t-SNE to plot all samples together |
|---|
|
 Image Removed Image Removed
|
...
| Numbered figure captions |
|---|
| SubtitleText | Accessing the t-SNE advanced options |
|---|
| AnchorName | Accessing advanced options |
|---|
|
 Image Removed Image Removed
|
...
| Combine all cells into one plot by unchecking the Split by sample box | | AnchorName | Multi-sample PCA setting |
|---|
|
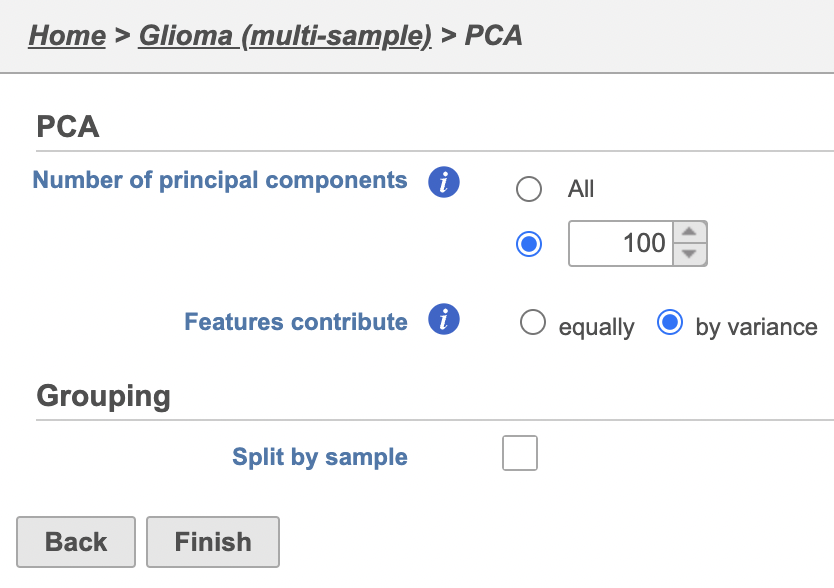 Image Added Image Added
|
The PCA task will run as a new green layer.
- Click the new PCA data node
- Select t-SNE from the Exploratory analysis section of the task menu
- Click Finish to run the t-SNE task with default settings
The t-SNE task will be added as a to the green layer in the analysis tab (Figure 21(Figure 23). Layers are created in Partek Flow when the same task is run on the same data node.
| Numbered figure captions |
|---|
| SubtitleText | All samples Multi-sample PCA and t-SNE task is tasks are added as a new layer |
|---|
| AnchorName | PCA and t-SNE task new layer |
|---|
|
 Image Removed Image Removed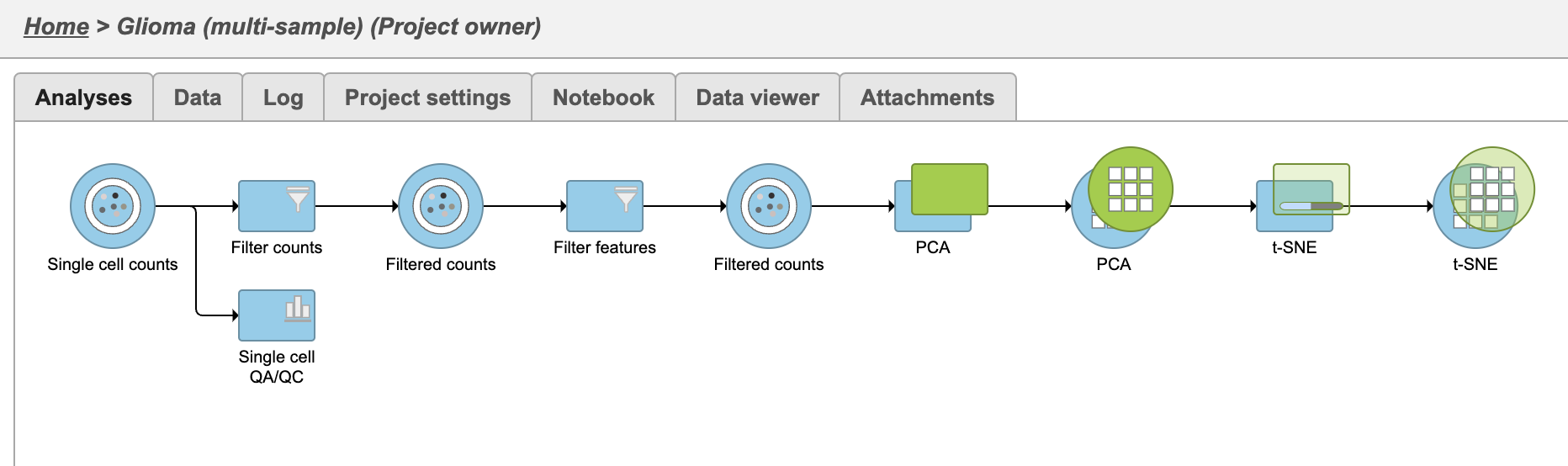 Image Added Image Added
|
Once the task has completed, we can view the plot.
- Select Double-click the green t-SNE plot node
- Select Task Report from the task menu
...
- data node to open the t-SNE scatter plot
- Click and drag the 2D scatter plot icon onto the canvas and replace the 3D scatter plot (Figure 24)
| Numbered figure captions |
|---|
| SubtitleText | Viewing the multi-sample t-SNE plot |
|---|
|
...
| | AnchorName | multi-sample t-SNE plot |
|---|
|
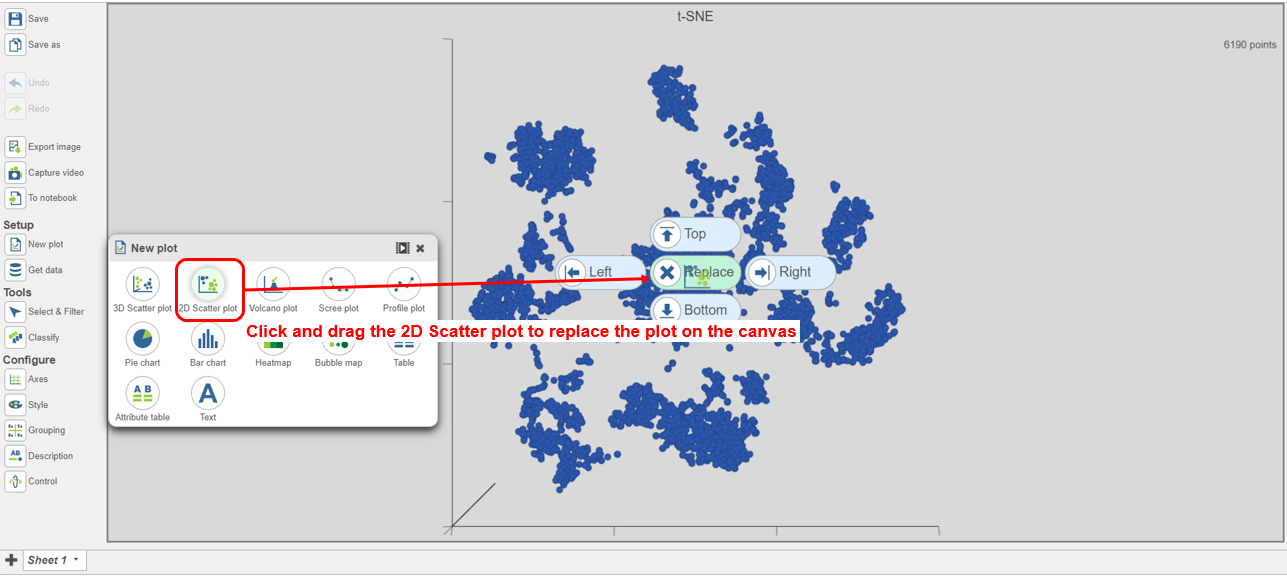 Image Added Image Added
|
- Search for and select green t-SNE data node (Figure 25)
| Numbered figure captions |
|---|
| SubtitleText | Viewing Select the green multi-sample t-SNE data node to draw the 2D t-SNE plot |
|---|
| AnchorName | Select multi-sample t-SNE plotdata |
|---|
|
 Image Removed Image Removed
|
...
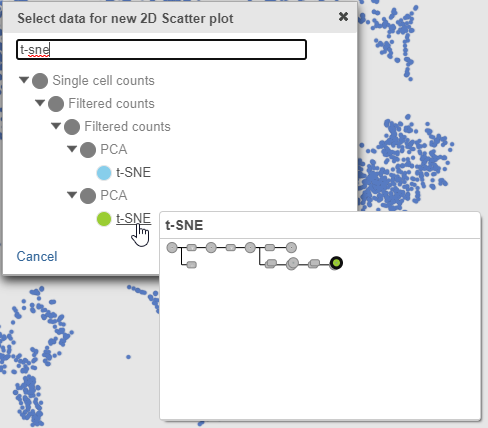 Image Added Image Added
|
- In the Style icon, choose Sample name from the Color by drop-down list under Color
Viewing the 2D t-SNE plot, while most cells cluster by sample, there are a few clusters with cells from multiple samples (Figure 23Figure 26).
| Numbered figure captions |
|---|
| SubtitleText | Viewing the multi-sample t-SNE plot in 2D |
|---|
| AnchorName | Viewing 2D t-SNE |
|---|
|
 Image Removed Image Removed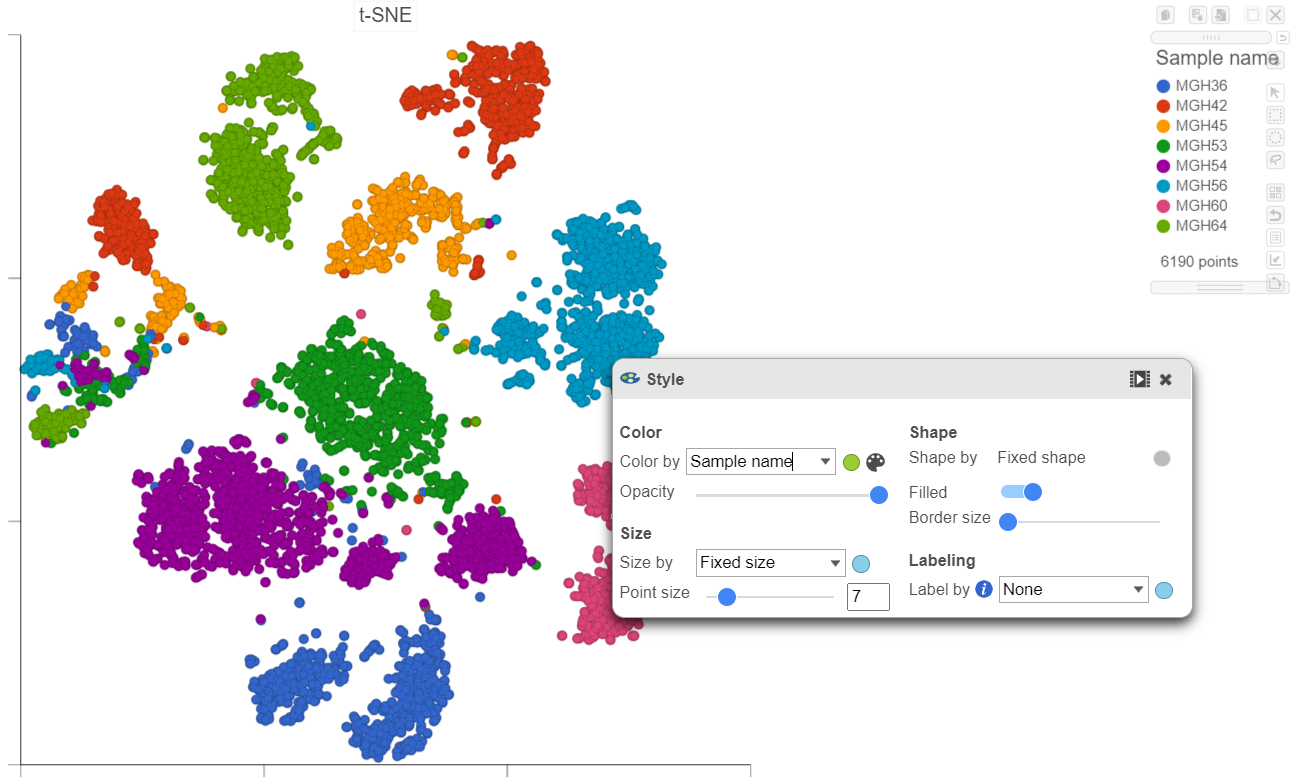 Image Added Image Added
|
Using maker marker genes, BCAN (glioma), CD14 (microglia), and MAG (oligodendrocytes), we can assess whether these multi-sample clusters belong to our known cell types.
- Select Gene expression from the Color by drop-down menu
- Type BCAN in the new Gene ID box
- Select BCAN from the list of genes in the data set
- Select the
 Image Removed icon next to BCAN
Image Removed icon next to BCAN - Type CD14 in the new Gene ID box
- Select CD14 from the list of genes in the data set
- Select the
 Image Removed icon next to CD14
Image Removed icon next to CD14 - Type MAG in the new Gene ID box
- Select MAG from the list of genes in the data setSelect any of the count data nodes from the Data card on the left (Single cell counts, or any of the Filtered counts)
- Search for the BCAN gene
- Click and drag the BCAN gene onto the plot and drop it over the Green (feature) option
- Search for the CD14 gene
- Click and drag the CD14 gene onto the plot and drop it over the Red (feature) option
- Search for the MAG gene
- Click and drag the MAG gene onto the plot and drop it over the Blue (feature) option
After coloring by these marker genes, three cell populations are clearly visible (Figure 24Figure 27).
| Numbered figure captions |
|---|
| SubtitleText | Overlaying marker gene expression on the multi-sample t-SNE plot |
|---|
| AnchorName | Overlaying gene expression |
|---|
|
 Image Removed Image Removed
|
...
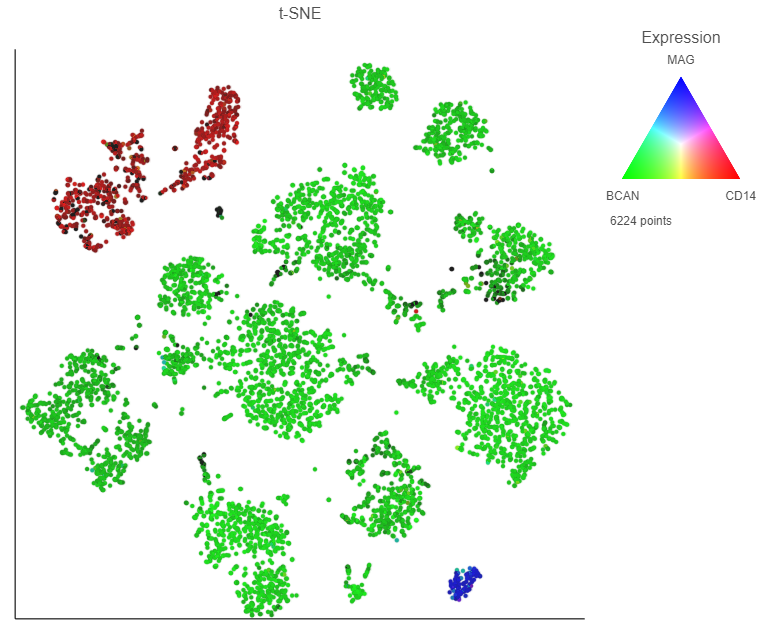 Image Added Image Added
|
The red cells are CD14 positive, indicating that they are the microglia from every sample.
- Switch to lasso mode by clicking the
 Image Added icon in the top right of the plot
Image Added icon in the top right of the plot - Draw the lasso around the cluster of red cells and click the circle to close the lasso (Figure 25Figure 28)
- Open the Classify tool and click Classify selection
- Name the classification Microglia
- Click Save
| Numbered figure captions |
|---|
| SubtitleText | Classifying microglia (red) |
|---|
| AnchorName | Classifying MOBP + cells |
|---|
|
 Image Removed Image Removed
|
...
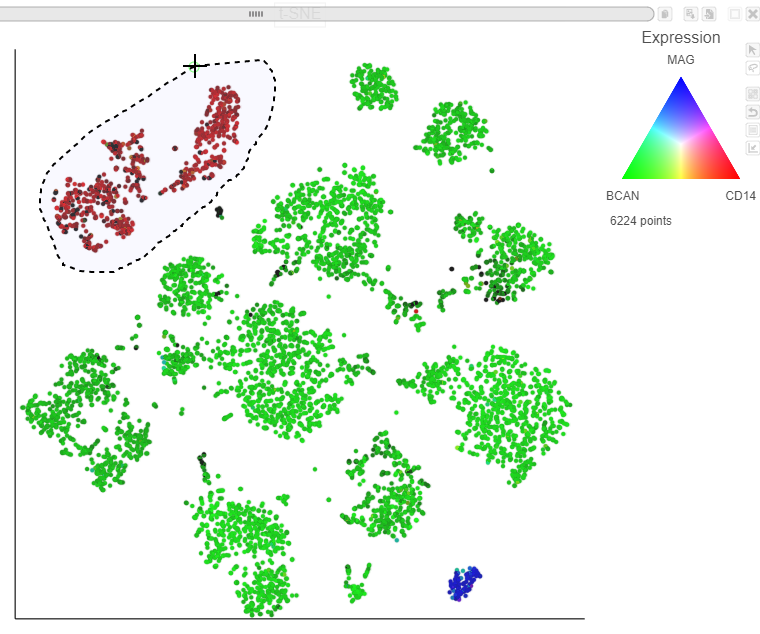 Image Added Image Added
|
The blue cells are CD14 MAG positive, indicating that they are the microglia oligodendrocytes from every sample.
- Name the classification Microglia
- Click Save
To clearly see the MAG expressing population, clear the current selection.
- Deselect by double clicking on any black Switch to pointer mode by clicking
 Image Added in the top right corner of the plot
Image Added in the top right corner of the plot - Deselect the cells by clicking on any blank space on the plot
...
- Switch to lasso mode again by clicking the
 Image Added icon in the top right of the plot
Image Added icon in the top right of the plot - Draw the lasso around the cluster of blue cells and click the circle to close the lasso (Figure 26)
| Numbered figure captions |
|---|
| SubtitleText | Classifying oligodendrocytes (blue) |
|---|
| AnchorName | Classifying microglia |
|---|
|
 Image Removed Image Removed
|
- Click Classify
- Open the Classify tool and click Classify selection
- Name the classification Microgliaclassification Oligodendrocytes
- Click Save
- Deselect by double clicking on any black space on the plot
Finally, we will classify the BCAN expressing cells on the plot as glioma cells from every sample.
- Switch to pointer mode by clicking
 Image Added in the top right corner of the plot
Image Added in the top right corner of the plot - Deselect the cells by clicking on any blank space on the plot
- Switch to lasso mode again by clicking the
 Image Added icon in the top right of the plot
Image Added icon in the top right of the plot - Draw the lasso around the cluster of green cells and click the circle to close the lasso (Figure 27)
| Numbered figure captions |
|---|
| SubtitleText | Classifying glioma cells (green) |
|---|
| AnchorName | Classifying malignant cells |
|---|
|
 Image Removed Image Removed
|
- Click Classify selection
- Name the classification
- Open the Classify tool and click Classify selection
- Name the classification Glioma
- Click Save
With every cell from every sample classified, we can view the number of cells classified into each cell type for each sample on the classification summary page.
...
- Switch to pointer mode by clicking
 Image Added in the top right corner of the plot
Image Added in the top right corner of the plot - Deselect the cells by clicking on any blank space on the plot
The number of cells classified as microglia, oligodendrocytes, and glioma are shown in Classify (Figure 29)
...
| The number of cells for each cell type | | AnchorName |
|---|
|
...
|
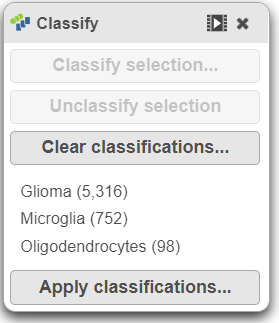 Image Added Image Added
|
...
- Click Apply classifications
- Click Apply to confirm classification
The pipeline view will open and the Classify cells tasks will run, generating a new green-layer Classified groups data node (Figure 29).
- classifications in the Classify icon (Figure 30)
| Numbered figure captions |
|---|
| SubtitleText | Apply classifications to the data project |
|---|
| AnchorName | Apply classifications to the data project |
|---|
|
 Image Added Image Added
|
- Name the classification attribute Cell type (multi-sample) (Figure 31)
- Click Run
| Numbered figure captions |
|---|
| SubtitleText | Classify cells tasks from Name the cell-level attribute |
|---|
| AnchorName | Classification attribute name multi-sample t-SNE plot |
|---|
| AnchorName | Classify cells task |
|---|
|
 Image Removed Image Removed
|
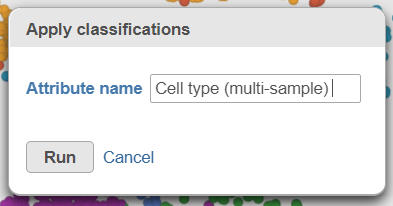 Image Added Image Added
|
The new attribute is now available for downstream analysis.
- Click on the Glioma (multi-sample) project name at the top to go back to the Analyses tab
- Your browser may warn you that any unsaved changes to the data viewer session will be lost. Ignore this message and proceed to the Analyses tab
|
...

























































