Analysis of The list, LCLs vs B cells, includes differentially methylated loci in humans and mice typically includes exclusion of the probes on X and Y chromosomes because of the difficulties caused by the inactivation of one X chromosome in female samples. To filter out probes from the X and Y chromosomes, we first need to annotate the ANOVA spreadsheet with chromosome locations.for locations across the genome; however, in many cases we may want to focus on loci located in particular regions of the genome. To filter our list to include only regions of interest, we can use the annotations provided by Illumina and the interactive filter in Partek Genomics Suite.
- Select LCLs_Vs_B_cells from the spreadsheet tree
- Right-click on the Gene Symbol column
- Select Insert Annotation (Figure 1)
| Numbered figure captions |
|---|
| SubtitleText | Adding an annotation column to the ANOVA results |
|---|
| AnchorName | Adding annotation to ANOVA results |
|---|
|
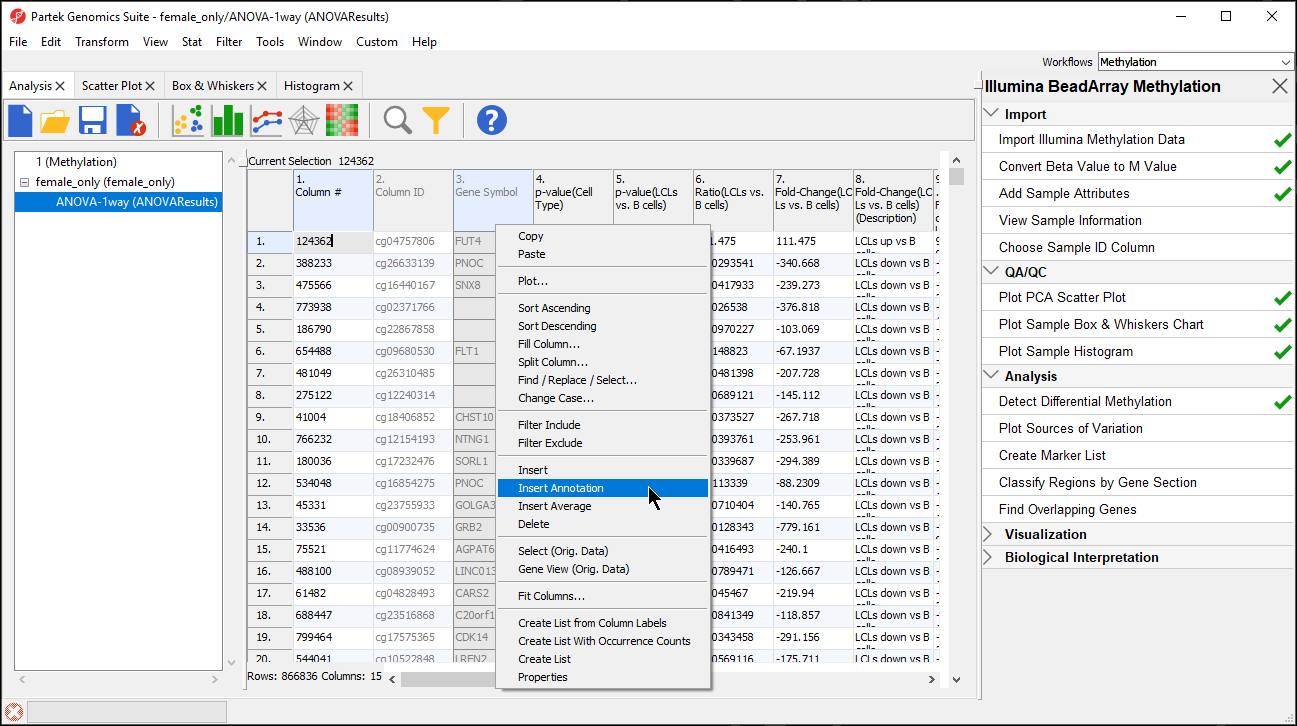 Image Removed Image Removed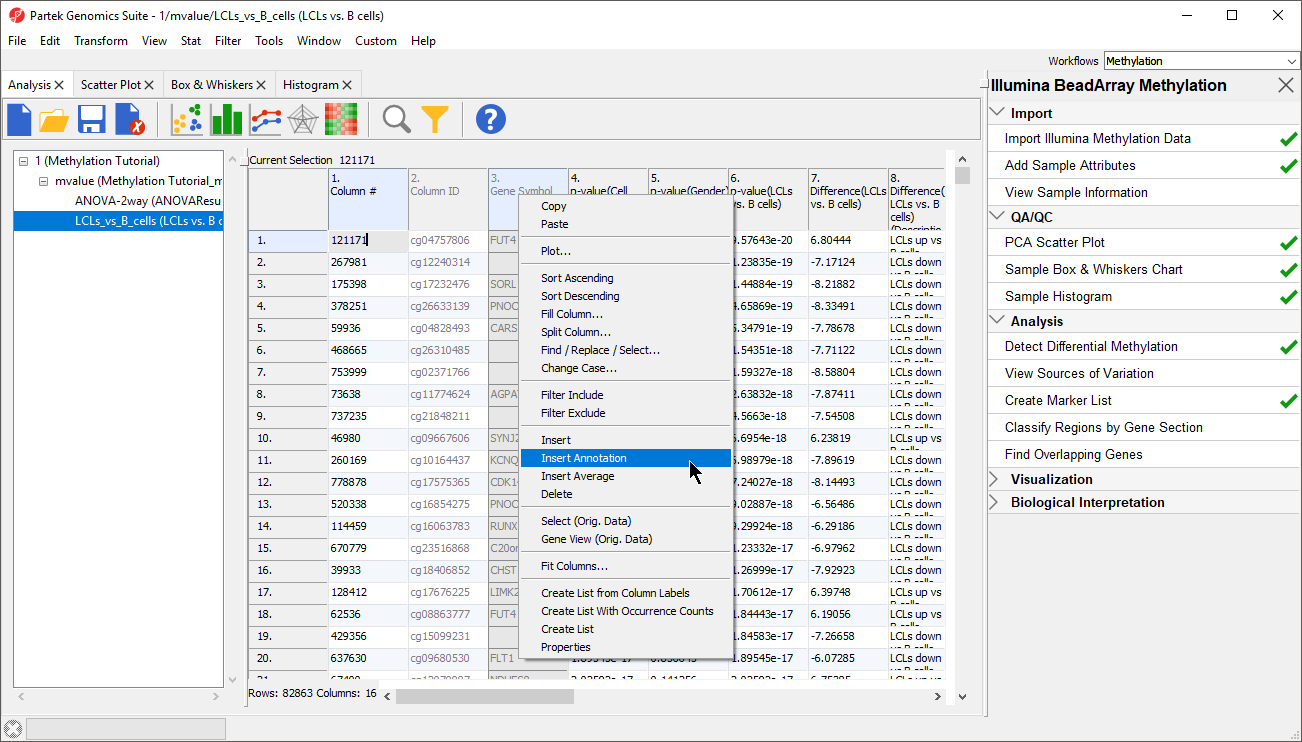 Image Added Image Added
|
- Select the Add as categorical option
- Select CHRRelation_to_UCSC_CpG_Island (Figure 2)
CpG islands are regions of the genome with an atypically high frequency of CpG sites. CpG islands and their surrounding regions (termed shelf and shore) include many gene promoters and altered methylation in these regions can have a disproportionate effect on gene expression. For example, hyper-methylation of promoter CpG islands is a common mechanism for down-regulating gene expression in cancer.
| Numbered figure captions |
|---|
| SubtitleText | Adding chromosome location to ANOVA results |
|---|
| AnchorName | Adding chromosome location to ANOVA results |
|---|
|
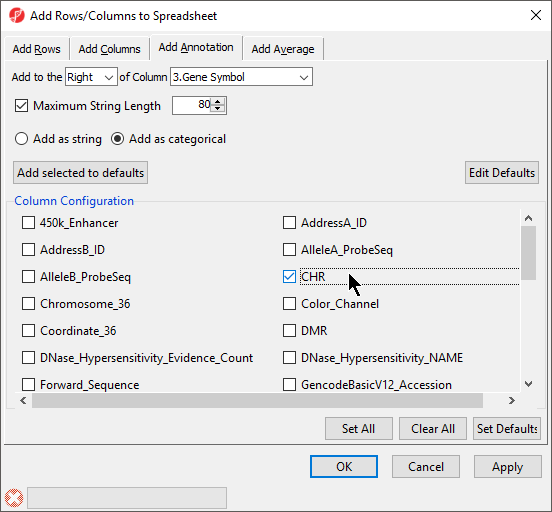 Image Removed Image Removed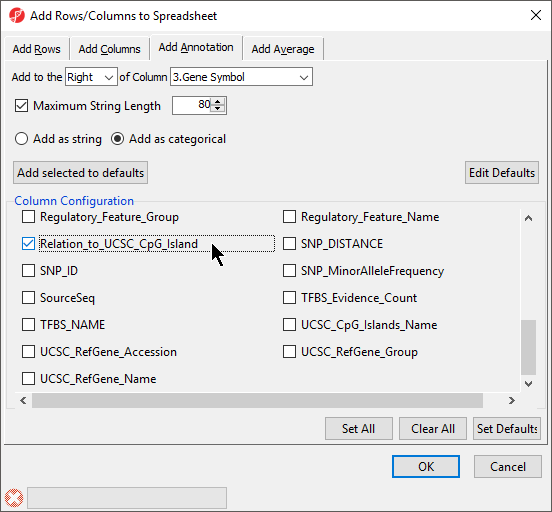 Image Added Image Added
|
- Select OK to add chromosome add Relation_to_UCSC_CpG_Island as a column in next to 3. Gene Symbol
- Select (
 ) from the quick action bar to save the ANOVA-1way 2way (ANOVA Results) spreadsheet with the added annotation
) from the quick action bar to save the ANOVA-1way 2way (ANOVA Results) spreadsheet with the added annotation
Now, we can filter probes by chromosome location.
| Comment |
|---|
There is a step missing here. When inserted, CHR column is Text and needs to be converted to Categorical. |
their relation to CpG islands.- Select (
 ) from the quick action bar to invoke the interactive filter
) from the quick action bar to invoke the interactive filter - Select 4. CHR Relation_to_UCSC_CpG_Island for Column
For categorical columns, the interactive filter displays each category of the selected column as a colored bar. For 4. CHRRelation_to_UCSC_CpG_Island, each bar represents a chromosome with the height of bar representing the number of probes from that chromosome in the selected spreadsheetone of the categories of the UCSC annotation . To filter out a category, left-click on its bar. Right clicking on a bar will include only the selected category. A pop up balloon will show the category label as you mouse over each bar.
- LeftRight-click the X and Y chromosome columns Island column to filter them out other columns (Figure 3)
| Numbered figure captions |
|---|
| SubtitleText | Using Interactive Filter tool to filter out probes by annotation. When pointed to a categorical column, the Interactive Filter tool summarises the content of the column by a column chart. Left-click to exclude a category (two columns were excluded, so they are grayed out), right-click to include only |
|---|
| AnchorName | Using the Interactive Filter |
|---|
|
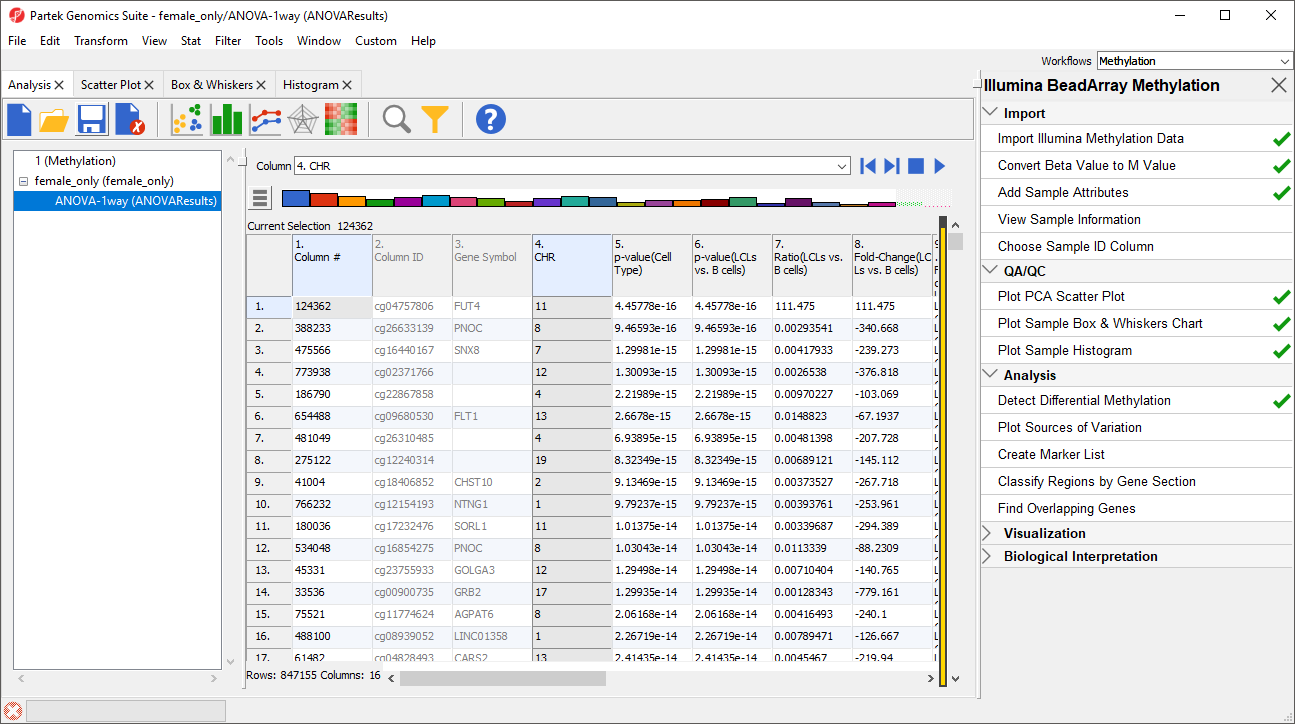 Image Removed Image Removed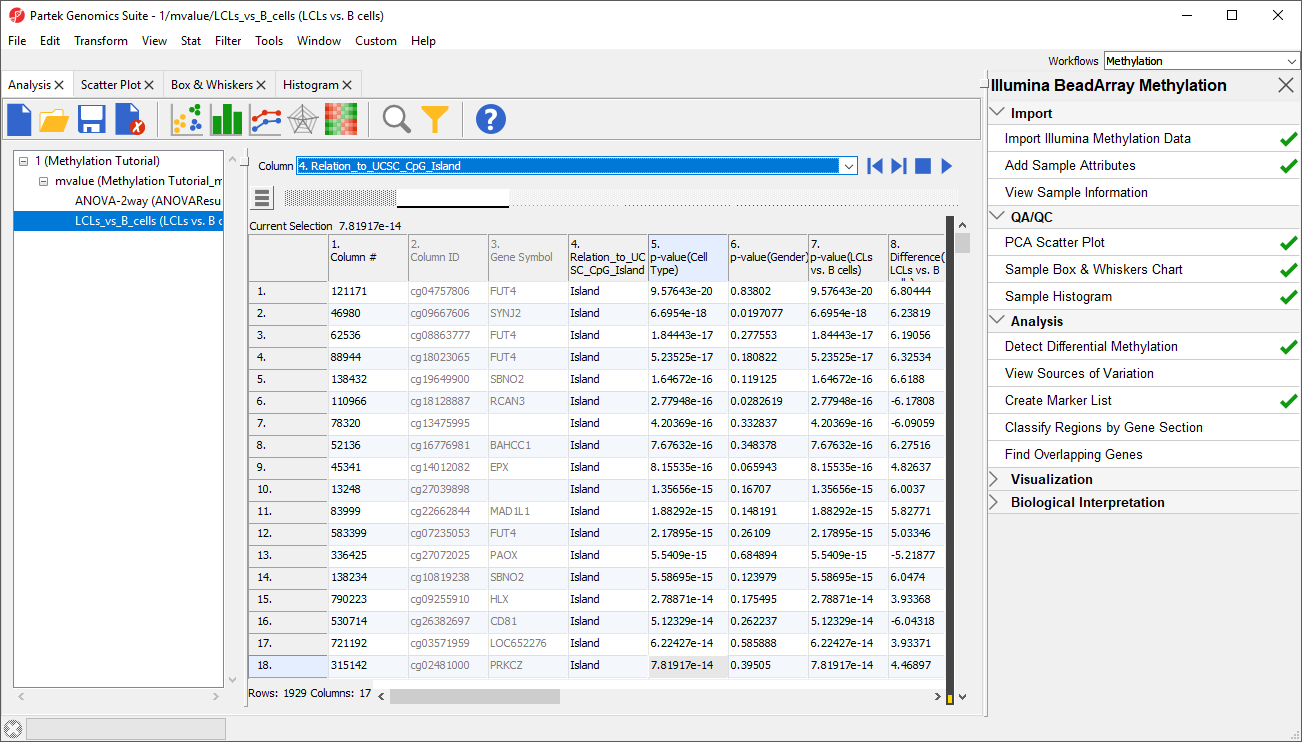 Image Added Image Added
|
The yellow and black bar on the right-hand side of the spreadsheet panel shows the fraction of excluded cells in black and included cells in yellow. Right-clicking this bar brings up an option to clear the filter.
Now that we have filtered out probes on the X and Y chromosomesthat are not in CpG islands, we will create a spreadsheet containing only these probes on the autosomes.
- Right click on the ANOVA-1way (ANOVAResults)the LCLs vs. B cells spreadsheet in the spreadsheet tree panel (Figure 4)
| Numbered figure captions |
|---|
| SubtitleText | Cloning a filtered spreadsheet creates a new spreadsheet with only the included cells |
|---|
| AnchorName | Cloning a filtered spreadsheet |
|---|
|
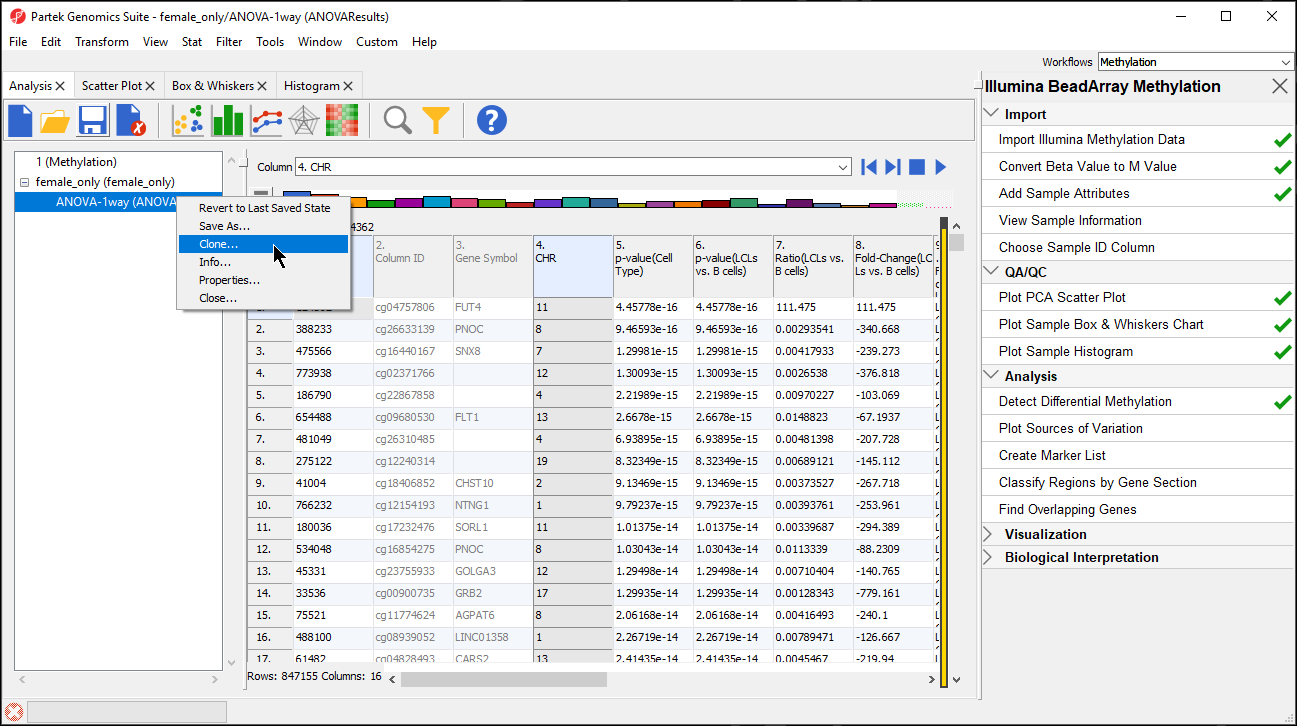 Image Removed Image Removed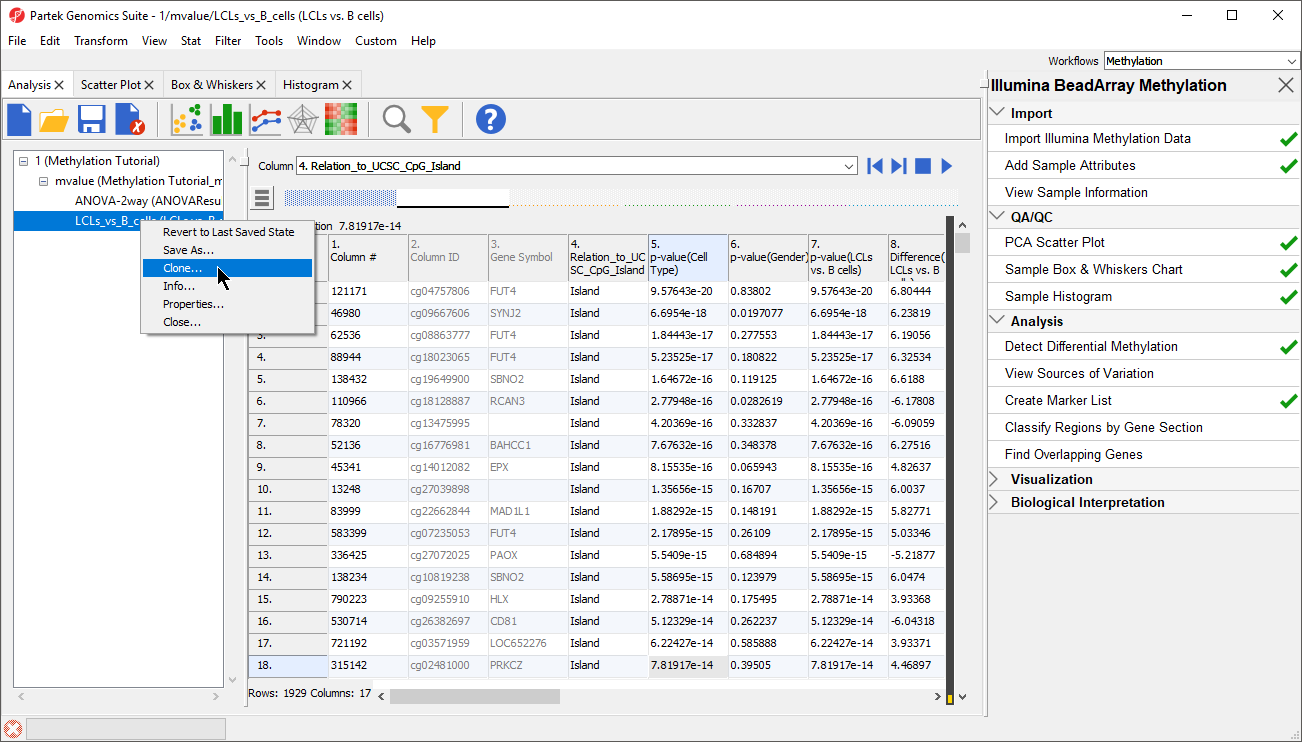 Image Added Image Added
|
- Select Clone
- Rename the new spreadsheet ANOVA-1way_autosomal LCLs_vs_B_cells_CpG_Islands using the Clone Spreadsheet dialog
- Select female_only mvalues from the Create new spreadsheet as a child spreadsheet: drop-down menu (Figure 5)
- Select OK
| Numbered figure captions |
|---|
| SubtitleText | Renaming and configuring filtered spreadsheet |
|---|
| AnchorName | Configuring Clone Spreadsheet |
|---|
|
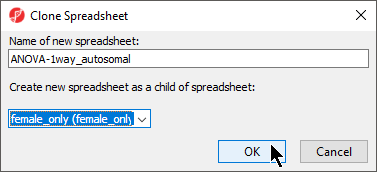 Image Removed Image Removed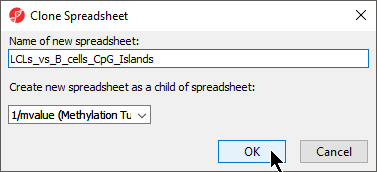 Image Added Image Added
|
- Select (
 ) from the quick action bar to save the filtered spreadsheet
) from the quick action bar to save the filtered spreadsheet - Specify a name for the spreadsheet, we chose autosomalLCLs_vs_B_cells_CpG_Islands, using the Save File dialog
- Select Save to save the spreadsheet
It is best practice to occasionally You may want to save the project you are working on. Let's take the opportunity to do this now.
- Select File from the main command toolbar
- Select Save Project...
- Specify a name for the project, we chose Methylation, using the Save File dialog
- Select Save to save the project
Saving the project saves the identity and child-parent relationships of all spreadsheets displayed in the spreadsheet tree. This allows us to open all relevant spreadsheets for our analysis by selecting the project file. before proceeding to the next section of the tutorial.
...









