Join us for a webinar: The complexities of spatial multiomics unraveled
May 2
Page History
...
- Select Create Marker List from the Analysis section of the Illumina BeadArray Methylation workflow
- Select anova-1way_autosomal (autosomal) as the source spreadsheet from the left-hand panel
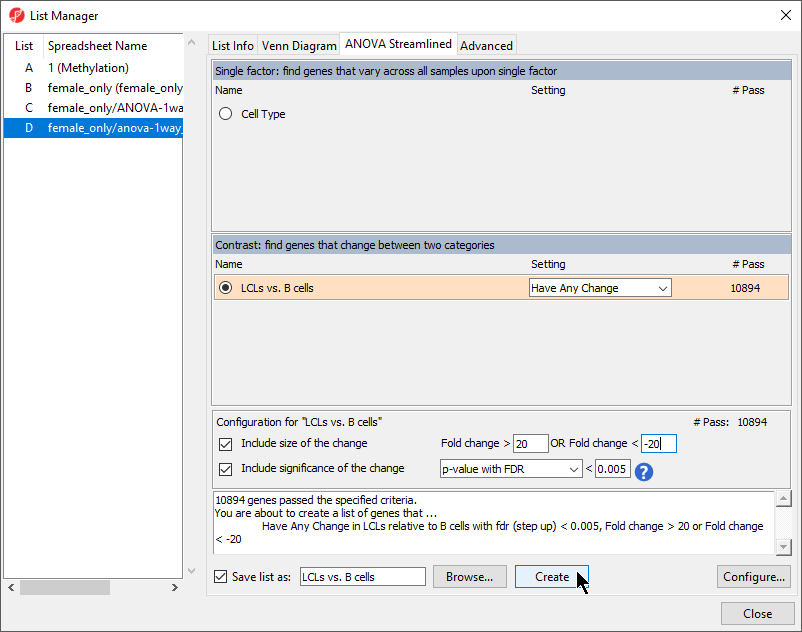
- Select LCLs vs. B cells from the Contrast: find genes that change between two categories section of the ANOVA Streamlined tab of the List Manager dialog
- Set Fold change > to 20
- Set OR Fold change < to -20
- Set p-value with FDR to 0.005
- Set file name as LCLs vs. B cells
- Leave the rest of the option set to defaults (Figure 1)
| Comment |
|---|
May I suggest filtering by Estimate, instead? Fold-change is not typical for methylation studies. |
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
Overview
Content Tools