Page History
...
There are many useful visualizations, annotations, and biological interpretation tools that can operate on a gene list. In order for these features work with an imported list, an annotation file must be associated with the gene list. Additionally, many operations that work with a list of significant genes (like GO- or Pathway-Enrichment) require comparison against a background of “non-significant” genes. The quickest way to accomplish both is to use the background of “all genes” for that organism provided by an annotation source like RefSeq, Ensembl, etc. in .pannot (Partek® annotationPartek annotation), .gff, .gtf, .bed, tab- or comma-delimited format. If the file is not already in a tab-separated or comma delimited format, you may import, modify, and save the file in the proper file format.
...
- Select the annotation file; in this example, we select a .pannot file downloaded from Partek distributed library file repository (www.partek.com/html/libIndex.txt) – – hg19_refseq_14_01_03_v2.pannot
- Delete or rearrange the columns as necessary; we have placed the column with identifiers (should be unique ID) that correspond to our gene list first
- Select File then Save As Text File... to save the annotation file; we have named it Annotation File (Figure 2)
...
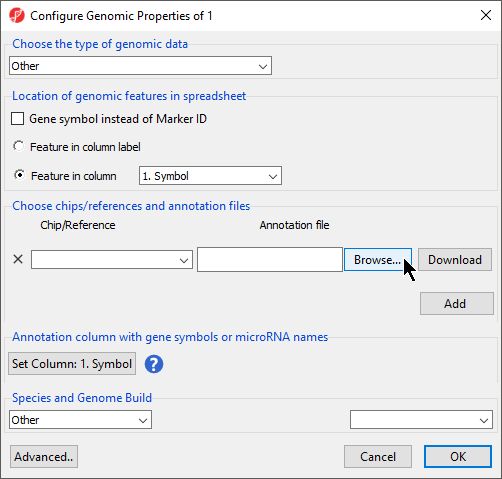
This brings up the Configure Genomic Properties dialog (Figure 3).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
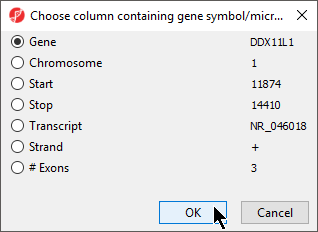
- Select the appropriate column; here the default choice of 1. Symbol is appropriate
- Select OK to return to the Configure Genomic Properties dialog
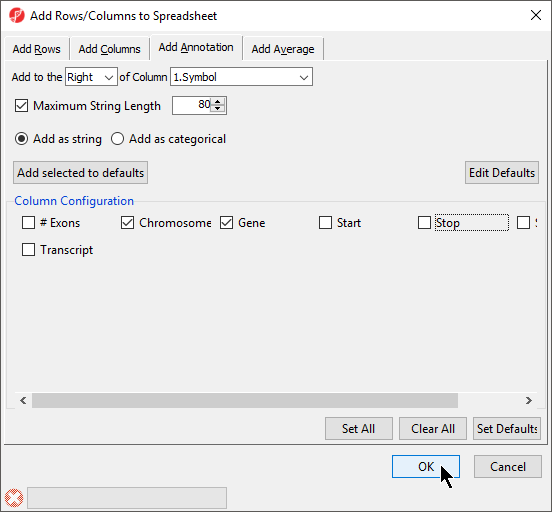
- Select the appropriate species and genome build options; we have selected Homo sapiens and hg19 (Figure 6)
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
|
| Page Turner | ||
|---|---|---|
|
| Additional assistance |
|---|
| Rate Macro | ||
|---|---|---|
|
...