...
| Numbered figure captions |
|---|
| SubtitleText | Location of the Settings link on the main page of Partek Flow |
|---|
| AnchorName | Getting to the settings |
|---|
|
 Image Removed Image Removed Image Added Image Added
|
...
| Numbered figure captions |
|---|
| SubtitleText | Tutorial data sets available through Partek Flow |
|---|
| AnchorName | Importing tutorial data |
|---|
|
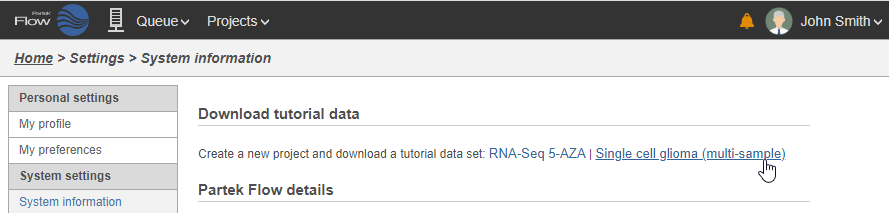 Image Removed Image Removed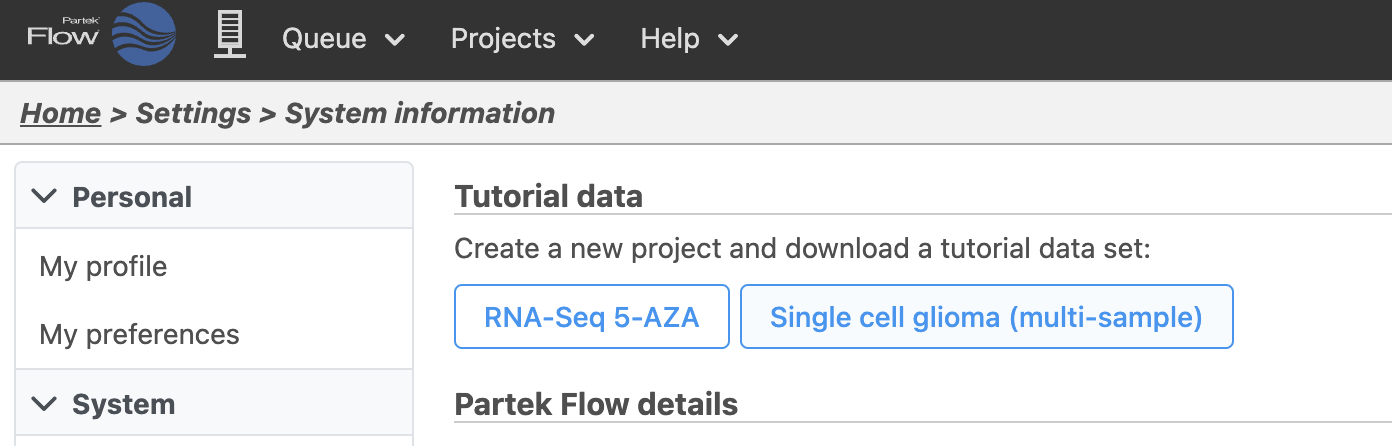 Image Added Image Added
|
- Click Single cell glioma (multi-sample)
...
| Numbered figure captions |
|---|
| SubtitleText | The data tab during tutorial data import |
|---|
| AnchorName | Importing tutorial data in progress |
|---|
|
 Image Removed Image Removed Image Added Image Added
|
You can wait a few minutes for the download to complete, or check the download progress by selecting Queue then View queued tasks... to view the Queue (Figure 4).
...
| Numbered figure captions |
|---|
| SubtitleText | Viewing the queue |
|---|
| AnchorName | Viewing the queue |
|---|
|
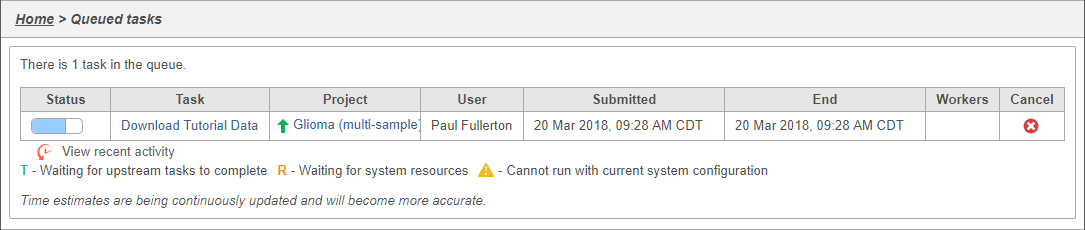 Image Removed Image Removed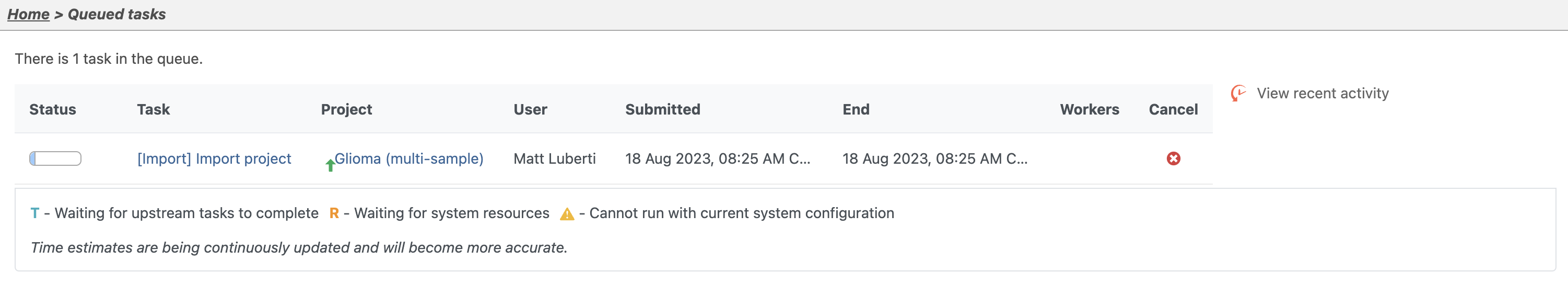 Image Added Image Added
|
Once the download completes, the sample table will appear in the Data Metadata tab, with one row per sample (Figure 5).
...
| Numbered figure captions |
|---|
| SubtitleText | Sample data table listing the name and the number of cells for each sample |
|---|
| AnchorName | Sample data table |
|---|
|
 Image Removed Image Removed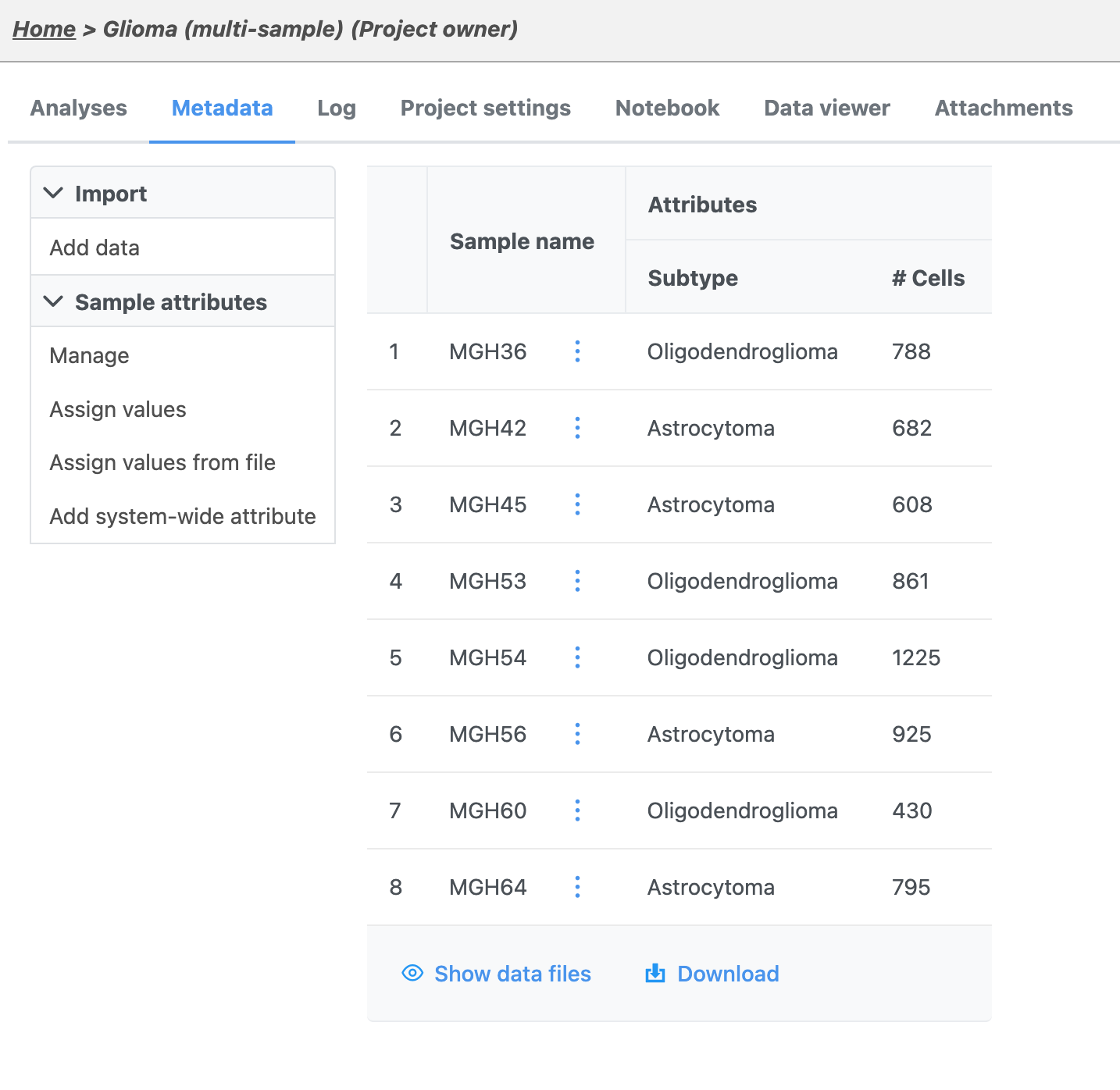 Image Added Image Added
|
The sample table is pre-populated with two sample attributes: # Cells and Subtype. Sample attributes can be added and edited manually by clicking Manage in the Sample attributes menu on the left. If a new attribute is added, click Assign values to assign samples to different groups. Alternatively, you can use the Assign values from a file option to assign sample attributes using a tab-delimited text file. For more information about sample attributes, see here.
For this tutorial, we do not need to edit or change any sample attributes.
...
| Numbered figure captions |
|---|
| SubtitleText | Selecting the Single cell QA/QC task from the task menu |
|---|
| AnchorName | Selecting a task |
|---|
|
 Image Removed Image Removed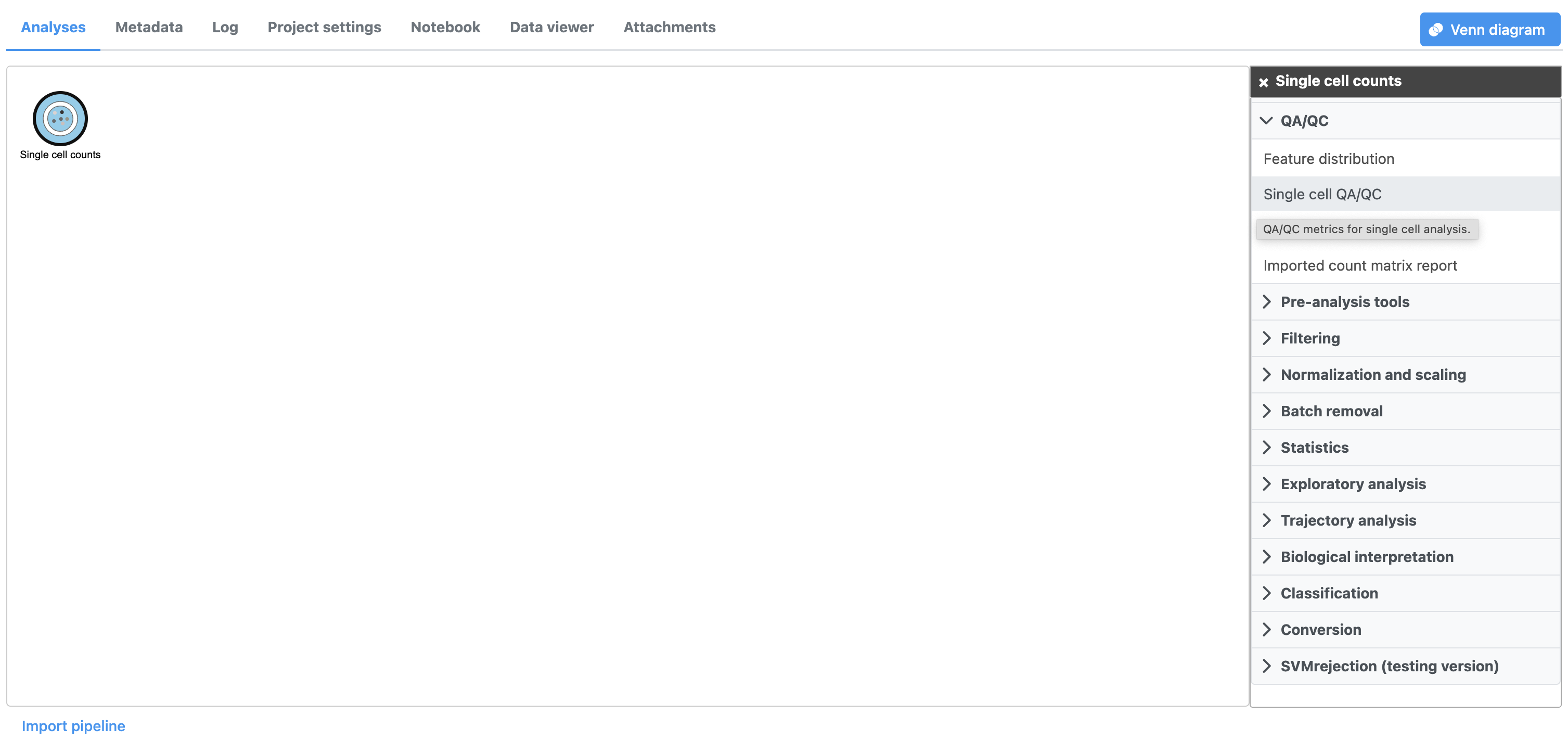 Image Added Image Added
|
A task node, Single cell QA/QC, is produced. Initially, the node will be semi-transparent to indicate that it has been queued, but not completed. A progress bar will appear on the Single cell QA/QC task node to indicate that the task is running.
...
| Numbered figure captions |
|---|
| SubtitleText | Selecting the task report for any task node opens a report with any tables or charts the task produced |
|---|
| AnchorName | Opening task report |
|---|
|
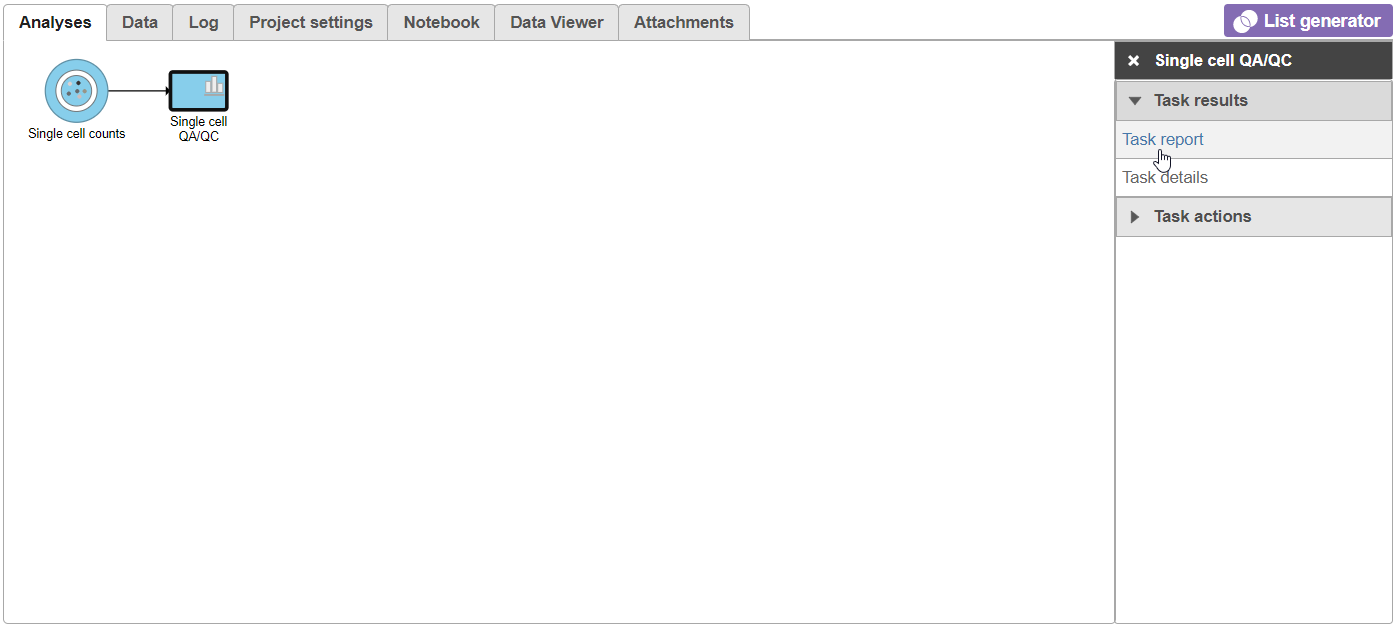 Image Removed Image Removed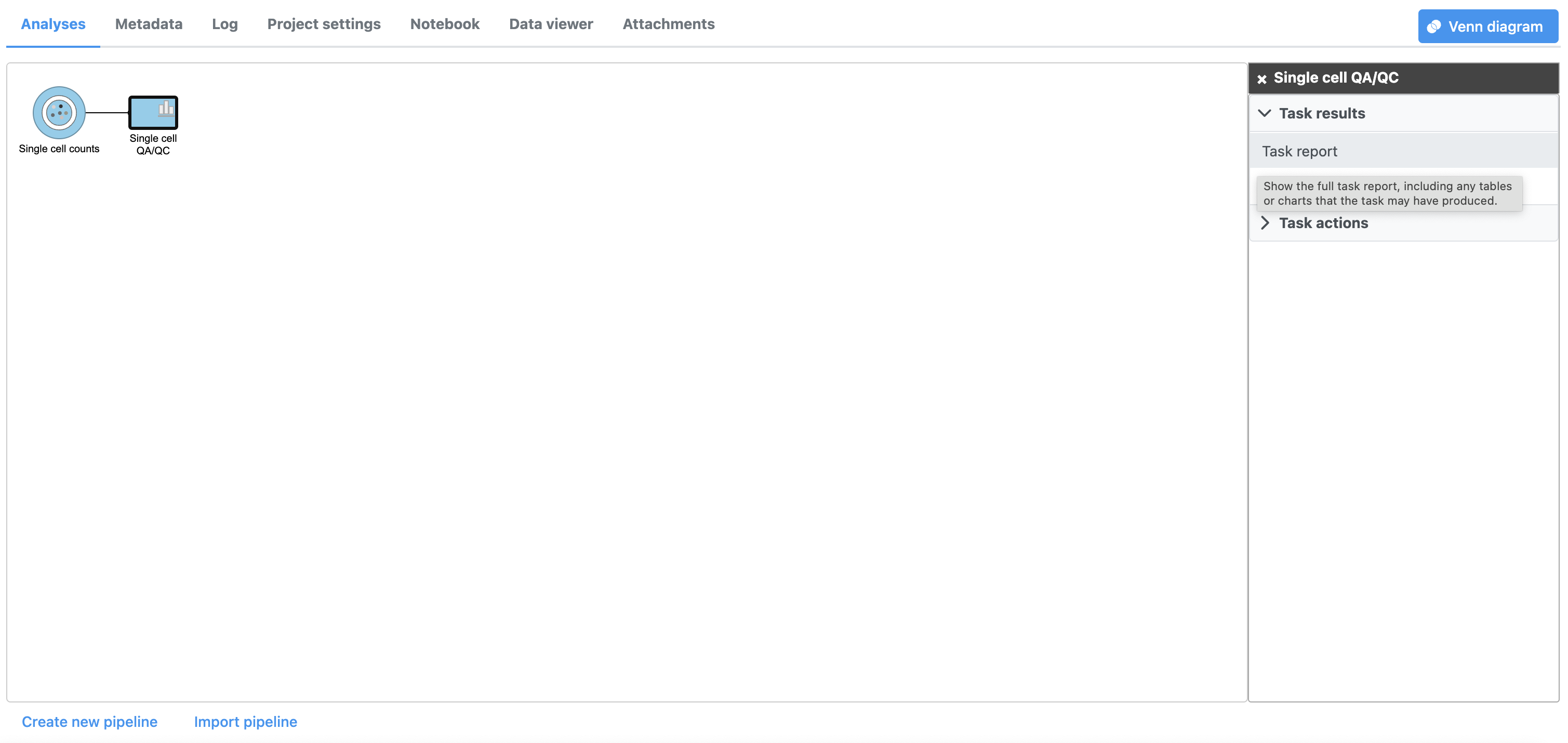 Image Added Image Added
|
The Single cell QA/QC report opens in a new data viewer session. There are interactive violin plots showing the most commonly used quality metrics for each cell from all samples combined (Figure 8). For this data set, there are two relevant plots: the total count per cell and the number of detected genes per cell. Each point on the plots is a cell and the violins illustrate the distribution of values for the y-axis metric. Typically, there is a third plot showing the percentage of mitochondrial counts per cell, but mitochondrial transcripts were not included in the data set by the study authors, so this plot is not informative for this data set.
...
| Numbered figure captions |
|---|
| SubtitleText | Each cell is shown as a point on the plot. Remove the % mitochondrial counts and empty text box using the X icons |
|---|
| AnchorName | Single cell QA/QC report |
|---|
|
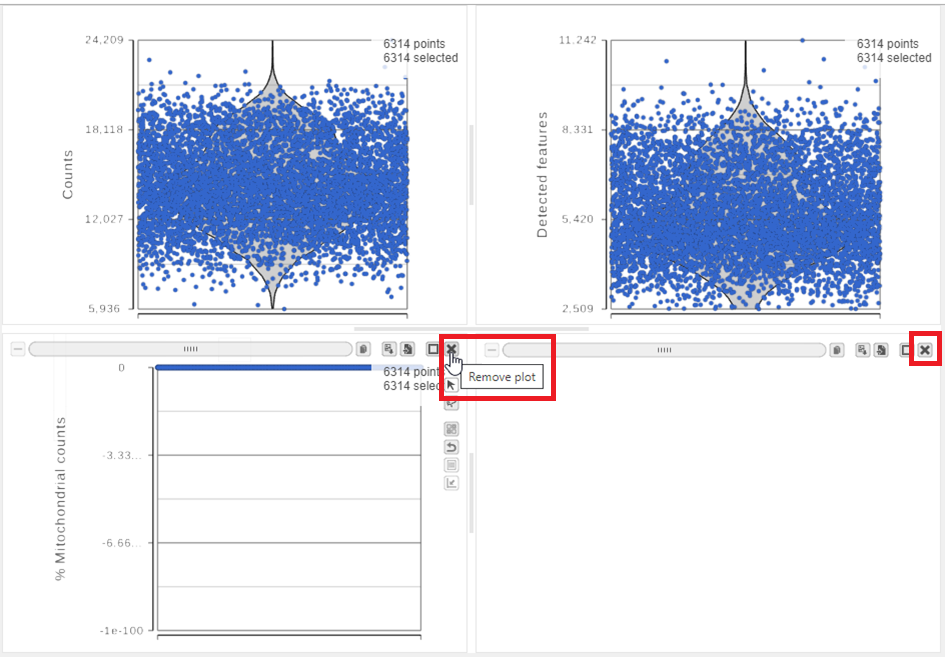 Image Removed Image Removed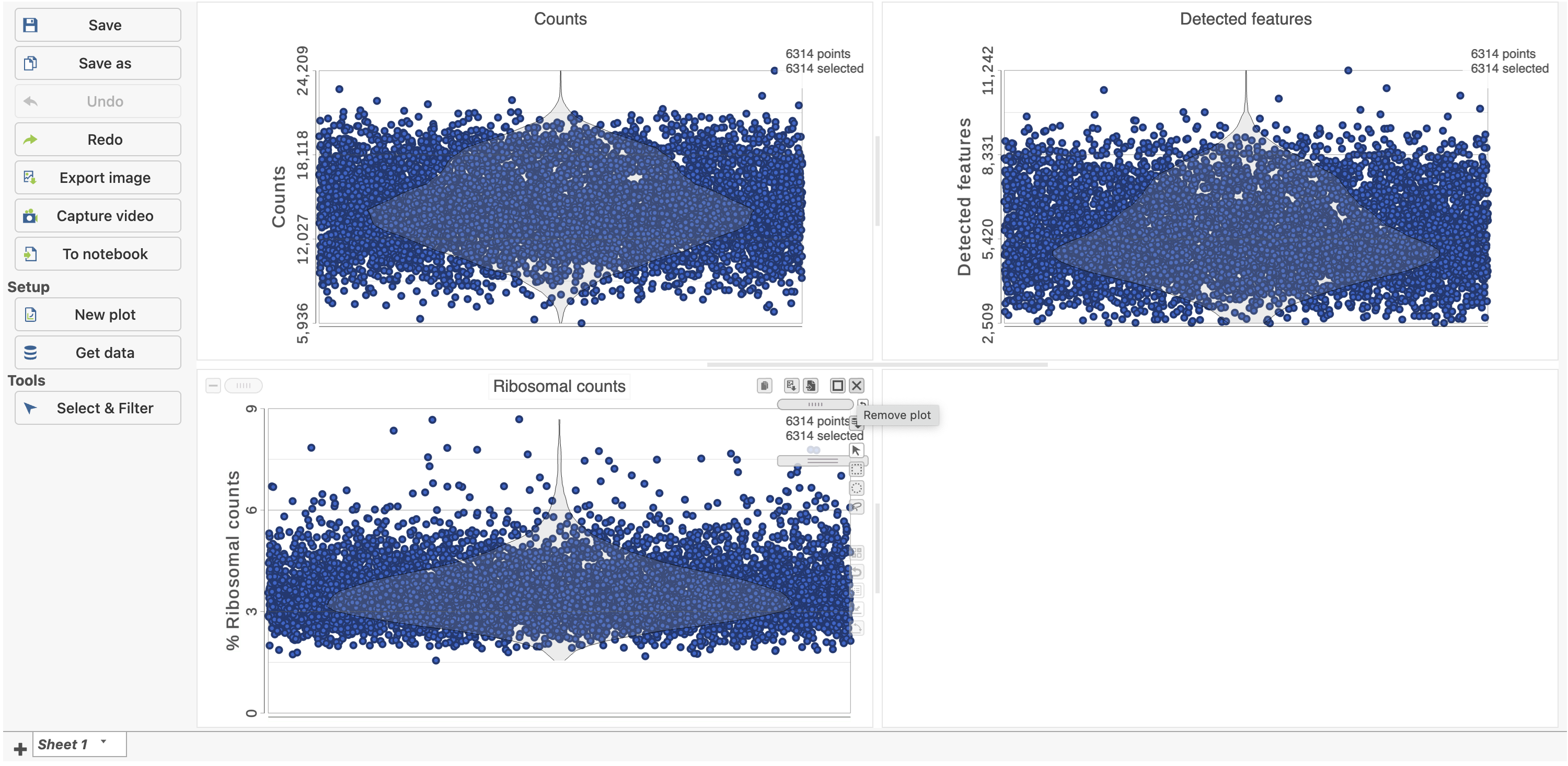 Image Added Image Added
|
The plots are highly customizable and can be used to explore the quality of cells in different samples.
...
| Numbered figure captions |
|---|
| SubtitleText | Click and drag the Sample name attribute onto the X-axis for each plot |
|---|
| AnchorName | Separating cells by sample |
|---|
|
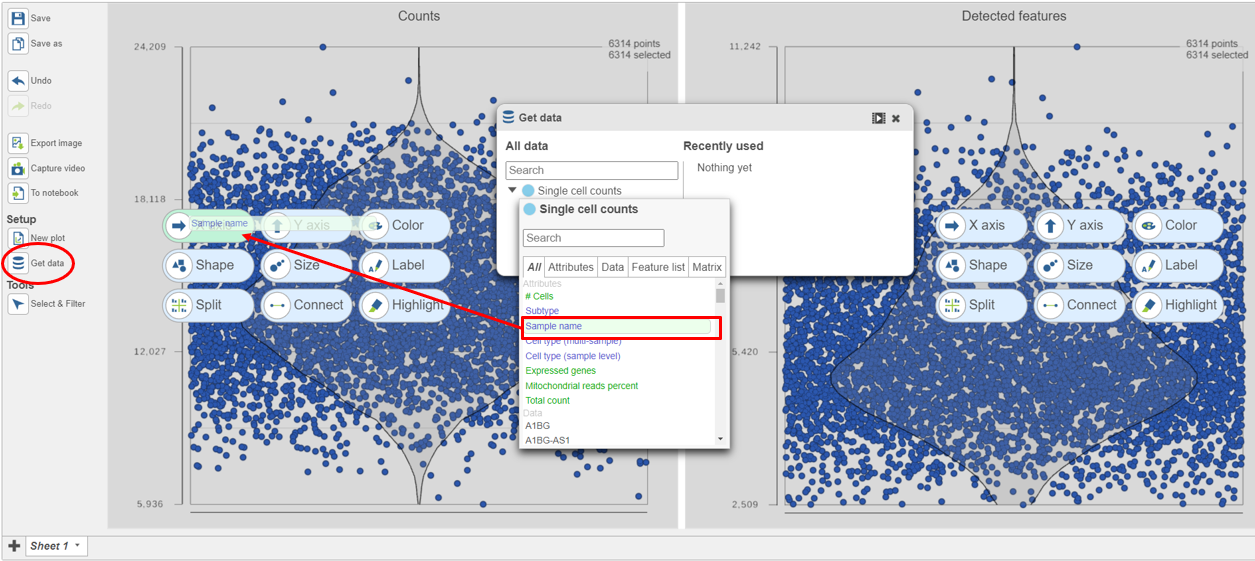 Image Removed Image Removed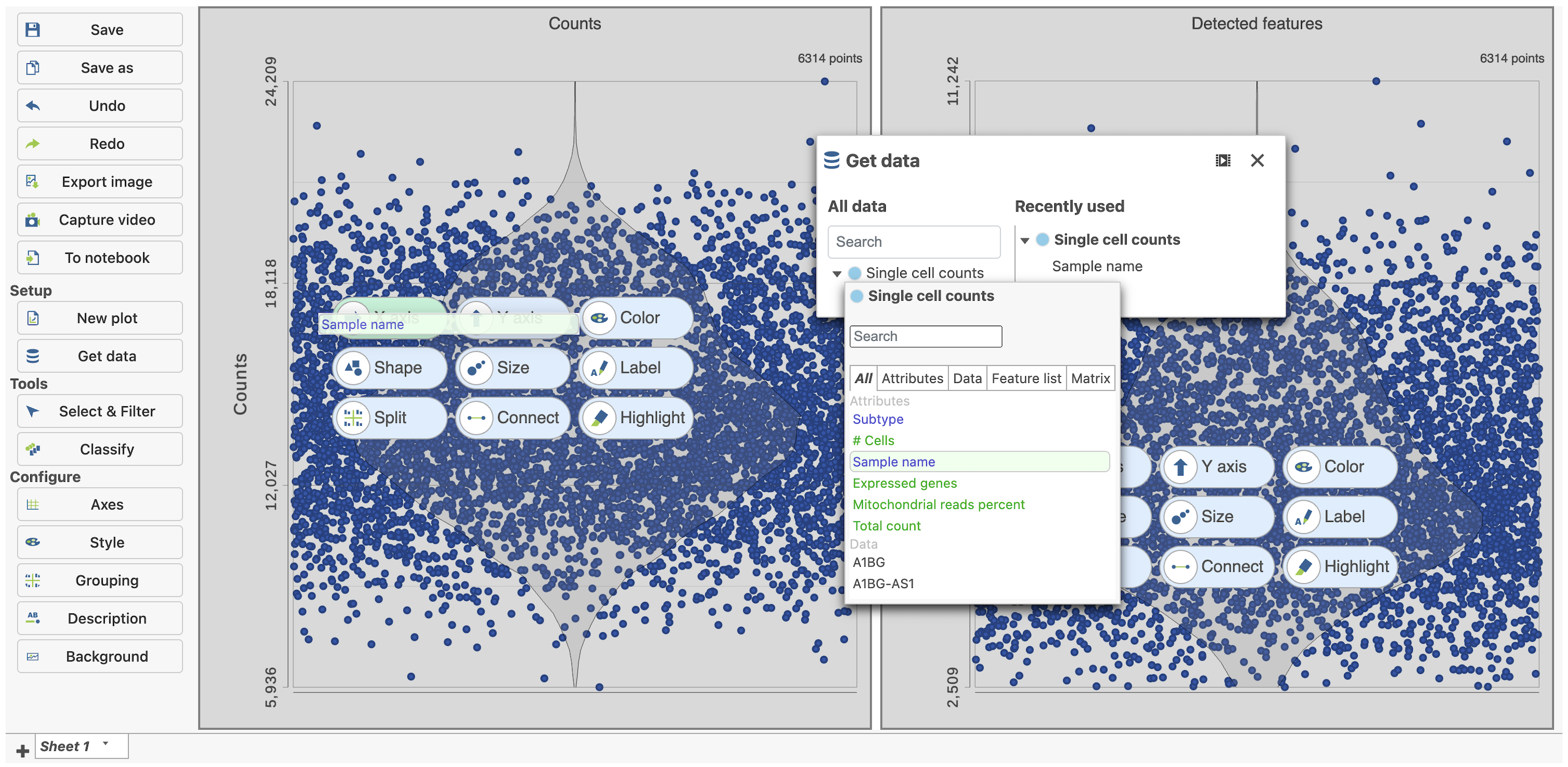 Image Added Image Added
|
The cells are now separated into different samples along the x-axis (Figure 10)
...
| Numbered figure captions |
|---|
| SubtitleText | Counts and detected genes plots can be customized to compare cells from different samples |
|---|
| AnchorName | Single cell QAQC samples on x-axis |
|---|
|
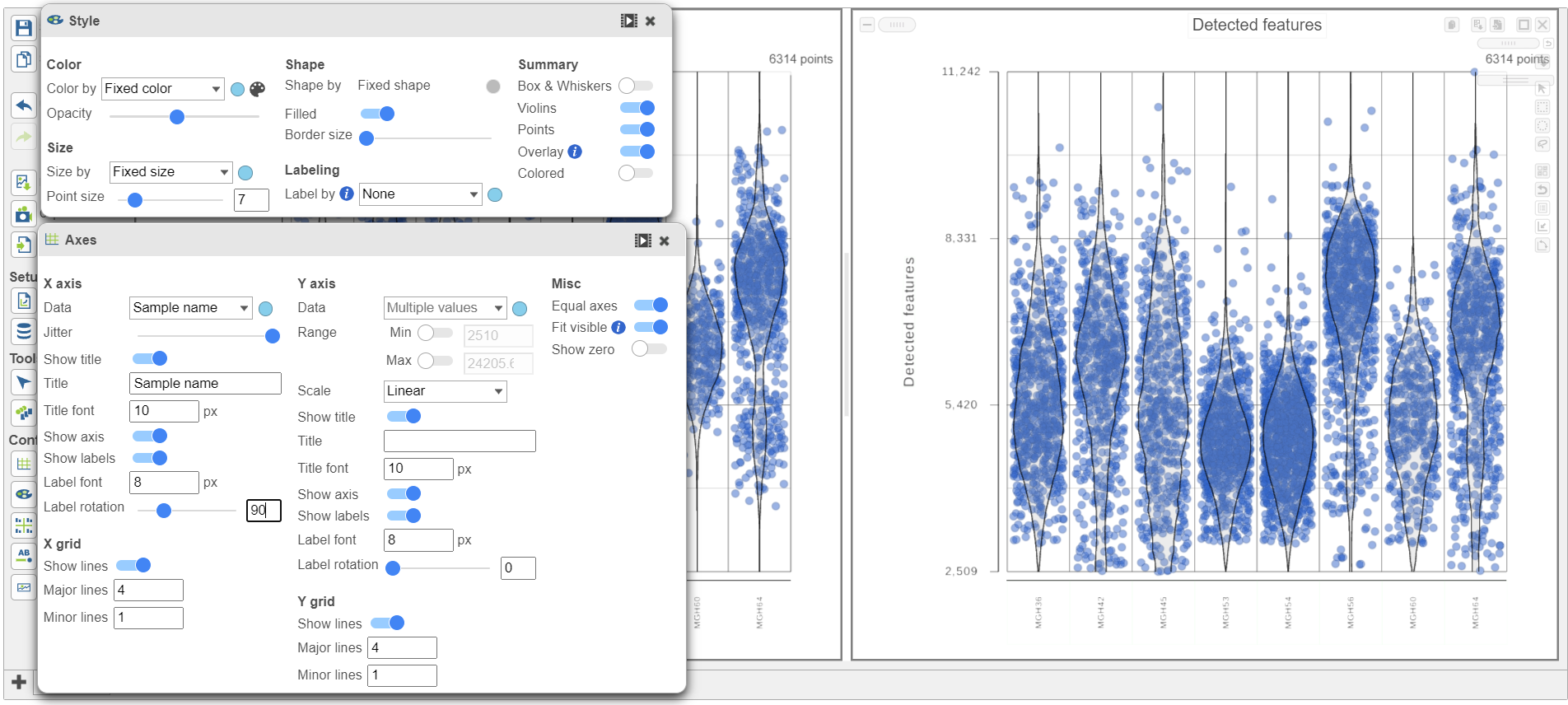 Image Removed Image Removed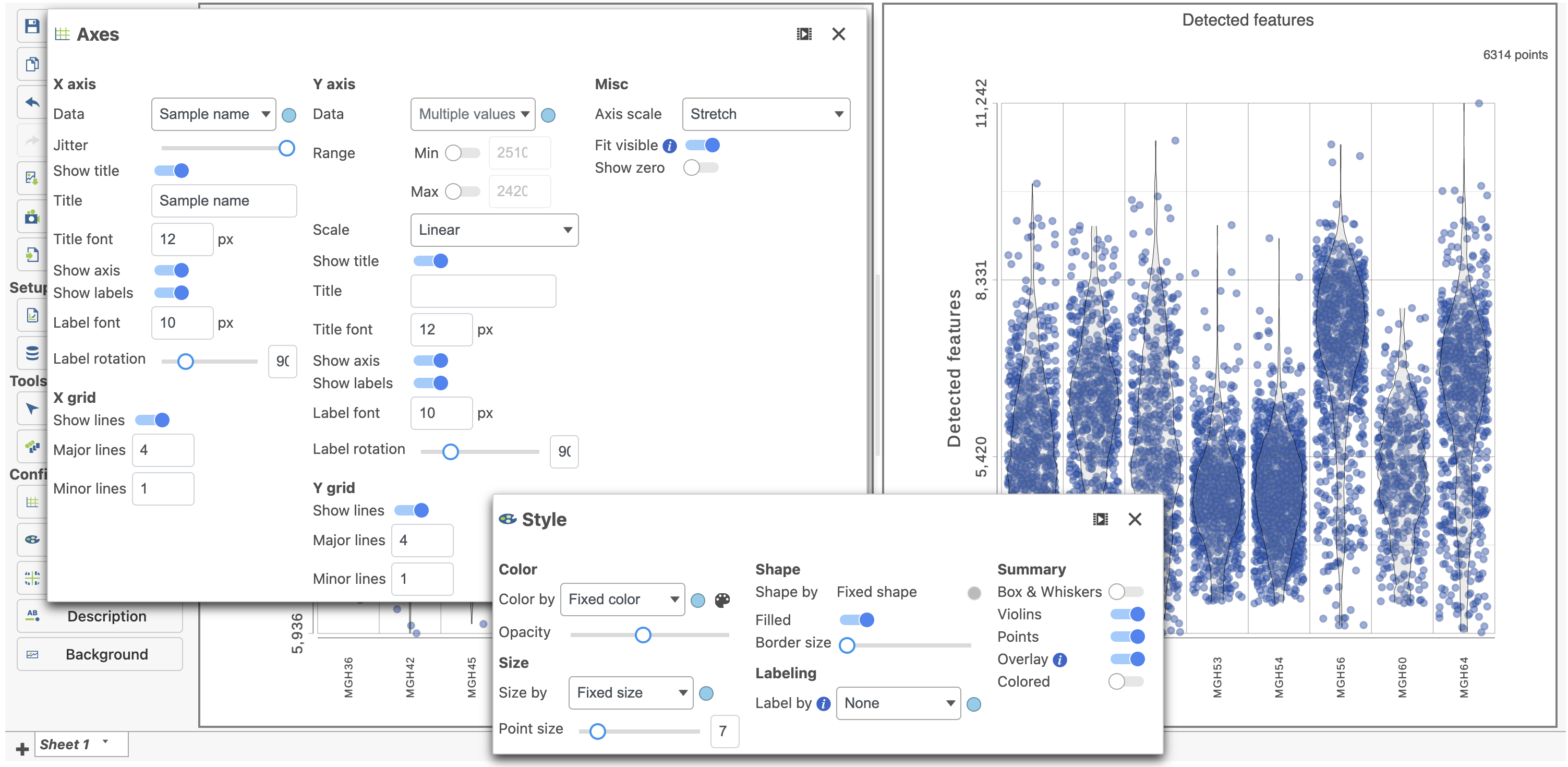 Image Added Image Added
|
Note how both plots were modified at the same time.
...
| Numbered figure captions |
|---|
| SubtitleText | Previewing a filter using the Single cell QA/QC violin plots |
|---|
| AnchorName | Filtering cells by read counts |
|---|
|
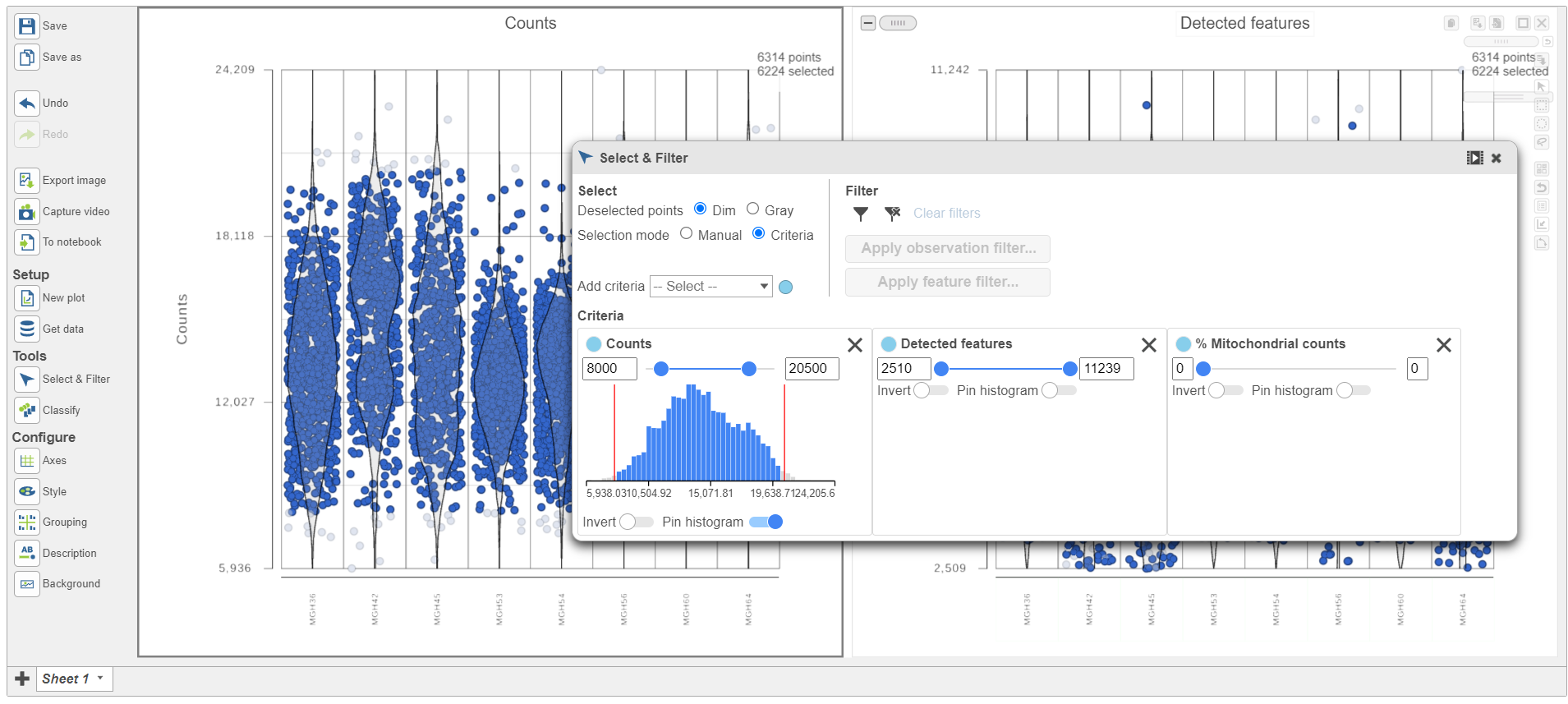 Image Removed Image Removed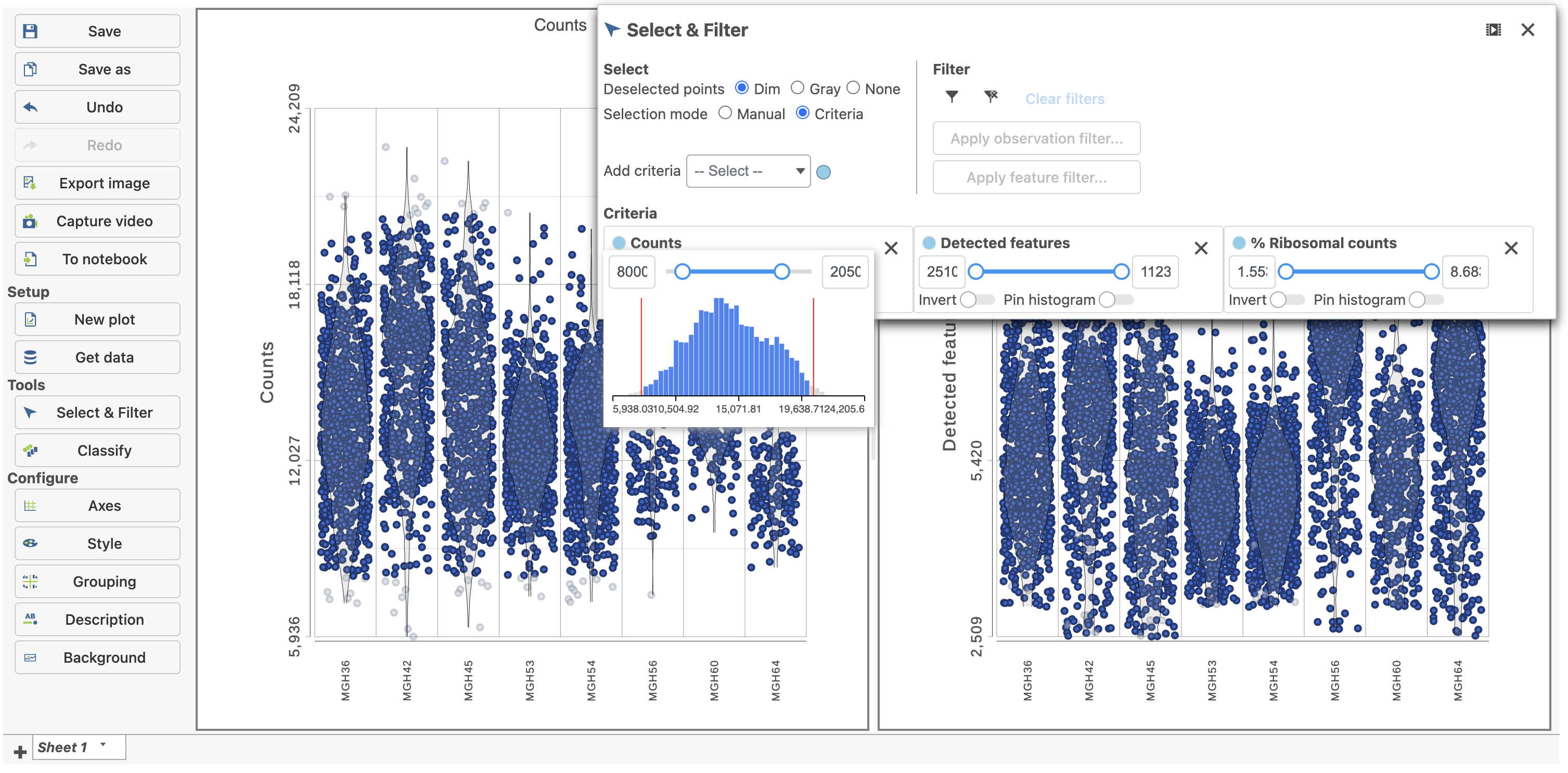 Image Added Image Added
|
Because this data set was already filtered by the study authors to include only high-quality cells, this count filter is sufficient.
...
| Numbered figure captions |
|---|
| SubtitleText | After the Apply filter button is selected, you will be presented with a preview of your pipeline. You need to select the appropriate data node to apply the filtering to |
|---|
| AnchorName | Select Single cell count data node as input for filtering task |
|---|
|
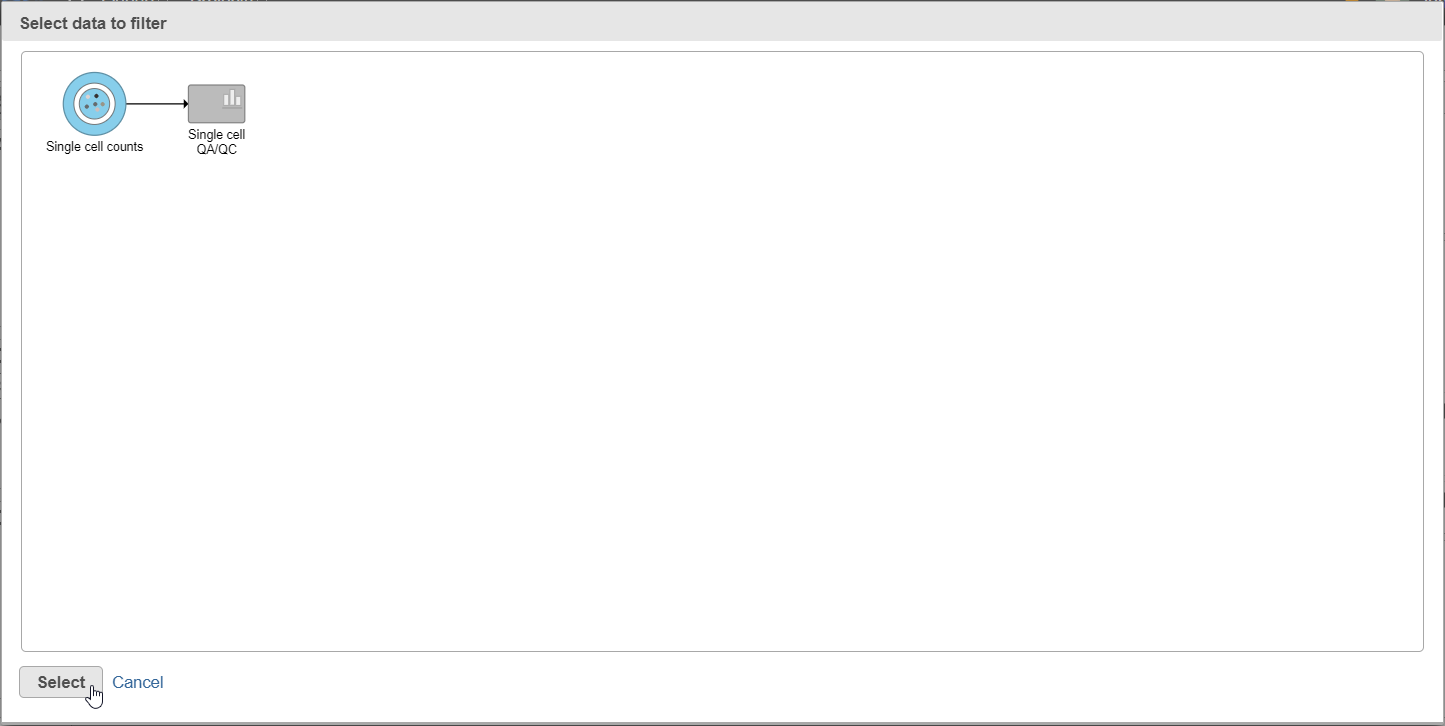 Image Removed Image Removed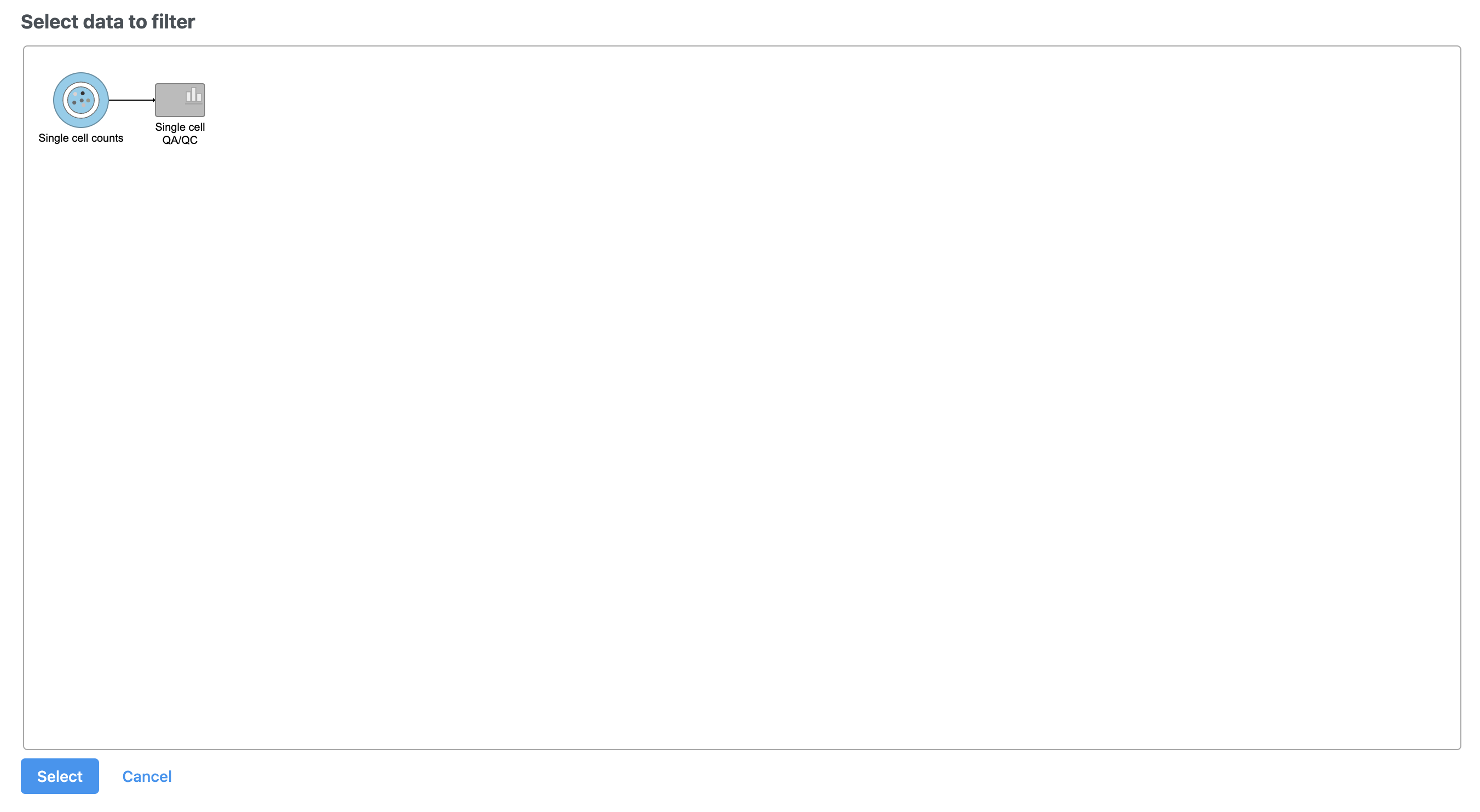 Image Added Image Added
|
A new task, Filter counts, is added to the Analyses tab. This task produces a new Filter counts data node (Figure 13).
...
| Numbered figure captions |
|---|
| SubtitleText | Applying a cell quality filter |
|---|
| AnchorName | Output of Filter cells |
|---|
|
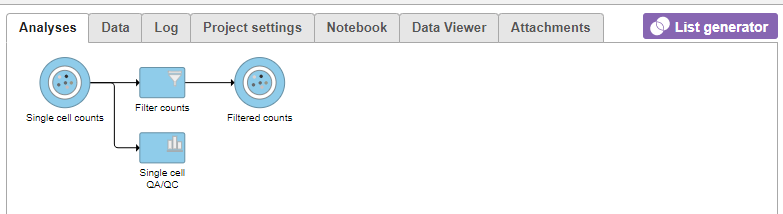 Image Removed Image Removed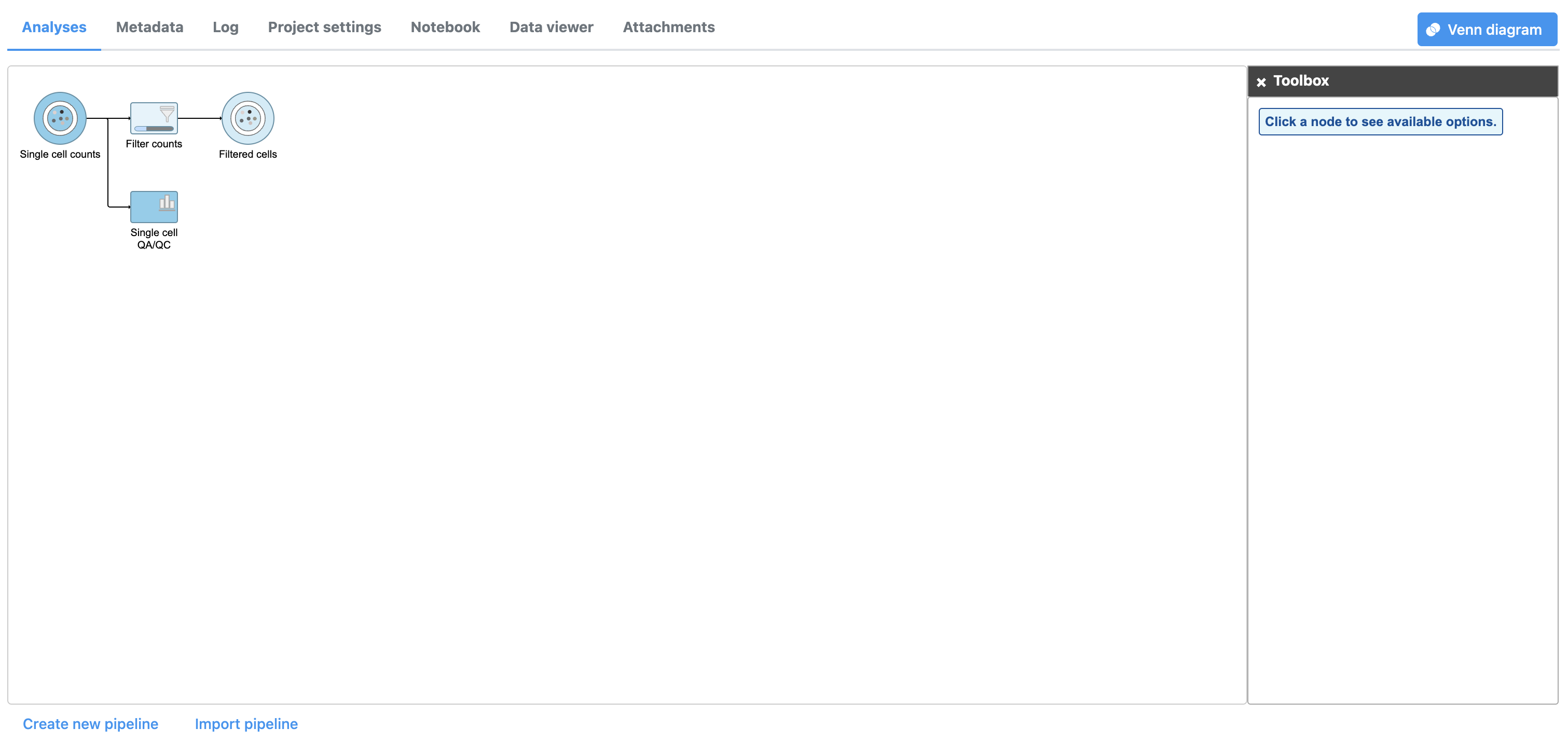 Image Added Image Added
|
Most tasks can be queued up on data nodes that have not yet been generated, so you can wait for filtering step to complete, or proceed to the next section.
...
| Numbered figure captions |
|---|
| SubtitleText | Invoking Filter features |
|---|
| AnchorName | Invoking Filter features |
|---|
|
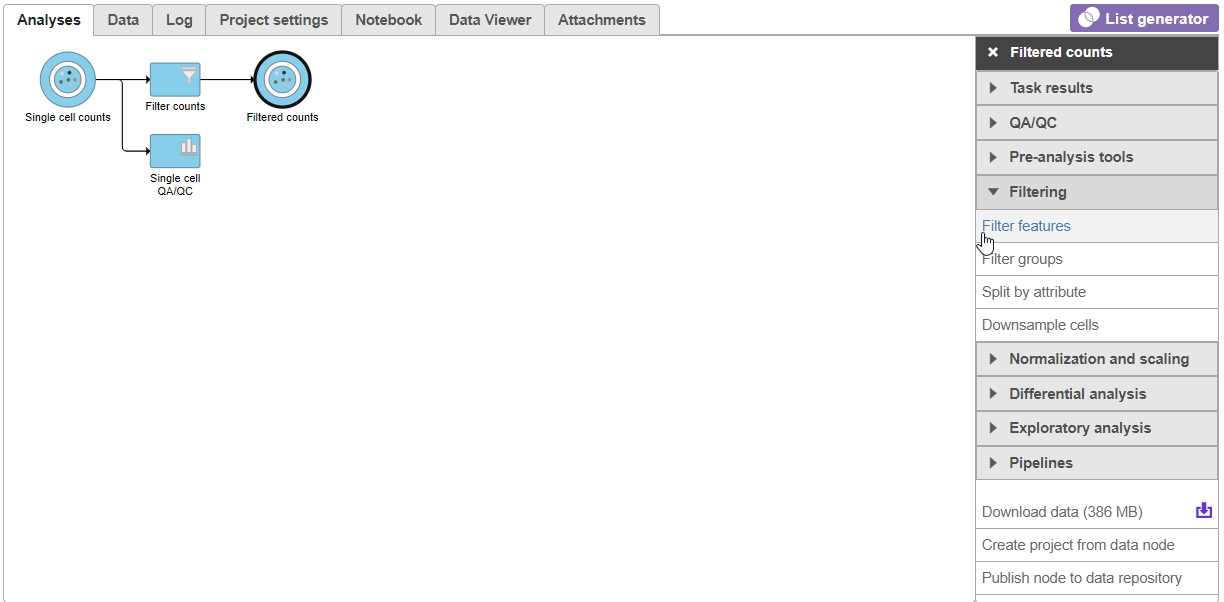 Image Removed Image Removed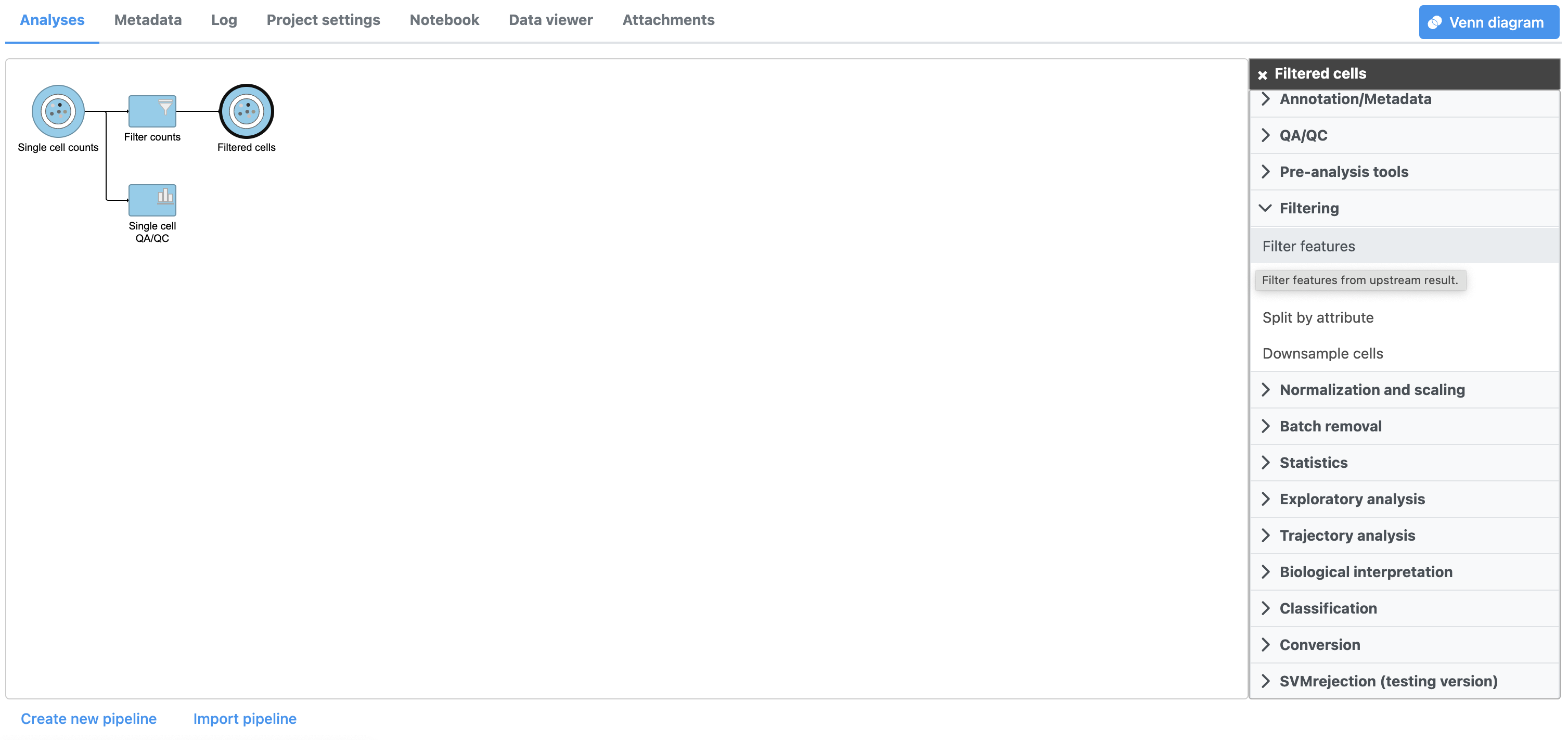 Image Added Image Added
|
There are four categories of filter available - noise reduction, statistics based, feature metadata, and feature list (Figure 15).
...
| Numbered figure captions |
|---|
| SubtitleText | Viewing the filtering options |
|---|
| AnchorName | Filter types |
|---|
|
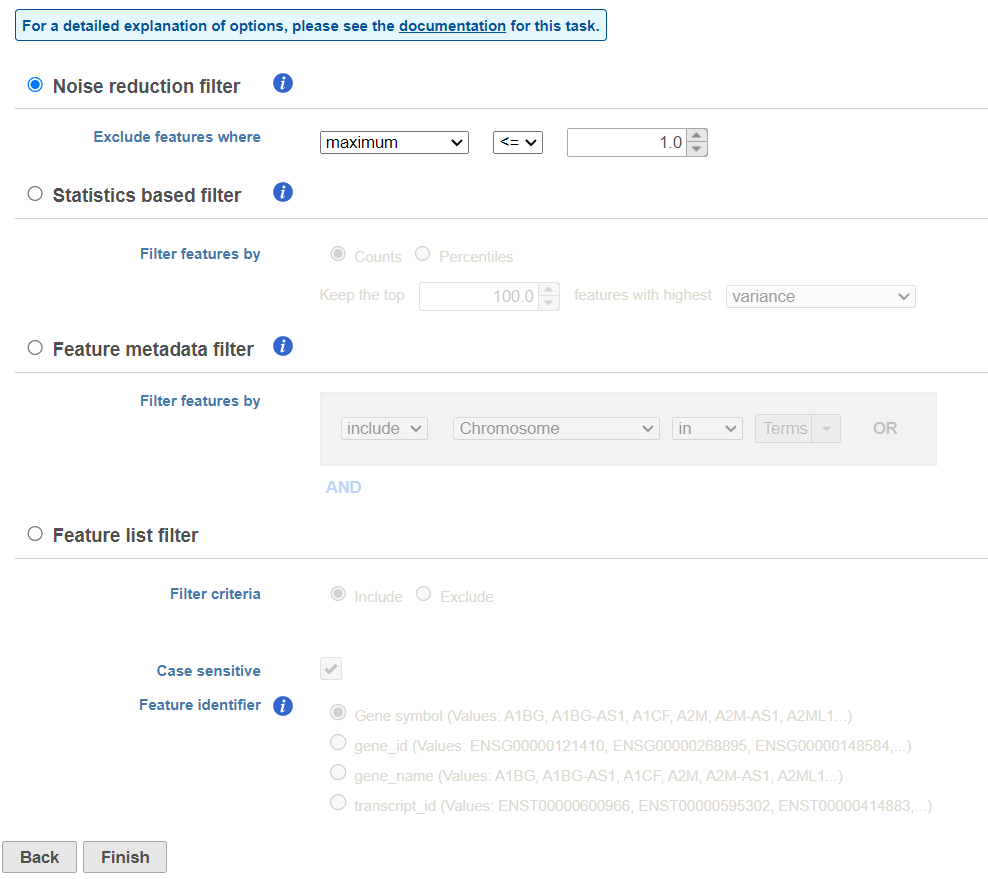 Image Removed Image Removed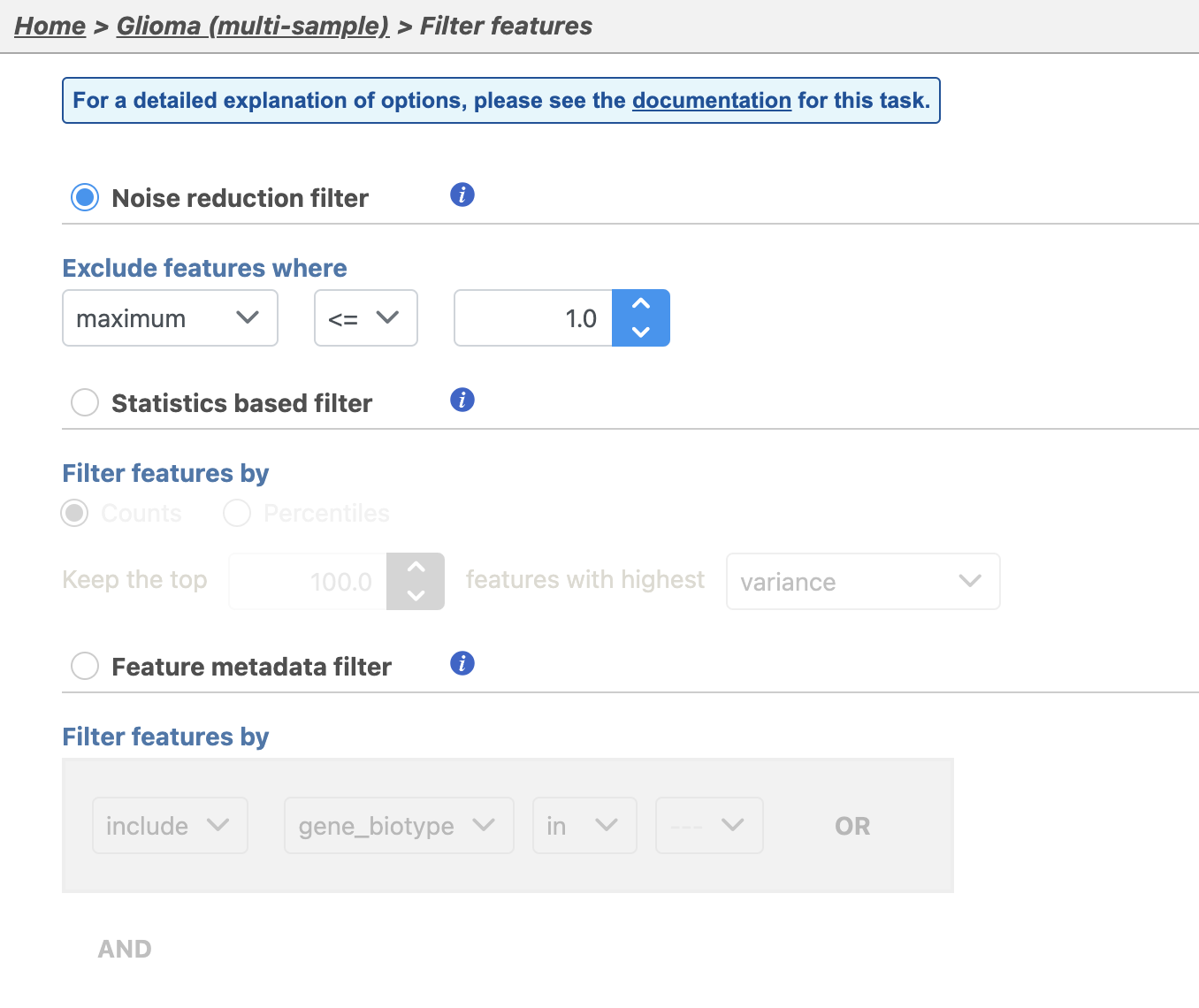 Image Added Image Added
|
The noise reduction filter allows you to exclude genes considered background noise based on a variety of criteria. The statistics based filter is useful for focusing on a certain number or percentile of genes based on a variety of metrics, such as variance. The feature list filter allows you to filter your data set to include or exclude particular genes.
...
| Numbered figure captions |
|---|
| SubtitleText | Configuring a noise reduction filter to exclude genes not expressed in the data set |
|---|
| AnchorName | Configuring a noise reduction filter |
|---|
|
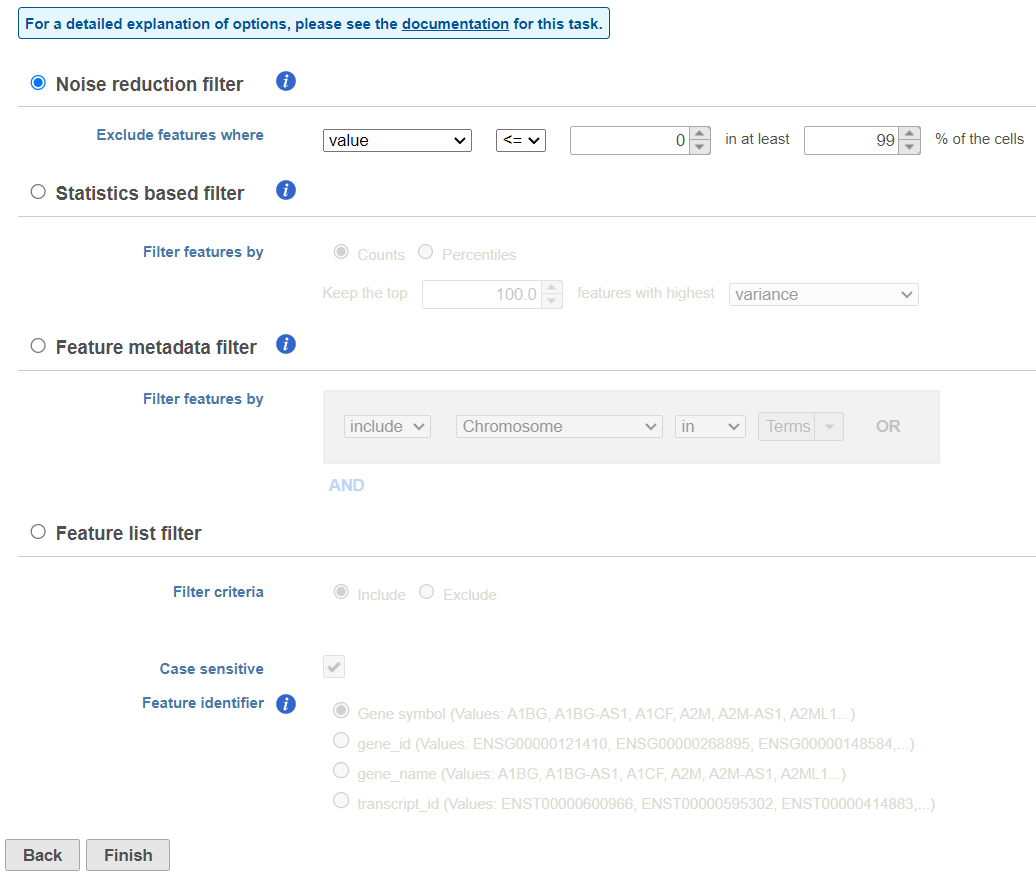 Image Removed Image Removed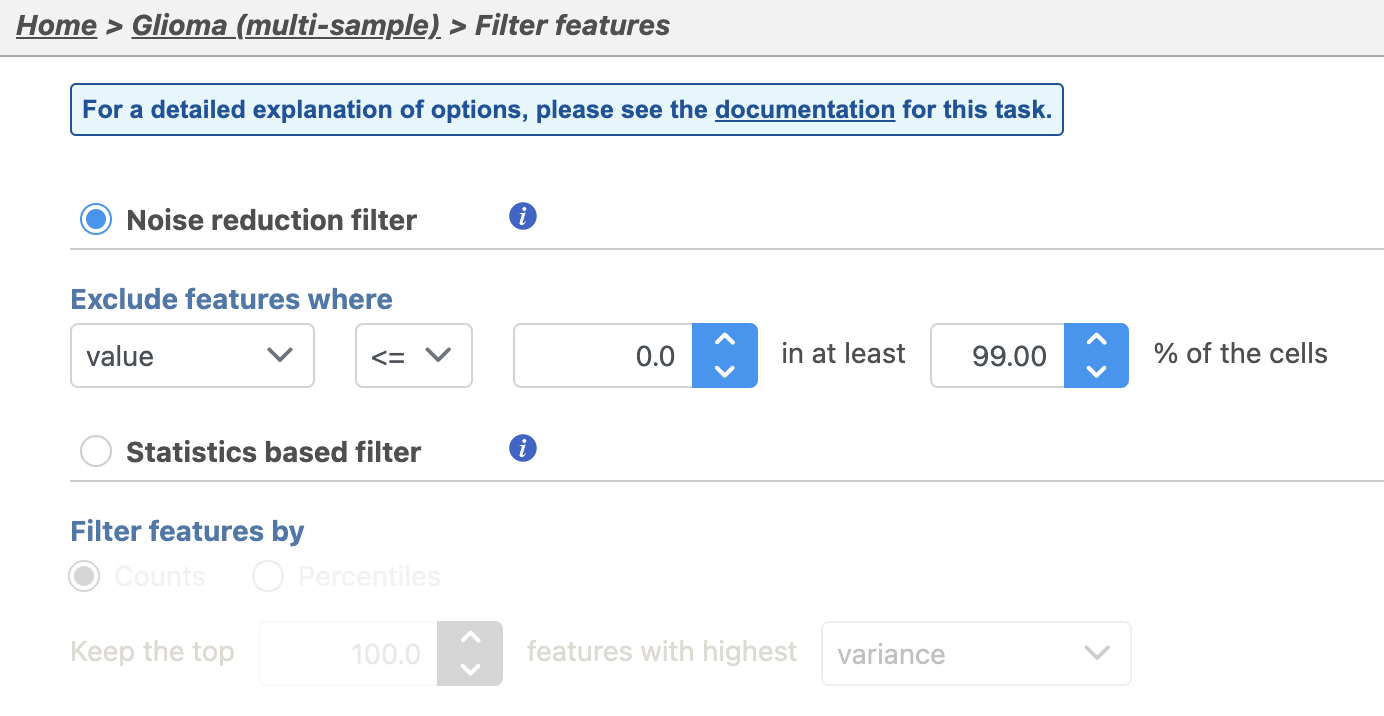 Image Added Image Added
|
This produces a Filtered counts data node. This will be the starting point for the next stage of analysis - identifying cell types in the data using the interactive t-SNE plot.
...































