| Table of Contents |
|---|
| maxLevel | 2 |
|---|
| minLevel | 2 |
|---|
| exclude | Additional Assistance |
|---|
|
Section Heading
Section headings should use level 2 heading, while the content of the section should use paragraph (which is the default). You can choose the style in the first dropdown in toolbar.
Next, we will perform some exploratory analysis on the merged mRNA and protein expression data and visualize the data in preparation to identify cell populations.
PCA
Because the merged count matrix has thousands of features, it is a good idea to reduce the dimensionality of the data for more efficient downstream processing.
- Click the Merged counts data node
- Click Exploratory analysis in the toolbox
- Click PCA
- Click Finish to run the PCA with default settings (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | Run PCA with default settings |
|---|
| AnchorName | PCA task set up |
|---|
|
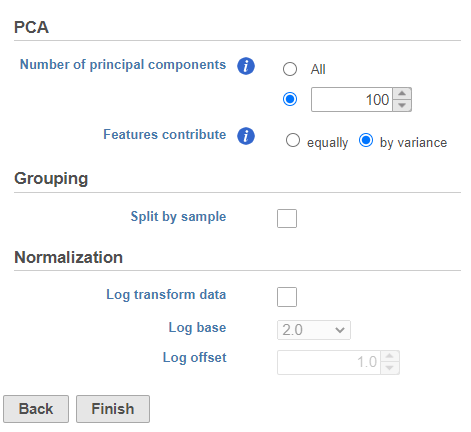 Image Added Image Added
|
A PCA task node will be added to the pipeline under the Analyses tab and a circular PCA output data node will be produced (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | PCA task run on the merged counts data node |
|---|
| AnchorName | PCA output node |
|---|
|
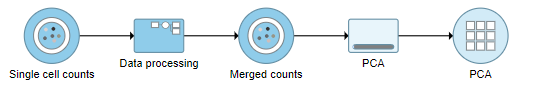 Image Added Image Added
|
Once the task completes, we will inspect the results to decide the optimal number of principal components (PCs) to use in downstream analyses. To do this, we will use a Scree plot.
- Double click the PCA data node to open the task report
The PCA plot will open in a new data viewer session. A 3D scatterplot will be displayed on the canvas (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | Each dot is a different cell. Cells are clustered based on how similar their expression profile is across the combined mRNA and protein data |
|---|
| AnchorName | PCA merged counts |
|---|
|
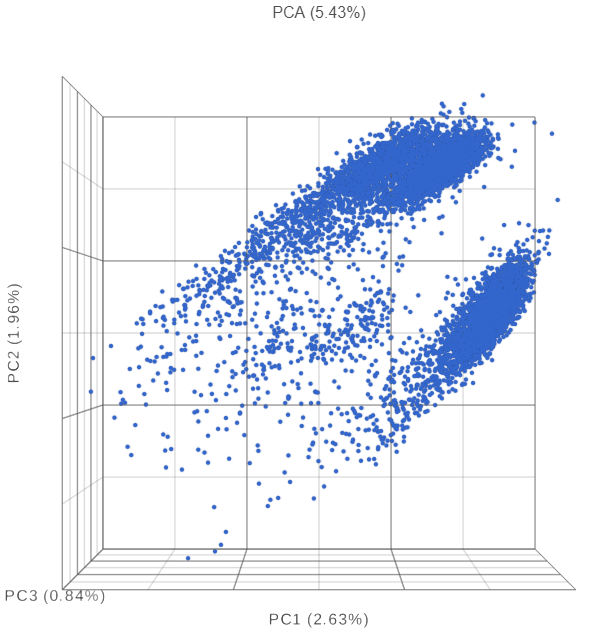 Image Added Image Added
|
- Click and drag the Scree plot from the Available plots card on the left onto the canvas
- Drop it over the Replace option (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | Click and drag the Scree plot to replace the PCA plot on the canvas |
|---|
| AnchorName | Replace PCA with Scree plot |
|---|
|
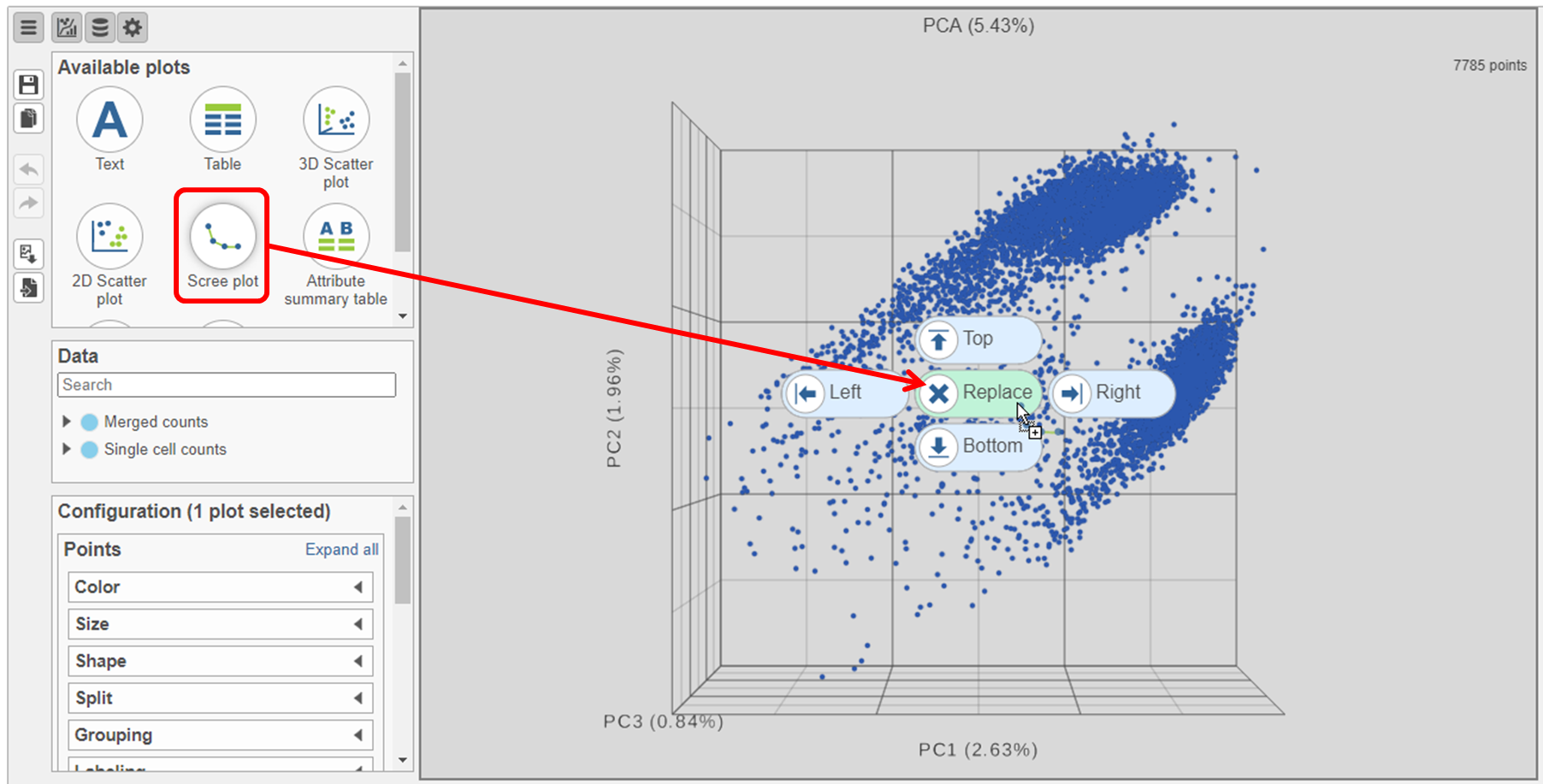 Image Added Image Added
|
- Select PCA as data for the new Scree plot
| Numbered figure captions |
|---|
| SubtitleText | The PCA data node contains the data to draw the Scree plot |
|---|
| AnchorName | Choose PCA data for Scree plot |
|---|
|
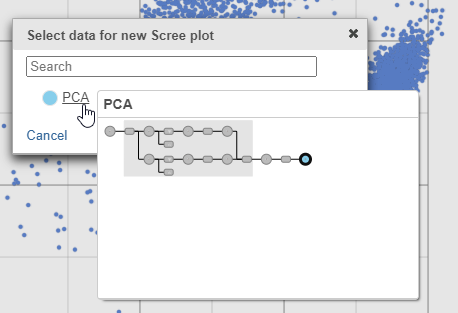 Image Added Image Added
|
The Scree plot (Figure ?) shows the eigenvalues on the y-axis for each of the 100 PCs on the x-axis. The higher the eigenvalue, the more variance explained by each PC. Typically, after an initial set of highly informative PCs, the amount of variance explained by analyzing additional components is minimal. By identifying the point where the Scree plot levels off, you can choose an optimal number of PCs to use in downstream analysis steps like graph-based clustering and UMAP.
| Numbered figure captions |
|---|
| SubtitleText | Scree plot shows the amount of variation explained by each principal component |
|---|
| AnchorName | Scree plot for merged CITE-Seq data |
|---|
|
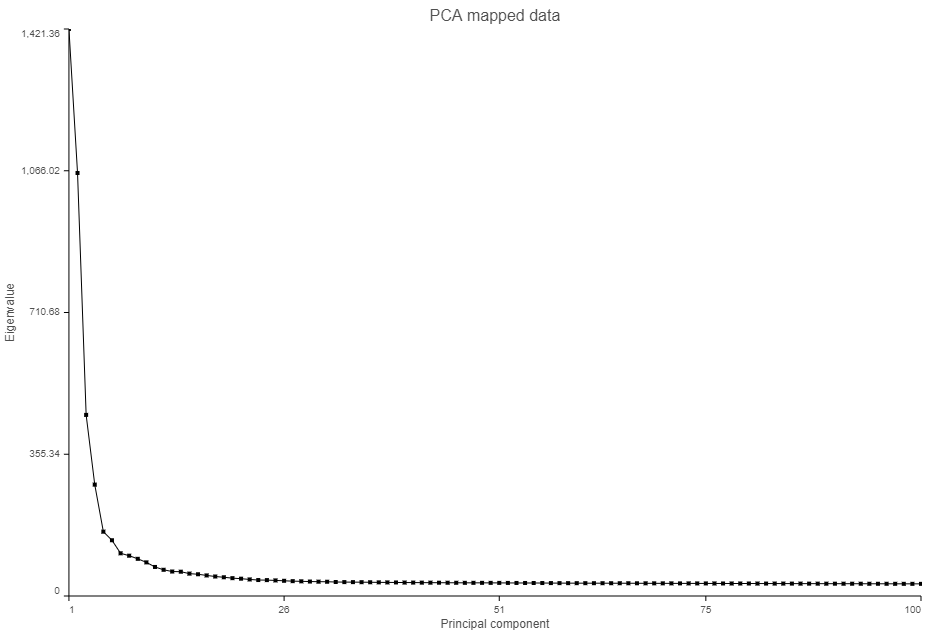 Image Added Image Added
|
- Click and drag over the first set of PCs to zoom in (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | Click and drag on the Scree plot to zoom in and see the first set of principal components |
|---|
| AnchorName | Scree plot zoom in |
|---|
|
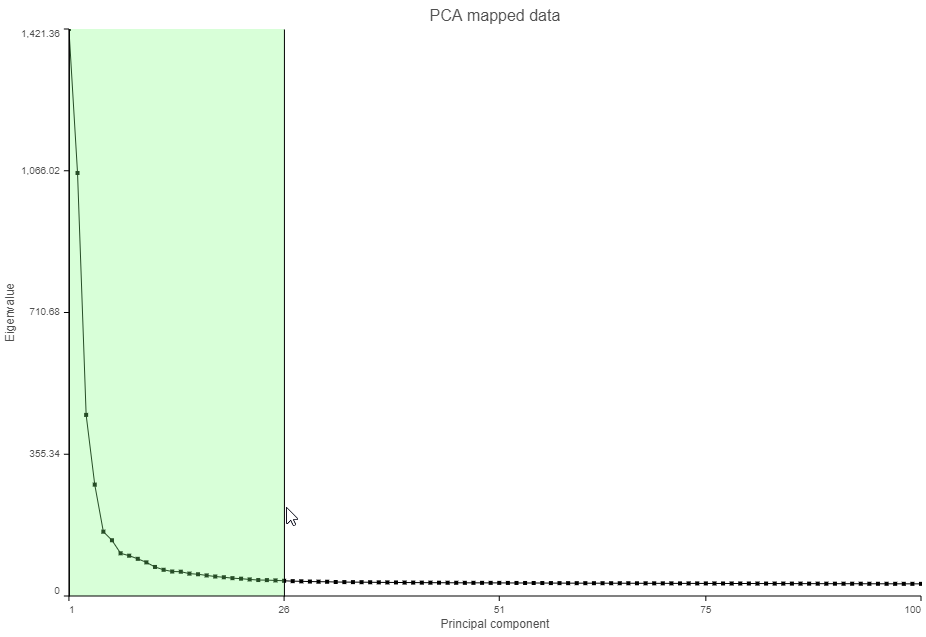 Image Added Image Added
|
- Mouse over the Scree plot to identify the point where additional PCs offer little additional information (Figure ?)
In this data set, a reasonable cut-off could be set anywhere between around 10 and 30 PCs. We will use 15 in downstream steps.
| Numbered figure captions |
|---|
| SubtitleText | Identifying the optimal number of PCs |
|---|
| AnchorName | Scree plot PC15 |
|---|
|
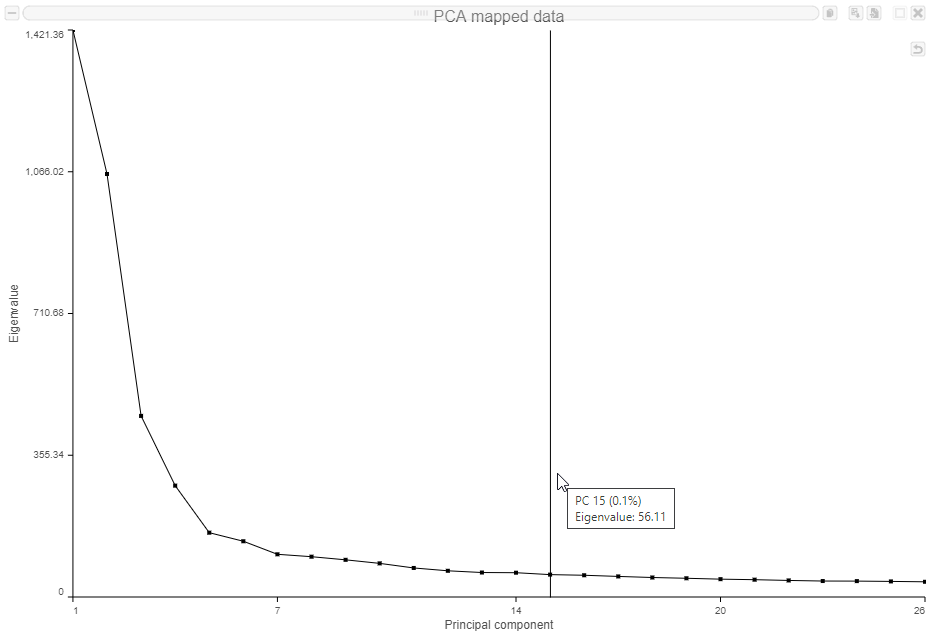 Image Added Image Added
|
Graph-based clustering
We can use Graph-based clustering to group similar cells together in an unsupervised manner.
- Click the project name near the top to go back to the Analyses tab
- Click the circular PCA data node
- Click Exploratory analysis in the toolbox
- Click Graph-based clustering
- Set the number of principal components to 15 (Figure ?)
- Click Finish to run the task
| Numbered figure captions |
|---|
| SubtitleText | Graph-based clustering task set up. Reduce the number of PCs to 15 |
|---|
| AnchorName | Graph-based clustering set up |
|---|
|
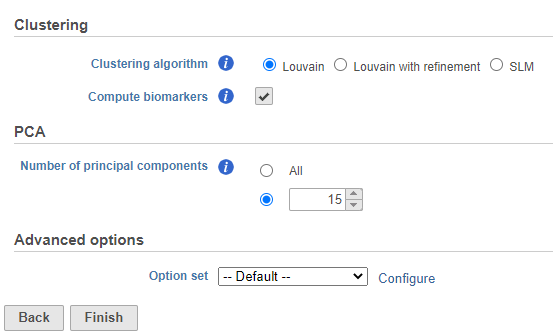 Image Added Image Added
|
A Graph-based clustering task node will be added to the pipeline under the Analyses tab and a circular Graph-based clusters output data node will be produced (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | Graph-based clustering task and output data nodes |
|---|
| AnchorName | Graph-based clustering output |
|---|
|
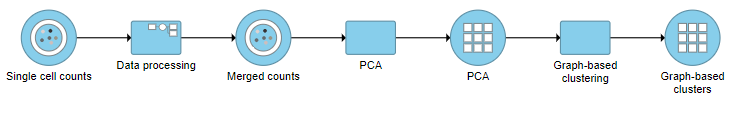 Image Added Image Added
|
UMAP
Once the graph-based clustering task hs completed, we can visualize the results with a UMAP plot.
- Click the circular Graph-based clusters data node
- Click Exploratory analysis in the toolbox
- Click UMAP
- Set the number of principal components to 15 (Figure ?)
- Click Finish to run the task
| Numbered figure captions |
|---|
| SubtitleText | UMAP task set up. Reduce the number of PCs to 15. |
|---|
| AnchorName | UMAP task set up |
|---|
|
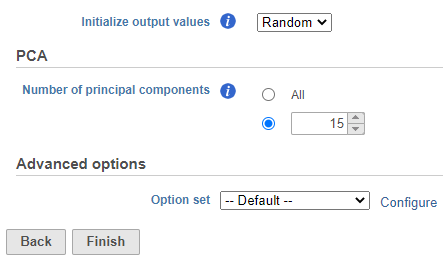 Image Added Image Added
|
A UMAP task node will be added to the pipeline under the Analyses tab and a circular UMAP output data node will be produced (Figure ?)
| Numbered figure captions |
|---|
| SubtitleText | UMAP task and output data node |
|---|
| AnchorName | UMAP output |
|---|
|
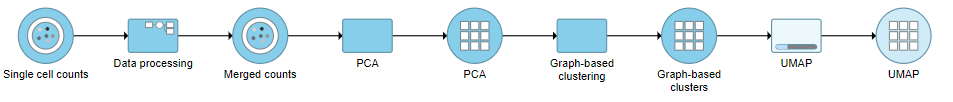 Image Added Image Added
|
...











