Page History
...
With a list of amplified or deleted regions in our cohort in hand, one of the more interesting questions to ask is what genes have recurrent amplifications or deletions in the data set. To address this question, we can use the Fine Find overlapping genes function to either add a column to our region list with the genes present in each region or create a new list of genes that overlap the regions.
...
- Select the amplified_or_deleted spreadsheet in the spreadsheet tree
- Select Find Overlapping Genes from the Copy Number Analysis section of the workflow
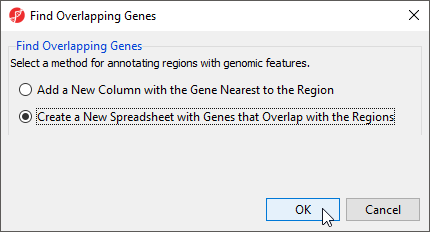
- Select Create a new spreadsheet with genes that overlap the regions from the Find Overlapping Genes dialog (Figure 1)
- Select OK
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
...
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
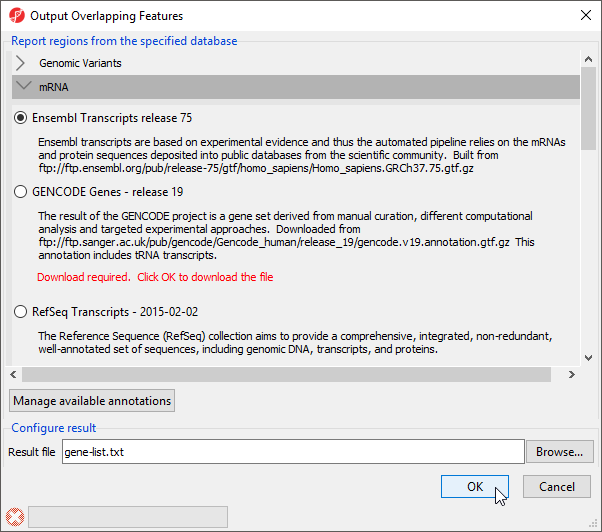
- Select Ensembl Transcripts release 75
- Select OK
...
This gene-list spreadsheet is gene-centric and enable enables genomic integration. For example, GO and Pathway encihment enrichment can be diretly directly invoked on the gene-list spreadsheet to detect the functional groups affected by copy number changes. Fo For rmore information on GO Enrichment enrichment analysis, please see the Gene Ontology Enrichment tutorial.
Another interesting use of this spreadsheet is to find possible fusion genes. If only a fraction of the gene as been amplified or deleted (column 8. Percent overlap with gene), it is possible that a translocation event took place and split the gene.
| Page Turner | ||
|---|---|---|
|
...