Page History
...
Partek Genomics Suite detects de novo motifs using the Gibbs motif sampler (Neuwald et al., Protein Science, 1995) and can search for known transcription factor binding sites using a database such as JASPAR.
Discover de novo
...
motifs
- Select Motif discovery from the Peak Analysis section of the ChIP-Seq workflow
- Select Discover de novo motifs
- Select OK
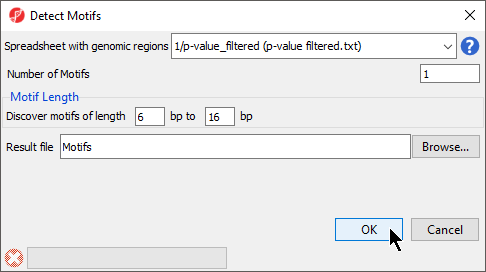
The Detect Motifs dialog will open to allow you to configure the search (Figure 1).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select 1/p-value_filtered from the Spreadsheet with genomic regions drop-down menu
- Set Number of Motifs to 1
- Set Discover motifs of lengthto 6 bp to 16 bp
- Set Result file to Motifs; the default save location is the folder you imported the .bam files from
- Select OK
...
7. score gives the log ratio of the probability that this sequence was generated by the motif versus the background distribution. A higher number indicates a better chance that the sequence is an instance of the motif.
Search JASPAR for known motifs
- Select Motif discovery from the Peak Analysis section of the ChIP-Seq workflow
- Select Search for known motifs
- Select OK
Search for known motifs will search the JASPAR database for motifs that are over-represented in the list of sequences in the significant regions list. The JASPAR database will download automatically if needed during the Search for known motifs step. Downloading the JASPAR database will create a spreadsheet in your experiment named JASPAR.txt that contains all of the species-specific motifs in the database. To visualize the motifs, right-click on a row in the JASPAR.txt spreadsheet and select Logo View.
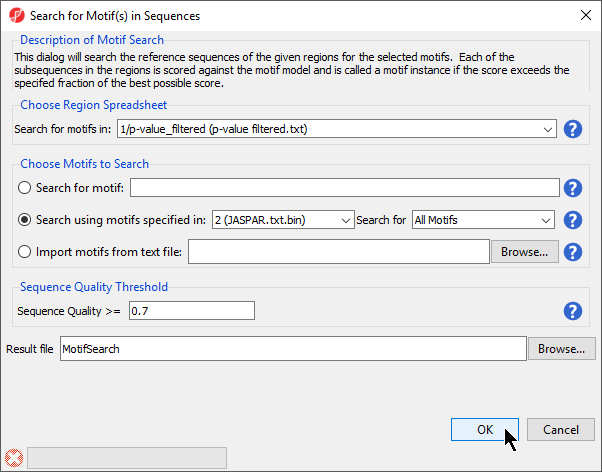
Before Search for known motifs runs, we need to configure the search (Figure 5).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select 1/p-value_filtered (p-value filtered.txt) from the Choose Region Spreadsheet drop-down menu
- Select Search using motifs specified in: for Choose Motifs to Search
- Set Search using motifs specified in: to 2 (JASPAR.txt) using the drop-down menu
- Set Search for to All Motifs using the drop-down menu
- Set Sequence Quality >= to 0.7
- Name the result file MotifSearch
- Select OK
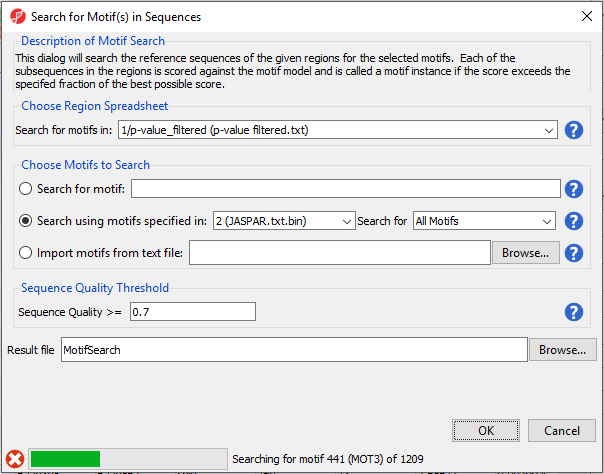
Because we are searching for around 1200 motifs, the process will take some time to complete. Progress is displayed in the progress bar in the lower left-hand side of the Search for Motif(s) in Sequences dialog (Figure 6).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
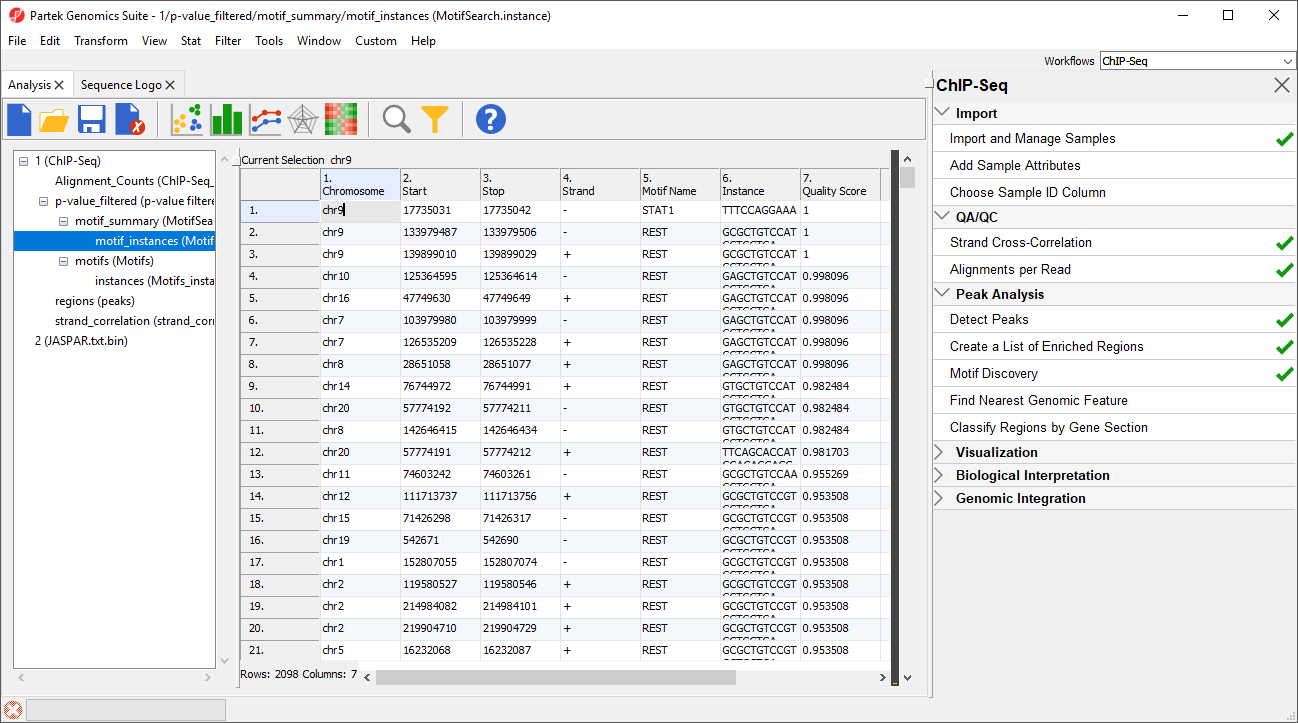
Two spreadsheets are created, similar to the spreadsheets in the de novo motif discovery, the motif_summary (MotifSearch) spreadsheet (Figure 6) and the motif_instances (MotifSearch.instance) spreadsheet.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
In the MotifSearch spreadsheet, each motif used in the motif search is shown. The columns detail the results of the search for each motif that was found in the reads.
1. Motif this is the name or ID of the motif
2. Probability of Occurrence gives the probability of detecting a false positive for this motif in a random DNA sequence
3. Expected Number of Outcomes gives the Probability of Occurrence multiple by the summed length of the reads
4. Actual Number of Occurrences gives a count of sequences that match the known motif in the reads
5. p-value is the uncorrected p-value (binomial test)
As you can see, REST, which is another name for NRSF, is near the top of the list as one of the most significantly over-represented motifs (Figure 6). This motif agrees with the motif found in the de novo motif detection step. Interestingly, other motifs appear a significant number of times in the ChIP-Seq peaks and may represent possible co-factors or regulators.
The motif_instances spreadsheet contains all instances of the motifs from the motif_summary spreadsheet in a format identical to the instances spreadsheet from de novo motif detection.
Generating a list of regions containing a motif
While the motif_instances spreadsheet contains every instance of every motif, it may be useful to create a spreadsheet with just instances of one motif or a select group of motifs. Let's do this for both REST motifs.
- Select the motif_instances spreadsheet in the spreadsheet tree
- Right-click the 5. Motif Name column
- Select Find / Replace / Select... from the pop-up menu (Figure 7)(
- Set Find What: to REST
- Select By Columns for Search:
- Select Only in column with 5. Motif Name selected form the drop-down menu
- Select Select All
This finds and selects every instance of REST in column 5. Motif Name.
- Select Close
In the motif_instances spreadsheet the selected columns are highlighted.
- Right-click on the first highlighted row visible; in this example, we see row 13196
- Select Filter Include from the pop-up menu
The spreadsheet will now include 2098 rows and a black and yellow bar will appear on the right-hand side of the spreadsheet (Figure 8). The black and yellow bar is a filter indicator showing the fraction of the spreadsheet currently visible as yellow and the filtered fraction as black.
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
To create a spreadsheet that contains only the REST instances, we can clone the motif_instances spreadsheet while the filter is applied.
- Right-click on motif_instances in the spreadsheet navigator
- Select Clone... from the pop-up menu
- Set the Name of resulting copy as REST
- Select 1/p-value_filtered/motif_summary (MotifSearch) from the Create as a child of spreadsheet drop-down menu
- Select OK
This creates a temporary spreadsheet rest from the filtered motif_instances spreadsheet. We can now save the new spreadsheet.
- Select rest from the spreadsheet tree
- Select () from the command bar
- Name the file REST
- Select Save
We can now remove the filter from the source motif_instances spreadsheet.
- Select motif_instances from the spreadsheet tree
- Right-click the filter bar
- Select Clear Filter
| Additional assistance |
|---|
|
| Rate Macro | ||
|---|---|---|
|