Page History
The significant CpG loci detected in the previous step actually form a methylation signature that differentiates , which we shall now visualise by a heat map.
- Select the
...
- (CpG
...
- of
...
- interest) spreadsheet in the spreadsheet pane on the left
- Select Cluster Based on Significant Genes from the Visualization panel of the Illumina BeadArray Methylation workflow
- Select Hierarchical Clustering for Specify Method (Figure 1)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select OK
- Verify that
...
- (CpG
...
- of
...
- interest) is selected in the drop-down menu
- Select Standardize for Expression
...
- normalization (Figure 2)
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
- Select OK
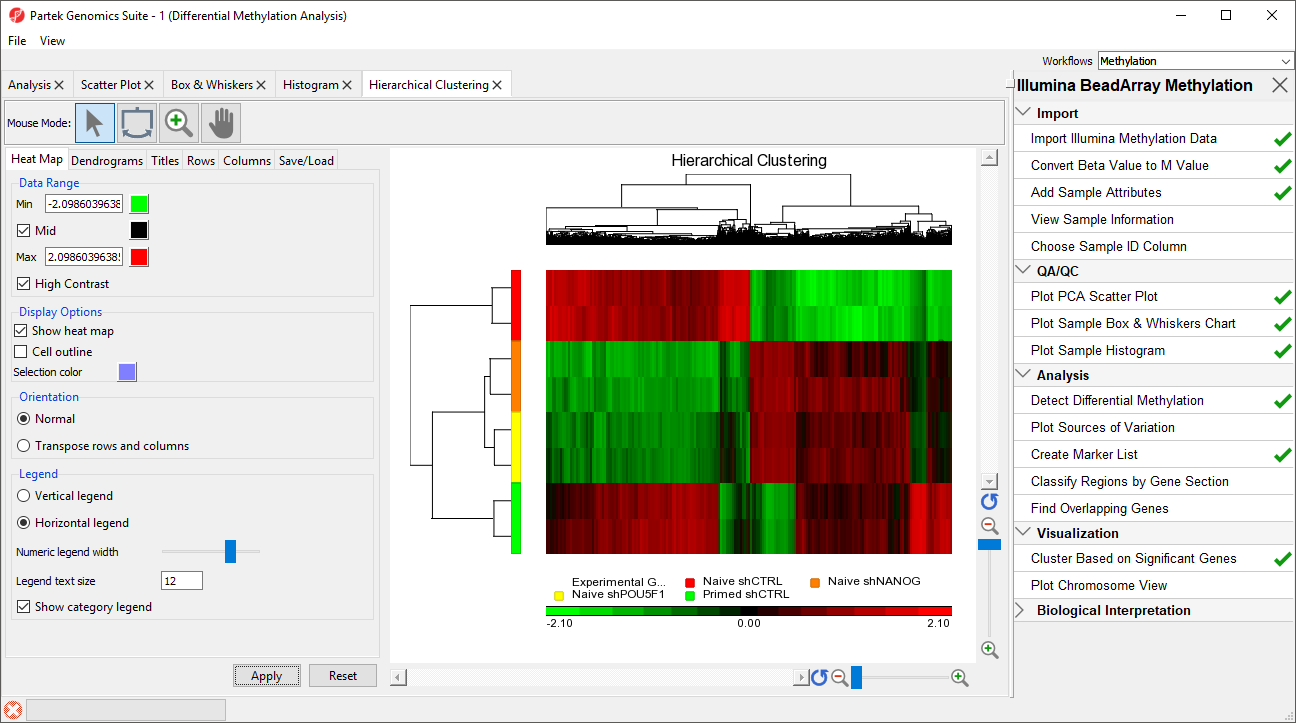
The heat map will be displayed on the Hierarchical Clustering tab (Figure 13).
| Numbered figure captions | ||||
|---|---|---|---|---|
| ||||
|
As expected, the Naive shPOU5F1, Naive shNANG, and Primed shCTRL experimental conditions cluster together; however, we can also see regions of similarity between Primed shCTRL and Naive shCTRL. This suggests that treatment with shRNA against POU5F1 or NANOG brings the methylation profile of naive hPSCs closer to that of primed hPSCs, but also causes additional changes not observed in either primed or naive hPSCs.
| Page Turner | ||
|---|---|---|
|
| Additional assistance |
|---|
|
| Rate Macro | ||
|---|---|---|
|
Overview
Content Tools