In addition to annotating regions with overlapping genes, other annotations can be to characterize the regions showing copy number variation.
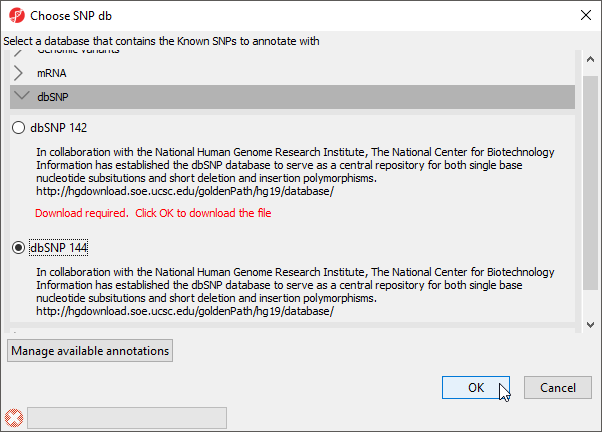
For example, Overlap with known SNPs in the Copy Number workflow gives the option of annotation regions with SNPs from dbSNP or a custom SNP database (Figure 1).
This task adds two column to the region list spreadsheet - the list of SNPs described in each region and the total number of SNPs in the region. If the list of SNPs is very long, you can output a separate list by right-clicking on the row header and select Create list of dbSNP from the pop-up menu.
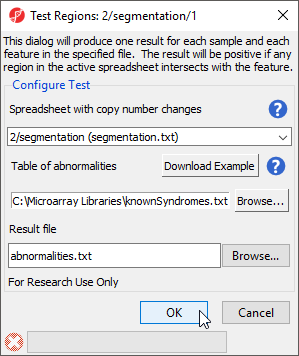
Another option in the workflow is Test for known abnormalities. Selecting this option compares the regions listed in the region list with a database of genomic abnormalities characteristic of particular diseases or syndromes to find possible matches. Annotation options include a Partek-distributed database of 60 syndromes or a custom database (Figure 2). Please note that the included table of known abnormalities is distributed for research use only.
If you like to add a custom database, organize the following information by column: the name of the abnormality, chromosome number, start location, and stop location. The input for the task should be a list of aberrations for every sample; do not include unchanged regions in the input or every syndrome will be shown as positive.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
35 | rates |