After samples have been added and associated with valid data files, in a Partek Flow project, a data node will appear in the Analyses tab (Figure 1). The Analyses tab is where different analysis tools and the corresponding reports are accessed.
Data and Task Nodes
The Analyses tab contains two elements: data nodes (circles) and task nodes (rounded rectangles) connected by lines and arrows. Collectively, they represent a data analysis pipeline.
Data nodes (Figure 2) may represent a file imported into the project, or a file generated by Partek Flow as an output of a task (e.g., alignment of FASTQ files generates BAM files).
Task nodes (Figure 3) represent the analysis steps performed on the data within a project. For details on the tasks available in Partek Flow, see the specific chapters of this user manual dedicated to the different tasks.
The Context Sensitive Menu
Clicking on a node reveals the context sensitive menu, on the right side of the screen.
Only the tasks that are available for the selected data node will be listed in the menu. For data nodes, actions that can be performed on that specific data type will appear.
In Figure 4, a node that contains Unaligned reads is selected (bold black line). The tasks listed are the ones that can be performed on unaligned data (QA/QC, Pre-analysis tools, and Aligners).
To hide the context sensitive menu, simply click the symbol on the upper left corner of the context sensitive menu. Clicking the triangles will collapse ( ) or expand ( ) the different categories of tasks that are shown.
After a task is performed on a data node, a new task node is created and connected to the original data node. Depending on the task, a new data node may automatically be generated that contains the resulting data. For details of individual tasks, see Task Menu.
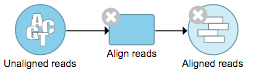
In Figure 5, alignment was performed on the unaligned reads. Two additional nodes were added: a task node for Align reads and an output data node containing the Aligned reads.
Running a Task
To run a task, select a data node and then locate the task you wish to perform from the context sensitive menu. Mouse over to see a description of the action to be performed. Click the specific task, set the additional parameters (Figure 6), and click Finish. The task will be scheduled, the display will refresh, and the screen will return to the project's Analyses tab.
In Figure 6, the STAR aligner was selected and the choices for the aligner index and additional alignment options appeared.
Tasks that are currently running (or scheduled in the queue) appear as translucent nodes. The progress of the task is indicated by the progress bar within the task node. Hovering the mouse pointer over the node will highlight the related nodes (with a thin black outline) and display the status of the task (Figure 7).
If a task is expected to generate data nodes, expected nodes appear even before the task is completed. They will have a lighter shade of color to indicate that they have not yet been generated as the task is still being performed. Once all tasks are done, all nodes would appear in the same shade.
Canceling and Deleting Tasks
Tasks can only be cancelled by the user that started the task or by the owner of the project. Running or pending tasks can be canceled by clicking the right mouse button on the task node and then selecting Cancel (Figure 8). Alternatively, the task node may be selected and the Cancel task selected from the context sensitive menu.
A verification dialog will appear (Figure 9) asking to confirm the task cancellation, the cancelled tasks will remain in the Analyses tab but will be flagged by gray x circles on the nodes (Figure 10).
Data nodes connected to incomplete tasks are also incomplete as no output can be generated (Figure 10).
To delete tasks from the project click the right mouse button on the task node and then select Delete (Figure 11). Alternatively, click the task node and select Delete task from the context sensitive menu.
A verification dialog will appear (Figure 12). A yellow warning sign will show up if there some downstream tasks performed by collaborators will be affected. Deleting the tasks output files optional. If this is not selected, the task nodes will disappear from the Analyses tab but the output files will remain in the project output directory.
Task Results and Task Actions
Selecting a task node will reveal a menu pane with two sections: Task results and Task actions (Figure 13).
Items from the Task results section inform on the action performed in that node. Certain tasks generate a Task report (Figure 14), which include any tables or charts that the task may have produced.
The Task details shows detailed diagnostic information about the task (Figure 15). It includes the duration and parameters of the task, lists of input and output files, and the actual commands (in the command line) that were run.
Additionally, the Task details page would contain the error logs of unsuccessful runs. The user can download the logs or send them directly to Partek. This page plays an important role in diagnosing and troubleshooting issues related to task.
Double clicking on a task node will show the Task report page. However, if no report was generated, the user will be directed to the Task details page.
In the Task actions sections, the selected task can be Re-run w/new parameters, and in case it is part of a pipeline that includes additional tasks after it, running the Downstream tasks is an option. Re-running tasks will result in a new layer being made in the Analyses tab.
Another action available for a task node is Add task description (Figure 16), which is a way to add notes to the project. The user can enter a description, which will be displayed when the mouse pointer is hovered over the task node.
Layers
It is common for next-generation sequencing data analysis to examine different task parameters for optimization. Users may want to modify an upstream step (e.g. alignment parameters) and inspect its effect on downstream results (e.g. percent aligned reads).
The implementation of Layers makes optimizations easy and organized. Instead of creating separate nodes in a pipeline, another set of nodes with a different color is stacked on top of previous analyses (Figure 17). To see the parameters that were changed between runs, hover the mouse icon over the set of stacked task nodes and a pop-up balloon will display them. The text color signifies the layer corresponding to a specific parameter.
Layers are formed when the same task is performed on the same data node more than once. They are also formed when a task node is selected and the Re-run it w/new parameters is selected in the context sensitive menu. This will allow the users to change the options only for the selected task. The user may choose to re-run the task to which the changes have been made, as well as all the downstream tasks until the analysis is completed. To do so, select Re-run w/new parameters, downstream tasks from the context sensitive menu.
To select a different layer, use the left mouse button to click on any node of the desired layer. All the nodes associated with the selected layer have the same color and when clicked will be displayed on the top of the stack.
Collapsing Tasks
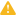
Addition of task and resulting data nodes to project may lead to creation of long pipelines, extending well beyond the border of the canvas (Figure 18).
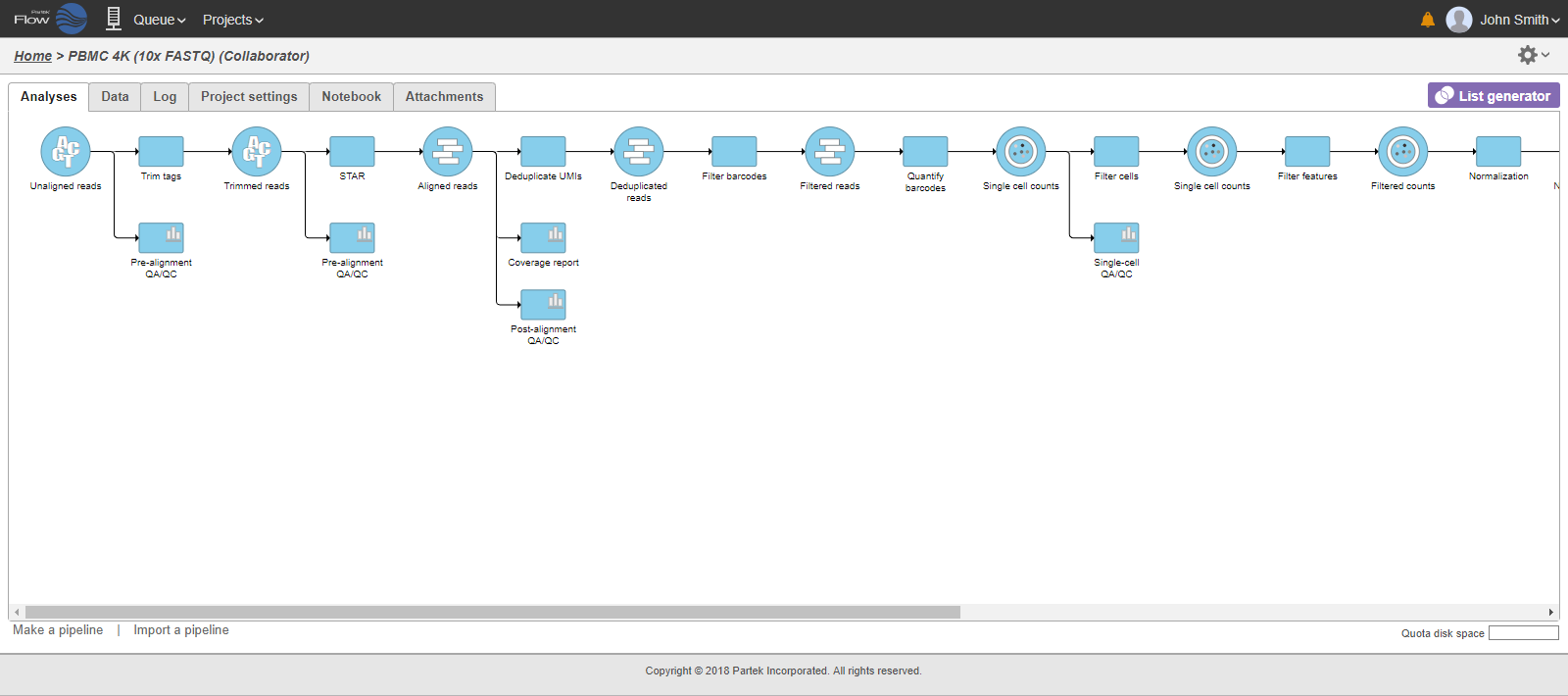
In that case, the pipeline can be collapsed, to hide the steps that are (no longer) relevant. For example, once the single-cell RNA-seq data has been quantified, Single cell counts data node will be a new analysis start point, as the subsequent analyses will not focus on alignment, UMI deduplication etc. To start, right-click on the task node which should become a boundary of the collapsed portion of your pipeline and select Collapse tasks (Figure 19).
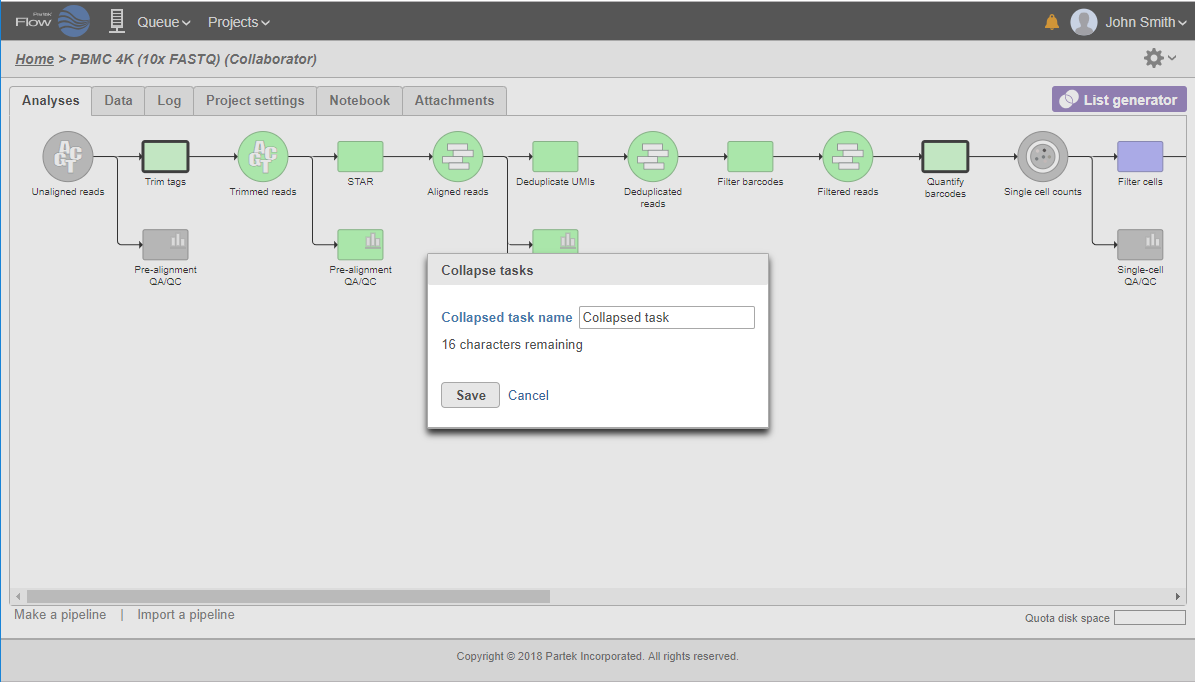
All the tasks on that layer will turn purple. Then left-click the task which should be the other boundary of the collapsed portion. All the tasks that will be collapsed will turn green and a dialog will appear (Figure 19). In the example shown in Figure 19, the tasks between Trim tags and Quantify barcodes will be collapsed. Give the collapsed section a name (up to 30 characters) and select Save (Figure 20).
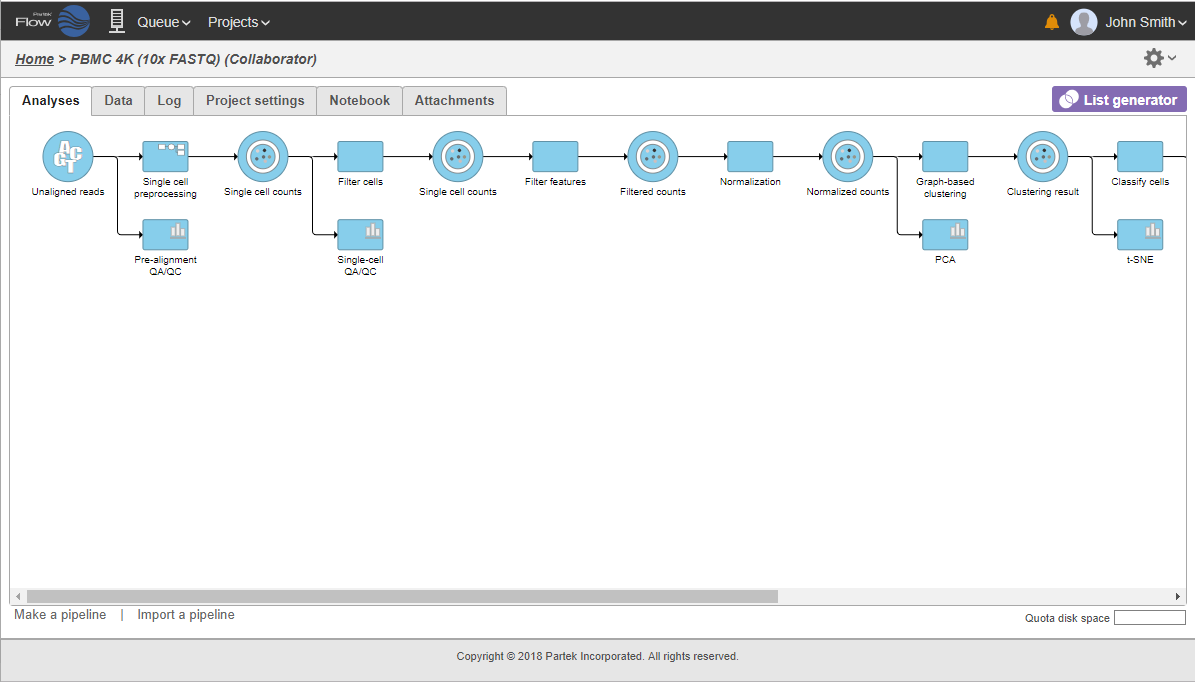
The collapsed portion of the pipeline is replaced by single task node, with a custom label ("Single cell preprocessing"; Figure 21).
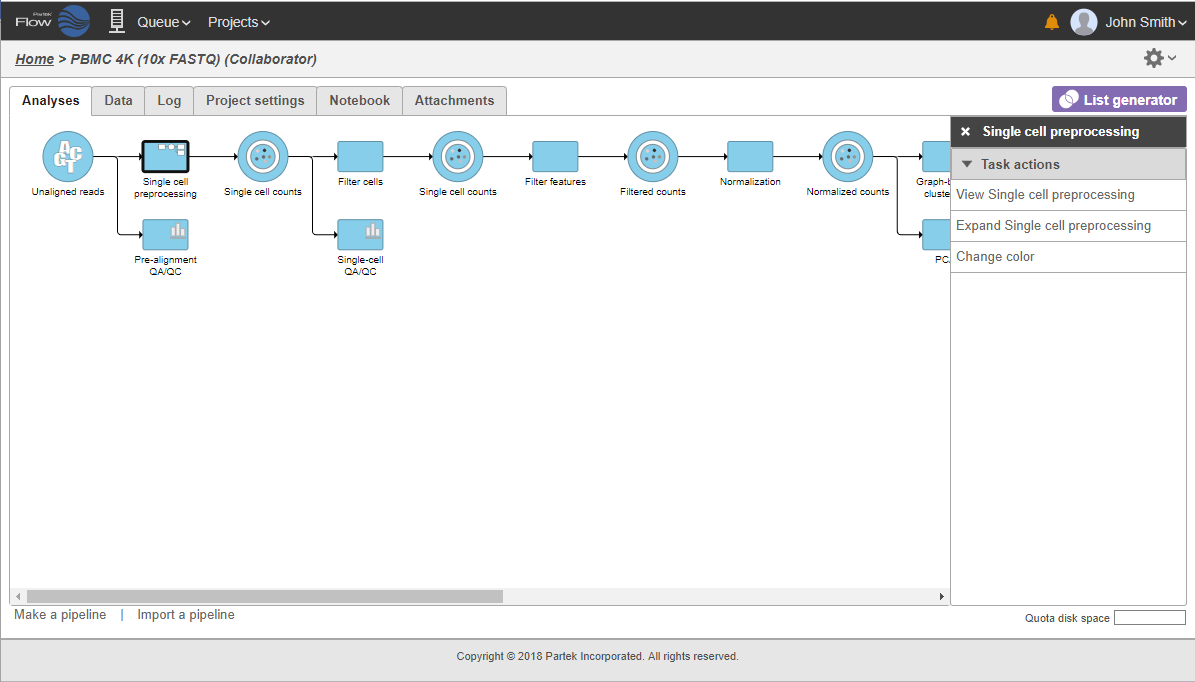
To re-expand the pipeline double click on the task node representing the collapsed portion of the pipeline. Alternatively, single click on the node and select Expand... on the context-sensitive menu. Within the same menu, you can also preview the contents of the collapsed task by selecting View... (Figure 22).
Downloading Data
Data associated with any data node can be downloaded using the Download data link in the context sensitive menu (Figure 23). Compressed files will be downloaded to the local computer where the user is accessing the Partek Flow server. Note that bigger files (such as unaligned reads) would take longer to download. For guidance, a file size estimate is provided for each data node. These zipped files can easily be imported by the Partek® Genomics Suite® software.
Additional Assistance
If you need additional assistance, please visit our support page to submit a help ticket or find phone numbers for regional support.


| Your Rating: |
    
|
Results: |
    
|
40 | rates |


















1 Comment
Melissa del Rosario
author: cignacio